
Concept explainers
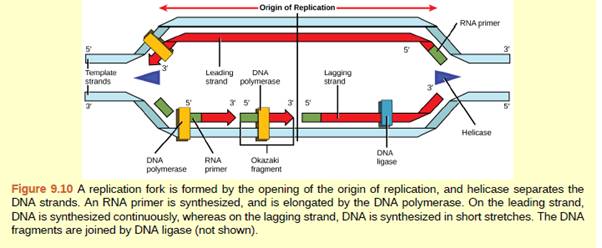
Figure 9.10 You isolate a cell strain in which the joining together of Okazaki fragments is impaired and suspect that a mutation has occurred in an enzyme found at the replication fork. Which enzyme is most likely to be mutated?
To write:
The enzyme which is most likely mutated when joining of Okazaki fragments is impaired in the replication.
Introduction:
DNA replication is the process in which the copy of DNA is made. This is done in 3 steps in a eukaryotic cell. They are initiation, elongation, and termination. It is a semiconservative type of replication. Enzymes involved in this process are helicase, DNA polymerase, and DNA ligase.
Explanation of Solution
DNA replication is started at the origin of replication where a Y shaped fork known as replication is formed when both strands are separated by helicase enzyme. Both the strands of DNA act as a template. One strand is known as a leading strand at which replication is continuous and other strand is known as a lagging strand at which replication occurs in fragments. This is because DNA polymerase can synthesize DNA only in 5' to 3' direction. These fragments are known as Okazaki fragments. These fragments at the lagging strands are then sealed by the enzyme DNA ligase. The joining of Okazaki fragments is impaired when the DNA ligase is not able to join the strands. This can occur due to the mutation in enzyme DNA ligase.
When joining of DNA fragments on the lagging strand is impaired in DNA replication, the mutation is most likely to occur in DNA ligase as it is an important enzyme which joins the strands.
Want to see more full solutions like this?
Chapter 9 Solutions
Concepts of Biology
Additional Science Textbook Solutions
College Physics
Human Biology: Concepts and Current Issues
Anatomy & Physiology
Essentials of Genetics (9th Edition) - Standalone book
Campbell Biology in Focus
Microbiology with Diseases by Body System (4th Edition)
- The sequence below shows the ends of one strand of a linear chromosome, with slashes representing the middle part, which is not shown. During replication of this one strand, on which side of the slashes will Okazaki fragments be made in the newly synthesized strand? 5' AGCCGTACGGTTATCTCCTAG //// GGGCCTATTGTGACCAGTGAGTCG 3' a) Both sides b) Neither side c) The right side d) The left sidearrow_forwardThe following diagrams represent DNA molecules that are undergoing replication. Draw in the strands of newly synthesized DNA and identify (a) the polarity of the newly synthesized strands, (b) the leading and lagging strands, (c) Okazaki fragments, and (d) RNA primers.arrow_forwardShown below is a long template strand of DNA where lagging strand DNA synthesis is occurring. The short horizontal lines represent two Okazaki fragments that have already been made. In the context of the replication fork, select the letter(d–g) that indicates where primase will synthesize the next RNA primer. Explain why did you choose that location?arrow_forward
- The speed of DNA replication at a replication fork is about 100 nucleotides per second in human cells. What is the minimum number of origins of replication that a human cell must have if it is to replicate its DNA once every 24 hours?arrow_forwardDescribe in order, the four repeating steps that repeat over and over on the discontinuous lagging strand of DNA replication and name the major proteins required to carry out each of these steps in E. coli The first function is given. I. Function: Create RNA primer Enzyme: II. Function: Enzyme: III. Function: Enzyme: IV. Function: Enzymearrow_forwardWhich of the following is NOT a function of DNA polymerase I in E. coli?Question 1 options: A) 5' exonuclease B) polymerase C) helicasearrow_forward
- Which of the following is not depicted in the diagram attached? A. Okazaki fragment B. Replication fork C. Leading strand D. Origin of replicationarrow_forwardA genetics instructor designs a laboratory experiment to study the effects of UV radiation on mutation in bacteria. In the experiment, the students spread bacteria on petri plates, expose the plates to UV light for different lengths of time, place the plates in an incubator for 48 hours, and then count the number of colonies that appear on each plate. The bacteria that have received more UV radiation should have more pyrimidine dimers, which block replication; thus, fewer colonies should appear on the plates exposed to UV light for longer periods. Before the students carry out the experiment, the instructor warns them that while the bacteria are in the incubator, the students must not open the incubator door unless the room is darkened. Why should the bacteria not be exposed to light?arrow_forward"In E. coli, where the replication fork travels at 500 nucleotide pairs per second, the DNA ahead of the fork-in the absence of topoisomerase-would have to rotate at nearly 3000 revolutions per minute" is true or false.arrow_forward
- Which DNA repair systems you think might be capable of repairing a situation where T is in one strand and G is in the complementary strand? Explain dramatically.arrow_forwardWhich process does NOT use a specific nucleotide sequence for activity? please explain the answer a.replication initiation b.DNA helicase unwinding c.transcriptional termination d.translational terminationarrow_forwardBelow is a sample of a segment of DNA…(copy from left to right) 3’ TACAATGGGCGACGCGCTTCGTTTCAGATT 5’ 5’ ATGTTACCCGCTGCGCGAAGCAAAGTCTAA 3’ 1.Assume the 6th amino acid is changed from T to G on the DNA template strand. What type of mutation is this? What effect would this have on the protein? Look up an example for this type of mutation. 2, Assume the 5th and 6th amino acids are removed from the DNA template strand. What type of mutation is this? How would this affect the protein? Look up an example of this type of mutation. 3.Which mutation changes the protein more...a point mutation or a frameshift mutation. Explain your reasoning. 4.What would be the problem if ATT was inserted into the DNA template strand after the second codon? (Be sure to consult the coding chart for amino acids). 5. What if the second amino acid was repeated over 5Ox. What amino acid is repeated? What type of mutation is this? If this is on chromosome 4, what genetic disorder is this?…arrow_forward
 Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning

