
Concept explainers
Core Skill: Modeling The goal of this modeling challenge is to increase the complexity of the model shown in Figure 13.8d by including sequences in the crRNA and bacteriophage DNA that bind to each other.
Modeling Challenge: For simplicity, the structure of the crRNAs in Figure 13.8 is shown shorter than it really is. Let’s suppose that the part of a crRNA that recognizes the bacteriophage DNA is 20
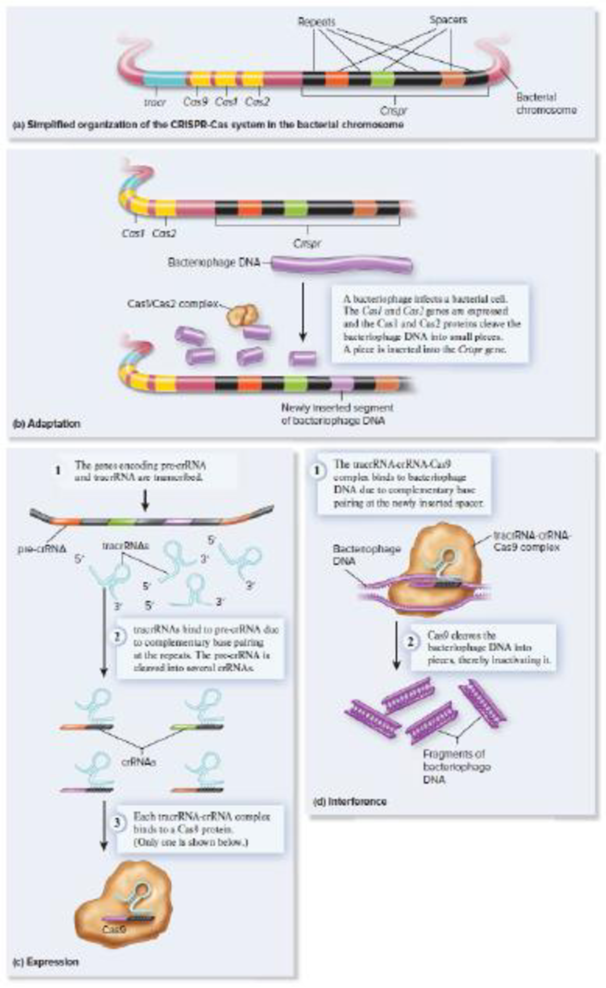
Figure 13.8 The CRISPR-Cas system of genome defense in bacteria. The system shown here is a type II system, which is found in the chromosome of certain bacterial species but not in archaea. (a) Organization of the CRISPR-Cas system in a bacterial chromosome. This drawing shows a typical organization, but different species have variations. (b–d) A simplified mechanism of the CRISPR-Cas system. The defense occurs in three phases, called adaptation, expression, and interference.
Want to see the full answer?
Check out a sample textbook solution
Chapter 13 Solutions
Biology
- Compare the possible differences between a eukaryotic protein-encoding gene cloned by PCR and the same gene cloned by reverse transcriptase PCR (RTPCR).arrow_forwardDiscuss how qPCR, DNA microarrays (DNA chips), and RNA-seq analysis are used to compare levels of gene expression among normal versus diseased tissues.arrow_forwardDescribe the features of a typical CRISPR locus in a bacterial cell.arrow_forward
- Describe how genome-wide association studies have led to new understandings about the structure and function of the genome.arrow_forwardExplain how targeted gene silencing and knockout mutations can give insights into the functions of a gene.arrow_forwardQ. When dsRNA is treated with Dicer enzyme siRNA and miRNA’s are produced. What role they will play in gene regulation and expression and how? (Subject Bioinformatics)arrow_forward
- Please help witht this homework question A- What are the topological parameters (linking number, twist, and writhe)for a relaxed, 4,200 base pair circular double-stranded DNA plasmid? LK=? Tw= ? Wr=? The CRISPR-Cas9 protein first forms a protein-RNA complex with a guide RNA, and then binds to DNA sequences that match a 20-base target sequence within the guide. When the CRISPR-Cas9 complex binds to DNA, it unwinds about 20 base pairs of the DNA double helix. B- Why is DNA unwinding required for CRISPR-Cas9 protein-RNA complexto recognize target sites? C-How would the topological parameters (linking number, twist, andwrithe) of the 4,200 bp plasmid in (A) change when the CRISPR-Cas9 complex binds the DNA and unwinds about 20 bp? Lk=? Tw=? Wr=? D- What is the linking number after the CRISPR-Cas9 complex cleaves theDNA?arrow_forwardNeed help ASAP. How do you use FISH(fluorescence in situ hybridization) to detect gene rearrangement? Describe an example with the detail of the disease and the procedure.arrow_forwardQuestion:- Define, compare, and contrast the utility of microarray and RNAseq while analyzing gene expression levels.arrow_forward
- Need help fast Explain briefly how bacteriophages causes mutation .arrow_forwardQ1. Genome-wide RNAi screens target expression of > 16,000 genes. Explain how each of these 16,000+ bacterial strains would be engineered in order that they only cause gene silencing of the intended target.arrow_forwardBIOC 385 RNA Processing and Silencing Q11.2: Describe the biochemical mechanism by which RNA splicing occurs precisely at the exon-intron borders considering that a single nucleotide offset by a mistake in splicing will reset the open reading frame register for the encoded protein.arrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
