
Concept explainers
If a restriction enzyme cuts between the G and the A whenever it encounters the sequence GAATTC, how many fragments will be produced when the enzyme is digested with DNA with the following sequence? TGAGAATTCAACTGAATTCAAATTCGAATTCTTAGC
- a. Two
- b. Three
- c. Four
- d. Five
Introduction:
Restriction enzymes are the enzymes that digest the higher molecular weight DNA in to smaller fragments. They are also known as restriction endonucleases.
Answer to Problem 1MCQ
Correct answer:
There are three sequences of GAATTC present on the DNA fragment and the fragments formed after the cleavage of the DNA are four in numbers. Therefore, option (c) is correct.
Explanation of Solution
Reason for correct statement:
Restriction endonucleases digest a double stranded DNA at specific sites. These specific sites are also called recognition site that present as palindrome. If the restriction enzyme given cuts between the G and A whenever it encounters the sequence GAATTC on the given DNA with the following sequence TGAGAATTCTGAATTCAAATTCGAATTCTTAGC, the cleaving of the DNA will be given as follows:
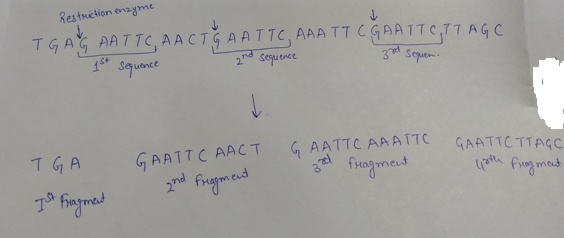
As shown, after the cleavage of the DNA by the help of the restriction enzyme, four fragments will be formed.
Option (c) is given as “Four”.
As, “the restriction enzyme recognizes three sequence on the given DNA, so it will make three cuts, that results in the production of the four DNA fragments,” is the right answer.
Hence, option (c) is correct.
Reasons for the incorrect statements:
Option (a), is given as “Two”.
The fragments of the DNA formed will be four after the action of the restriction enzyme. Hence, it is a wrong answer.
Option (b), is given as “Three”.
The DNA fragment formed after the cleavage will be four in numbers. Hence, it is a wrong answer.
Option (d), is given as “Five”.
For DNA fragments will be formed after the cutting action of the restriction enzyme at the given recognition site. Hence, it is a wrong answer.
Hence, options (a), (b), and (d), are incorrect.
Restriction enzymes are also known as molecular scissors that could cut double stranded DNA molecules at specific sites. It is an important tool that is used in the manipulation of DNA.
Want to see more full solutions like this?
Chapter 11 Solutions
BIOLOGY:CONCEPTS &...(LL) W/ACCESS +LM
Additional Science Textbook Solutions
Essentials of Genetics (9th Edition) - Standalone book
Microbiology: An Introduction
Essentials of Human Anatomy & Physiology (12th Edition)
- Choose the one answer that fits best. Which statement regarding Molecular Biology is NOT correct (videos)? a. Taq Polymerase was isolated from an organism found in Yellowstone Park b. Restriction enzymes leave sticky ends c. DNA sequencing allows us to read DNA sequences d. Restriction enzymes cut DNA at specific sites e. EcoRI and HindII are commonly used polymerasesarrow_forwardSelect the sequences that would be recognized by a restriction enzyme? (There can be more than one answer) A) 3'-TAGCTA-5' B) 5'CGATTC-3' C) 5'GAATTC- 3' D) 5'GAGCTC-3'arrow_forwardseparates molecules by movement due to size and electrical charge [ Choose ] a. DNA ligase b. gene cloning c. gel electrophoresis d. restriction enzymesarrow_forward
- Choose the combination of answers that most accurately completes the statement. Which gene is incorporated into plasmids to detect recombinant cells? a. restriction endonuclease c. a gene for antibiotic resistance b. virus receptors d. reverse transcriptasearrow_forwardExplain how electrophoresis separates DNA strands. a. How is a DNA fingerprinting test interpreted? b. Define plasmid and how plasmids can change a bacteria’s activity. c. How do we digest/cleave plasmids? Explain the role of a restriction enzyme. d. Define sticky end and blunt end and which one is useful in molecular biology.arrow_forwardFranklin and Wilkins made diffraction patterns of DNA using ___. A. X-ray crystallography B. microscopes C. electron microscopy D. X-ray microscopyarrow_forward
- Which type of cells were used to extract the DNA that was sequenced? a. red blood cells c. white blood cells b. intestinal epithelium d. cheek swab What type of mutation caused Nicholas’s disease? a. frameshift c. nonsense b. missense d. insertionarrow_forwardWhat is the purpose of the low temperature step in the PCR reaction? a. To allow DNA polymerase to synthesize new DNA in the 3' to 5' direction b. To permanently deactivate DNA polymerase c. To allow primers to anneal to DNA templates d. To allow DNA polymerase to synthesize new DNA in the 5' to 3' directionarrow_forwardFrom where do we get primers for sequencing DNA? A) they are synthesized by reverse transcriptase B) they are cut out of plasmids using restriction endonucleases C) DNA primase is added to the sequencing reaction and synthesizes the primers D) biotechnology companies synthesize them using organic chemistryarrow_forward
- Using the data in Table, identify restriction enzymes that (a) produce blunt ends; (b) recognize and cleave the same sequence (called isoschizomers); (c) produce identical sticky ends.arrow_forwardWhy must a genetically engineered plasmid contain a genetic marker? a.) to prevent the construction of an artificial chromosome b.) to separate cells that contain recombinant DNA from those that do not c.) to produce multiple copies of the recombined plasmid after heat treatment d.) to break apart the circular plasmid and introduce another DNA fragmentarrow_forwardIf a restriction enzyme cuts between the G and the A whenever itencounters the sequence GAATTC, how many fragments will beproduced when the enzyme is digested with DNA with the followingsequence? TGAGAATTCAACTGAATTCAAATTCGAATTCTTAGCa. Two c. Fourb. Three d. Fivearrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College

