
Concept explainers
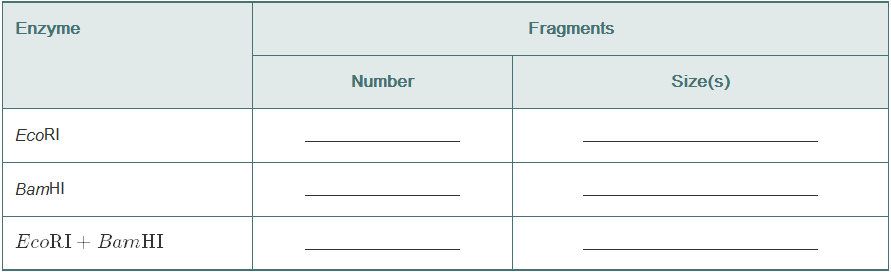
Using this map of pLAB30 (18 kb), give the number and lengths of the restriction fragments that would result from digesting pLAB30 with EcoR1, with BamHI, and with both enzymes together.

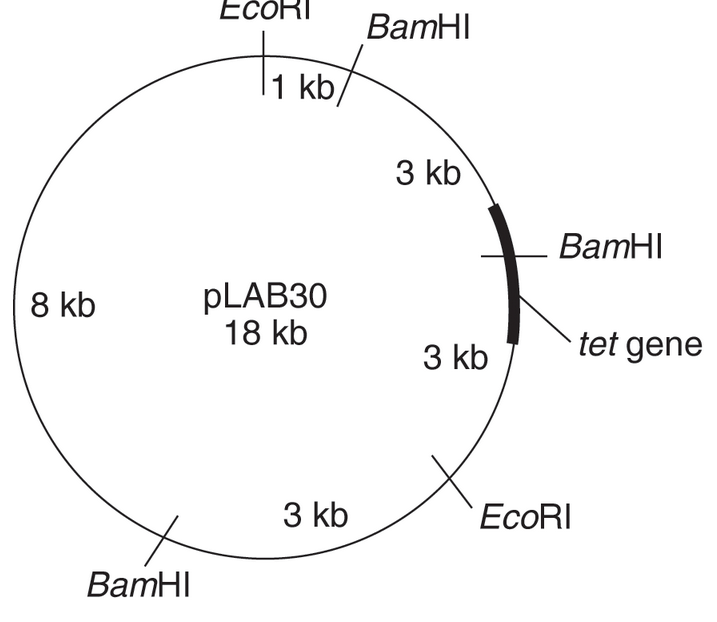
Which enzyme will give the smallest piece containing the tetracycline-resistance gene?
To determine:
The size and number of DNA Fragments given by restriction enzymes Eco RI, Bam HI, and both these enzymes together.
Introduction:
Restriction enzyme is an enzyme that cuts the DNA fragment at the specific site by recognizing the DNA sequence at that site. The site at which the DNA is cut is termed as the restriction site.
Explanation of Solution
pLAB30 is an 18 kb map, which is cut by two restriction enzymes, Eco RI, and Bam HI. When Eco RI acts, then it cuts the DNA fragment at two sites, giving an 11 kb fragment and 7 kb fragment of the DNA.
When Bam HIacts, then it cuts DNA fragment at three sites, giving three fragments. The size of these fragments is 9 kb, 3 kb, and 6 kb.
When both these restriction enzymes act together, then 5 DNA fragments are obtained. The size of these fragments is 1 kb, 3 kb, 3 kb, 3kb, and 8 kb.
The table showing the size and number of DNA fragments after the DNA is acted upon by restriction enzymes is shown below:
| Restriction enzyme | Number of DNA fragments obtained | Size of DNA fragments |
| Eco RI | 2 | 11 kb and 7 kb |
| Bam HI | 3 | 9 kb, 3 kb, and 6 kb |
| Eco RI + Bam HI | 5 | 1 kb, 3 kb, 3 kb, 3kb, and 8 kb |
Want to see more full solutions like this?
Chapter 30 Solutions
Laboratory Experiments in Microbiology (12th Edition) (What's New in Microbiology)
- Knowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardYour cloning vector has restriction recognition sites for two restriction endonucleases, EcorI and BamHI. However, the DNA to be manipulated does not have recognition sites for these two restriction endonucleases. How would you construct a recombinant DNA for the given DNA?arrow_forwardHow would you expect your results to change if the enzyme did not have time to cut all of the sites? (Hint, remember that there are many pieces of the lambda DNA in the tube – draw yourself a simplified model where there are four restriction sites, and imagine what would happen if the enzyme only cut at one, two, three or all four, randomly, each time). How would you expect your results to change if you treated them with both EcoRI and HindIII in the same tube? Why might it be safer for the virus to have its DNA become circular when it enters the host, rather than remain linear?arrow_forward
- Give the recognition sequences for each of the restriction enzymes (a) PagI, (b) AluI, (c) PstI, and (d) RcaI. Show the sequence in double-stranded DNA (dsDNA) format, indicate the cleavage position with a ^, and mention which types of ends are generated in each case (blunt or sticky; 5’ or 3’ overhang)? Recognition sequence for a)PagI cleaves DNA at the recognition sequence 5' ---T ^CATGA--- 3' b) AluI cleaves DNA at the recognition sequence 5' AG^CT 3' c) PstI cleaves DNA at the recog.arrow_forwardA group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardWhat detection method could be used to determine a different restriction enzyme that is used in splicing the gene of interest was then used for splicing the vector to be used in producing a recombinant molecule which will produce an enzyme X? What results could be achieved from that?arrow_forward
- Describe the process of cloning a DNA fragment into theBamHI and PstI sites of the vector pUC18. How would youscreen for clones that contain an insert? and explain the process(steps) by drawingarrow_forwardAssume that a circular plasmid is 3200 base pairs in length and has restriction sites for HindIII restriction enzyme at the following locations: 400, 700, 1400, 2600. Give the expected sizes of the restriction fragments following complete digestion.arrow_forwardExplain the process of how X-gal screening works with pUC19, you may build a model with boxes and arrows. 2. You are utilizing BamHI (GGATCC) restriction site and HindIII (AAGCTT) restriction site. Within pUC19, BamHI is at position 263, while HindIII is at position 233. (Hint: position is like coordinate on a map). If you manage to insert GTF2H5 in pUC19 vector, what are the sizes of fragments if you digest the pUC19-GTF2H5 (this is after insertion) with the following restriction enzyme combination after gel electrophoresis: BamHI and HindIII There is an NdeI site right in the middle of the GTF2H5 that has been inserted in pUC19, what would be the fragment sizes, if you digest with BamHI and NdeI. HindIII and NdeI BamHI, HindIII, and NdeI. 3. Design an experiment to confirm the presence of insert GTF2H5 in pUC19 vector using the following method (besides restriction analysis above), assuming that you know the sequence of GTF2H5: Southern Blot Polymerase Chain Reactionarrow_forward
- A 2.0kb bacterial plasmid ‘BS1030’ is digested with the restriction endonuclease Sau3A; the plasmid map is depicted in the diagram below and the Sau3A (S) restriction sites are indicated. Which of the following DNA fragments do you expect to see on an agarose gel when you run Sau3A-digested plasmid ‘BS1030’ DNA? a. 250 bp, 450 bp, 550 bp, 1.1 kb, 1.5 kb and 2.0 kb b. 2.0kb c. 250 bp, 400 bp, 450 bp, 500 bp and 550 bp d. 100 bp, 200 bp, 250 bp, 400 bp, 500 bp and 550 bparrow_forwardThe restriction enzymes Kpn I and Acc 65I recognize and cleave the same 6-bp sequence. However, the sticky end formed from Kpn I cleavage cannot be ligated directly to the sticky end formed from Acc 65I cleavage. Explain why.arrow_forwardDraw and explain how the recombinant DNA molecules can be produced in a pUC19 vector using a linear DNA (dummy) at Pst1 5'_CTGCA/G_3' restriction site?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





