
Concept explainers
To estimate: The phylogeny of the given taxa by plotting the changes on each of the possible unrooted trees and determine the tree with fewest evolutionary changes.
Introduction: Three species named 1, 2, and 3 are endemic to a group of islands. They all share a common ancestor named species 4. Species 4 serves as an out group. It has a huge
Explanation of Solution
Deoxyribonucleic acid consists of four nucleotides, these are: adenine (A), thymine (T), guanine (G), and cytosine(C). These nucleotides can be paired and labeled for the given condition as follows:
- GC - a
- AG - b
- GT - c
- AC -d
- AT - e
- TG - f
- CA - g
The data of the nucleotide bases at the ten sequenced sites in the given four species are as follows:
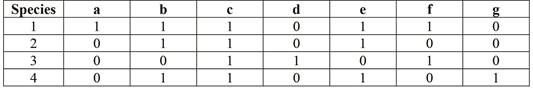
Pictorial representation: Fig.1, Fig.2 and Fig.3 represent the possible unrooted trees for phylogeny of the given taxa.
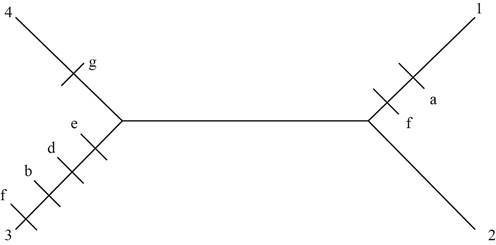
Fig 1: Seven changes observed in this unrooted tree
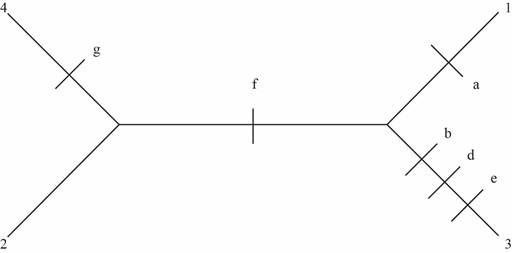
Fig 2: Six changes observed in this unrooted tree

Fig 3: Seven changes observed in this unrooted tree
Therefore, the tree with fewest evolutionary changes is the second unrooted tree represented by Fig.2. This tree has only six changes, whereas the other two have seven changes which are higher than the second one.
Want to see more full solutions like this?
Chapter 2 Solutions
Evolution
- Shown above are three possible phylogenetic trees for species I, II and III reconstructed based on the 4-nucleotide DNA sequences given in the righthand table. In every tree, each hatchmark on a branch represents a single base-change event. The most parsimonious tree would be - A. Both X and Y. B. X. C. Y. D. Both Y and Z. E. Z.arrow_forwardCan you answer all the parts to this diagram Species 1 and 2 are sister species from which you’ve cloned related genes. On the gene tree on the top of the next page, use labels to answer the following questions: (a) Label the node that represents a gene duplication with “D,” (b) Label the nodes that represent speciation events with “S,” (c) Pick a pair of genes that are paralogs and label them both “P.” (d) Pick a pair of genes that are orthologs and label them both “O.”arrow_forwardFor each of the following examples, discuss whether the observed result is due to neutral mutations or mutations that have been acted on by natural selection, or both: A. When comparing sequences of homologous genes, differences in the coding sequence are most common at the wobble base (i.e., the third base in each codon). B. For a protein-encoding gene, the regions that encode portions of the polypeptide that are vital for structure and function are less likely to display mutations than other regions of the gene. C. When comparing the sequences of homologous genes, introns usually have more sequence differences than exons.arrow_forward
- If not all mutations that contribute to species evolution are passed down, what conditions must be met for a mutant trait to be inherited by the next generation?arrow_forwardImagine that you have the DNA sequences from the intron of a gene in three species called A, B, and C. Species A and B are most closely related, while C is more distantly related. The sequences of A and B differ by 18 base pairs, A and C differ by 26 base pairs, and B and C differ by 28 base pairs. Fossils show that species A and B diverged about 1.2 Mya, but there is no fossil evidence as to when the most recent common ancestor of all three species lived. (Draw a simple tree to help you think about the problem) Use the genetic data to estimate that date (most recent common ancestor). HINT = use Eqn 7.1, several times- first to estimate mutation rate. Then to estimate the unknown time since divergencearrow_forwardImagine that you have the DNA sequences from the intron of a gene in three species called A, B, and C. Species A and B are most closely related, while C is more distantly related. The sequences of A and B differ by 18 base pairs, A and C differ by 26 base pairs, and B and C differ by 28 base pairs. Fossils show that species A and B diverged about 1.2 Mya, but there is no fossil evidence as to when the most recent common ancestor of all three species lived. (Draw a simple tree to help you think about the problem) Use the genetic data to estimate that date (most recent common ancestor). What assumptions are you making to get this estimate?arrow_forward
- By comparing DNA sequence of a specific gene, we can determine the evolutionary relationship of two organisms. Based on the sequence listed below, which two species would you expect to be more closely related? Organism No.1 with ATG CAA TAC GCC, organism No. 2 with ATG CAT GAC ACC and organims No. 3 with ATG CAT TAC GCC A. Organims No.2 and No.3 are likely more closely related. b.Organims No.1 and No.2 are likely more closely related. c.Organims No 1 and No.3 are likely more closely related.arrow_forwardThere 27 sequences from 27 individuals belonging to an unidentified group of organisms. Eleven (11) sequences were mined from NCBI while the rest are unpublished sequences from Mindanao. Sequences labeled with "SSL" are from Agusan Marsh while sequences labeled with "CKL/CITLR" and "CWL" are from Camiguin Island and Dinagat Islands respectively. Finally, sequences labeled with "MSLA" are from Mt. Magdiwata. (a) What species is considered as the outgroup? (b) What genetic marker is utilized to generate the sequences? (c) Which specimen group is more closely related to "SSL"? CKL or MSLA? Justify your answer. (d) Based on the BLAST results, what possible species name can be assigned to the MSLA group?arrow_forwardWere any of your physical traits autapomorphic or synapomorphic when plotted on the gene trees? Which ones and for which species? Phylogeneticists often refer to these physical traits as “evolutionarily significant,” what do you suppose they mean by this? Were any of your physical traits analogous? Which ones? Why do you suppose some traits can occur multiple times on a tree while others don’t? Given your results from two genes and physical traits, what relationships between species are you certain of? Which ones are you uncertain of? Why? If gene trees have more information in terms of base pairs for generating phylogenies, why do you suppose phylogeneticists even bother using and including physical traits in their analyses?arrow_forward
- If you were comparing the karyotypes of species that are closely related evolutionarily, what types of similarities and differences would you expect to find?arrow_forwardWere any of your physical traits autapomorphic or synapomorphic when plotted on the gene trees? Which ones and for which species? Phylogeneticists often refer to these physical traits as “evolutionarily significant,” what do you suppose they mean by this?arrow_forwardDo the data in the graph indicate that the mutation rates per base-pair in some taxa, such as mammals, are not at the lowest possible rate? Is this evidence that a certain level of mutations is an adaptation? Why or why not?arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
