What are Nucleic Acids?
Nucleic acids are essential biomolecules present in prokaryotic and eukaryotic cells and viruses. They carry the genetic information for the synthesis of proteins and cellular replication. The nucleic acids are of two types: deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). The structure of all proteins and ultimately every biomolecule and cellular component is a product of information encoded in the sequence of nucleic acids. Parts of a DNA molecule containing the information needed to synthesize a protein or an RNA are genes. Nucleic acids can store and transmit genetic information from one generation to the next, fundamental to any life form.
Discovery of Nucleic Acids
Nucleic acids were first isolated from the nuclei of white blood cells in 1868 by Swiss physician Friedrich Miescher. He called this phosphorous-containing material "nuclein" isolated from the nucleus, a darkly stained part of the cell. The material had acidic properties and is hence termed "nucleic acid." The "nuclein" was later on known to be DNA. In the 1940s, following scientists such as Ostwald T.Avery, Colin MacLeod, and Maclyn McCarty, it was understood that DNA was the cell's genetic material. In 1952, Alfred D. Hershey and Martha Chase confirmed that DNA and not proteins carried the genetic information. They performed experiments and studied bacterial infection using radioactively labeled DNA or protein-containing virus (bacteriophage).
In the late 1940s, Erwin Chargaff and his colleagues' experiments gave the world insights into DNA base composition's properties and quantitative relationships. Their studies revealed that the DNA base composition a) varied between species, b) remained the same for DNA isolated from different cell types or tissues of a species, c) remained the same in a species and did not alter with age, nutritional or environmental factors, and d) in all DNA, regardless of the species the number of adenosine (A) bases was always equal to thymidine (T) (A=T), and the number of guanines (G) residues was always equal to cytosine (C) residues (G=C). Thus, the sum of the purine residues is equal to the sum of the pyrimidine residues (A+G = T+C), known as the "Chargaff's rule."
In the early 1950s, Rosalind Franklin and Maurice Wilkins conducted X-ray diffraction of DNA fibers. They showed that DNA had a distinct helical structure with two periodicities along its long axis. Then, in 1953, James Watson and Francis Crick proposed the double-helical 3D structure of DNA.
Chemical Composition of the Nucleic Acids
Nucleic acids are polymers composed of nucleotide units. Nucleotides are made of three distinct components:
- a nitrogenous base
- a pentose (5-carbon monosaccharide/simple sugar)
- one or more phosphate groups
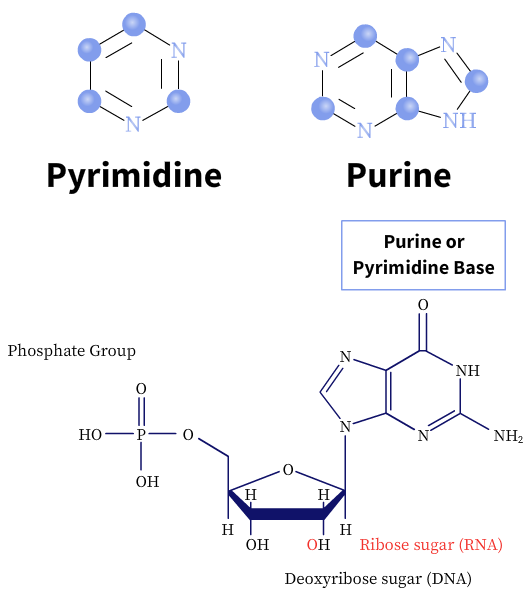
The nitrogenous bases are heterocyclic and derived from two-parent compounds, purine or pyrimidine. There are two major purine bases, adenine and guanine, and three commonly known pyrimidine bases, cytosine, thymine, and uracil (U). While adenine and cytosine are present in both DNA and RNA, thymine is found mostly in DNA molecules and uracil in RNA molecules.
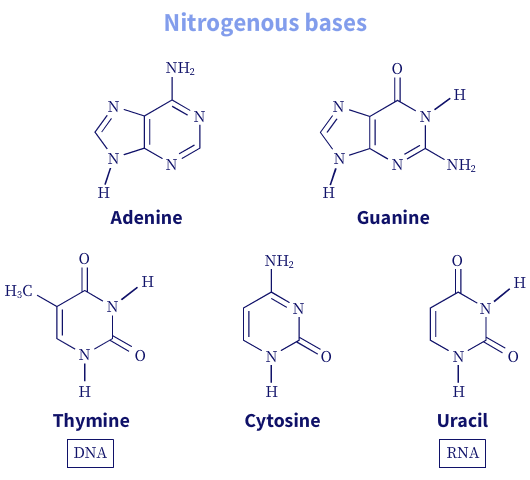
The carbon and nitrogen atoms of the nitrogenous bases are conventionally numbered (1-n). In contrast, the carbon atoms of the pentose sugar rings are given a prime (1′-5′) designation to distinguish them from the nitrogenous base atoms.
Pentoses are simple sugar molecules made of 5 carbon atoms. Nucleic acids are made of two kinds of pentoses: 2′-deoxy-D-ribose or D-ribose. The type of pentose defines the nucleic acid. For example, DNA is made of 2′-deoxy-D-ribose, while RNA is made of D-ribose molecules. In deoxyribonucleotides, the -OH group on the 2′ carbon is replaced with H.
The successive nucleotides in DNA and RNA are linked by phosphodiester linkages between the 5′-phosphate group of one nucleotide unit to the 3′-hydroxyl group of the next nucleotide. In addition, nucleotides have other functions in the cell, such as being the energy carriers, cofactors of enzymes, and chemical messengers.
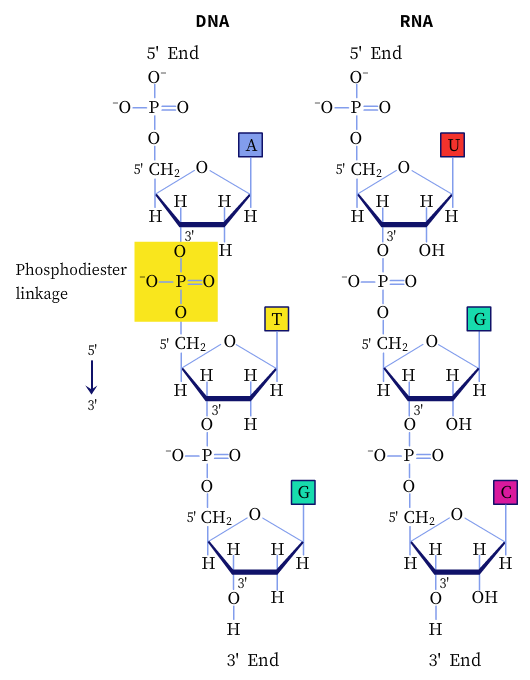
Structure of Nucleic Acids
Deoxyribonucleic acid
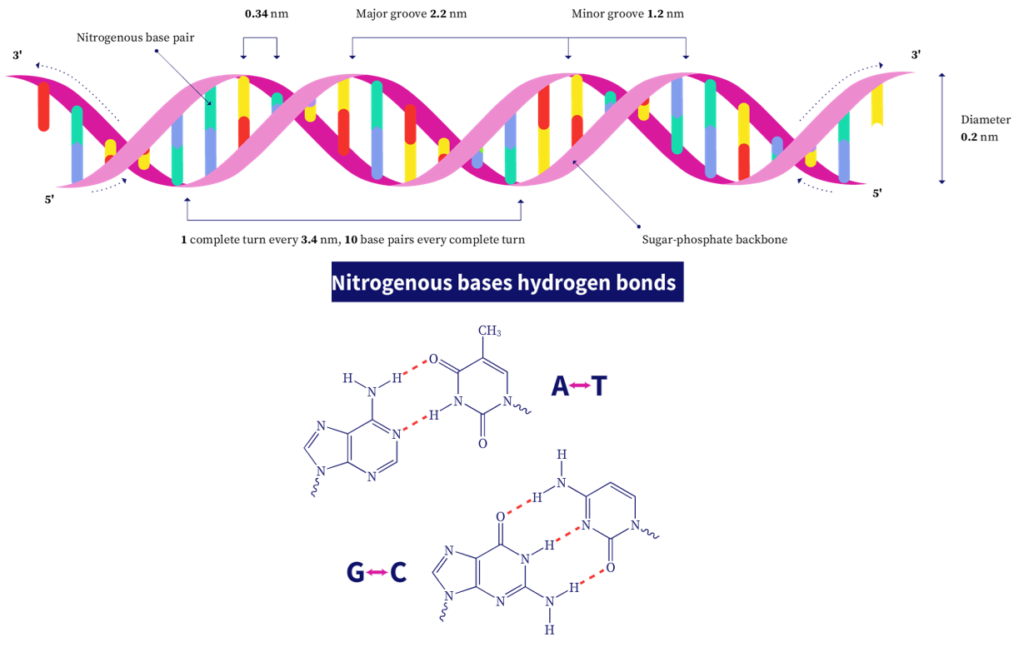
The 3D structure of DNA consists of two complementary or antiparallel strands of DNA wound around the same axis to form a right-handed double helix. The hydrophilic backbone consists of alternating deoxyribose and phosphate groups outside the double helix, facing the surrounding water. The nitrogenous bases are stacked on the inside of the double helix with their hydrophobic planar ring structures perpendicular to the long axis. There are about 10.5 bases per helical turn. Every nucleotide base of one strand is paired in the same plane with a base of the other strand as per Chargaff's rule. This base-pairing produces major and minor grooves throughout the structure of the duplex. These groves allow binding proteins to the DNA molecule during replication and transcription (described later).
The adenine forms two hydrogen bonds with thymine and is denoted by A=T. Guanine residues form three hydrogen bonds with cytosine residues, denoted by G≡C. This is why GC-rich DNA strands require a higher temperature to separate than AT-rich DNA.
The two helices of the DNA are held together by two forces:
- Hydrogen bonding between complementary base pairs.
- Van der Waals forces or weak non-specific interactions between molecules or atoms.
This DNA double helix model is the most stable form of DNA and is also known as the B-form. DNA is, however, a highly flexible molecule and many other conformations are possible. Two well-characterized forms include the A-form and the Z-form. The A-form is a right-handed helix, favored in non-aqueous solutions or short DNA strands, and has about 11 bases per helical turn. The Z-form is a left-handed helix and is slenderer and more elongated than the B-form, with 12 basepairs between each turn. In addition, the backbone has a more zig-zag appearance.
In addition to these structures, DNA can also form bends (in stretches of adenine residues), hairpins (on complementary regions within a strand), or cruciform (cross-shaped structures). Besides, three to four DNA strands can also interact to form additional structures such as a triplex DNA (for example, certain atoms of purines called Hoogsteen positions that participate in hydrogen bonding with three DNA strands, through non-Watson-Crick pairing called Hoogsteen pairing) or tetraplex DNA structures (for example, in guanosine rich DNA strands).
Some of these unique structures allow binding proteins and help regulate various cellular processes, including DNA replication.
Ribonucleic acid
The second most common form of nucleic acids is RNA. Double-stranded RNA molecules have a similar base pairing to that of DNA molecules except for adenine, which pairs with uracil instead of thymine (A=U), and guanine pairs with cytosine (G≡C). RNA molecules play a central role in protein synthesis and gene expression. The genetic information in the DNA is copied by a process called transcription into an RNA molecule called the "messenger RNA," or the mRNA. An mRNA can code for one (monocistronic) or more polypeptides (polycistronic). Eukaryotes typically have monocistronic mRNA, while bacteria and archaea produce both mono and polycistronic mRNA. The length of the mRNA is dependent on the length of the polypeptide chain.
The central dogma of molecular biology states that genetic information is encoded in the DNA can be perpetuated by replication or transferred to an RNA and subsequently used to make a DNA→RNA→Protein. Certain viruses have RNA as the genetic information. In these cases, the RNA molecule is reverse transcribed to the DNA before transcription. However, once information is transferred to make a protein, it is irreversible.
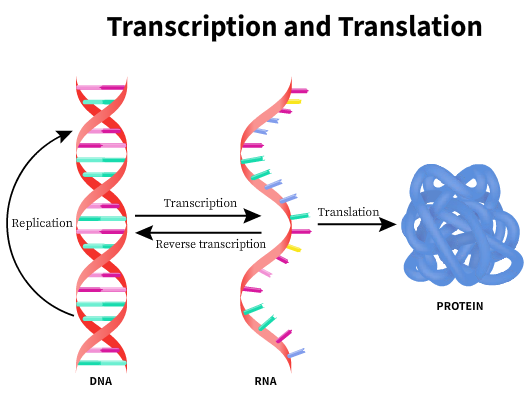
The DNA is usually located within the nucleus of eukaryotic cells or nucleoid of prokaryotic cells. Protein transcription occurs in the ribosomes located outside the nuclear region in the cytoplasm. The mRNA carries the information needed to synthesize the proteins to the site of protein synthesis. The genetic code is a codon, a triplet of nucleotides that codes for a specific amino acid. Multiple genetic codes can code for the same amino acid.
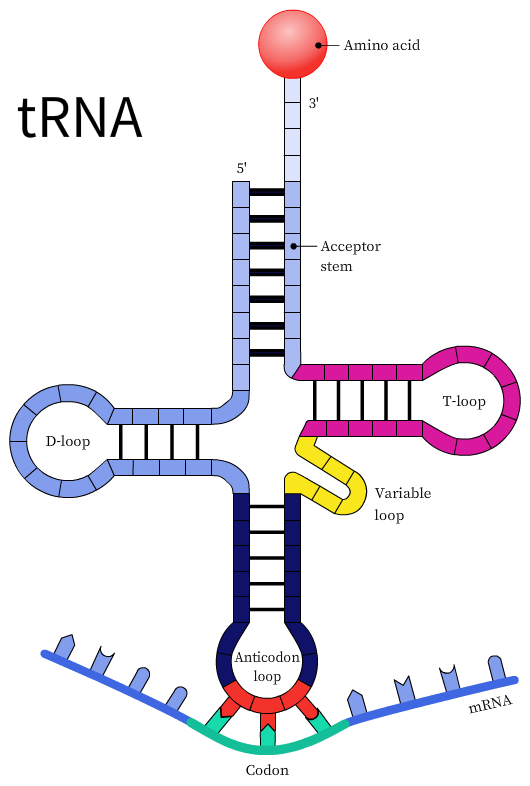
The transfer RNA or the tRNA is another RNA macromolecule that acts as an adapter for protein synthesis. The tRNA contains a three-base sequence known as the anticodon sequence that can recognize and bind to its complementary codon. During protein synthesis, the tRNAs base pair with the mRNA codons using the anticodon sequence. However, the codon and anticodon base-pairing is loose or wobbly and is not precise, especially at the third nucleotide on the codon or the first nucleotide of the anticodon. This model is known as the wobble hypothesis. The wobble pairing permits the rapid dissociation of the tRNA from its codon during protein synthesis.
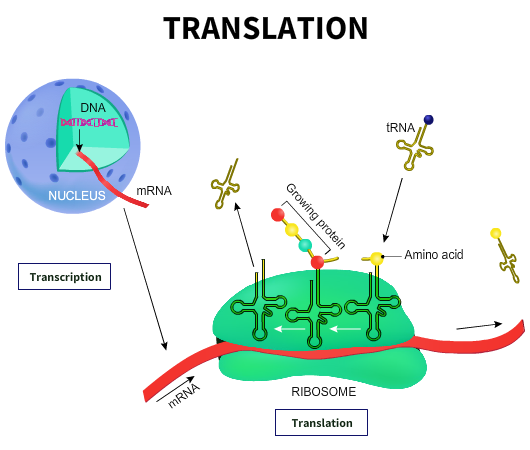
The ribosomes are RNA molecules involved in protein synthesis. There are typically thousands of ribosomes in a cell. The ribosomes in prokaryotes are smaller than eukaryotes. For example, bacterial ribosomes exist as the 70S (Svedberg) particles made up of 50S and 30S subunits. Eukaryotic ribosomes, on the other hand, are 80S particles made of 60S and 40S subunits. The ribosomes contain specific sites for the mRNA, various enzyme complexes, proteins, cofactors, and aminoacyl-transfer RNA to bind. The process of conversion of information from the mRNA to produce proteins or polypeptides is known as translation.
Context and Applications
This topic is significant in the professional exams for both undergraduate and graduate courses, especially for
- Bachelors in Zoology
- Bachelor of Science in Genetics
- Bachelor of Science in Molecular Biology
- Masters in Biochemistry
- Masters in Evolutionary Biology
Want more help with your biology homework?
*Response times may vary by subject and question complexity. Median response time is 34 minutes for paid subscribers and may be longer for promotional offers.
Nucleic Acids Homework Questions from Fellow Students
Browse our recently answered Nucleic Acids homework questions.