
Concept explainers
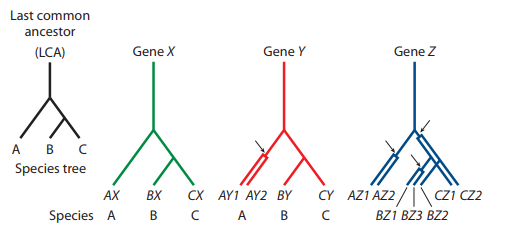
Consider the phylogenetic tree below with three related species (A, B, C) that share a common ancestor (last common ancestor, or LCA). The lineage leading to species A diverges before the divergence of species B and C.
For gene X, no gene duplications have occurred in any lineage, and each gene X is derived from the ancestral gene X via
For gene Y, a gene duplication occurred in the lineage leading to A after it diverged from that leading to B and C. Are genes
For gene Z, gene duplications have occurred in all species. Define orthology and paralogy relationships for the different Z genes.
Learn your wayIncludes step-by-step video

Chapter 16 Solutions
GENETIC ANALYSIS: INTEGRATED - ACCESS
Additional Science Textbook Solutions
Campbell Essential Biology with Physiology (5th Edition)
Campbell Biology (10th Edition)
Campbell Essential Biology (7th Edition)
Biology: Life on Earth with Physiology (11th Edition)
Campbell Biology: Concepts & Connections (8th Edition)
- In sexually reproducing species, each individual begins life with DNA inherited from both parent organisms. , Apply this idea to what occurs when organisms of two species that have homologous chromosomes mate and produce ( F1 ) hybrid offspring. What percentage of the DNA in the F1 hybrids' chromosomes comes from each parent species? As the hybrids mate and produce F2 and later-generation hybrid offspring, describe how recombination and natural selection may affect whether the DNA in hybrid chromosomes is derived from one parent species or the other.arrow_forwardYou want to make a phylogenetic tree of a group of three related species of lizards that live on an island. Their genome sequences are highly similar except for a gene that controls body size. In that region of the genome, one of the lizard species has one copy of the growth control gene (L1), the second species has a duplication of the growth control gene (L2) and the third species has three copies of the same gene (L3). The lizard species show an increase in size depending on how many copies of the growth control gene they have (L1 is smallest, L2 is medium-sized and L3 is largest). Is this enough information to determine the phylogenetic relationships between the species, and predict which of the species arrived on the island first (and is the ancestral species)? Yes, because the ancestral lizard genome probably had a single copy of the growth control gene and after arriving on the island it was duplicated, resulting in species L2, and then another duplication occurred resulting in…arrow_forwardWhich of the following best explains the number of similarities between the amino acid sequences of the Drosophila Hedgehog protein and the Chicken Indian Hedgehog protein? O A. The Drosophila hedgehog gene evolved from hedgehogs, which are distantly related to birds. O B. Both genes evolved from a gene present in the last common ancestor of Drosphila and chickens, and the number of differences reflects the amount of time that has elapsed during the evolution of these two lineages. a During the evolution of Drosophila and chickens, a hedgehog like gene arose independently in each lineage, then the gene that arose in chickens diversified. A These genes are unrelated, and the fact that they are similar is only because the proteins need to have similar biochemical properties. They are unrelated because chickens don't have segments and Drosophila larvae don't have limb buds.arrow_forward
- Suppose species 1, 2, and 3 are endemic to a group of islands (such as the Galápagos) and are all descended from species 4 on the mainland (which will serve as an outgroup; its large population size means that no new mutations have become fixed in its population in the time since the islands were colonized). You sequence a gene and find ten nucleotide sites that differ among the four species (among many other loci that do not vary). The nucleotide bases at these sites are: Species 1: GCTGATGAGT Species 2: ATCAATGAGT Species 3: GTTGCAACGT Species 4: GTCAATGACA Estimate the phylogeny of these taxa by plotting the changes on each of the three possible unrooted trees and determining which tree requires the fewest evolutionary changes.arrow_forwardVestigial wings Trait Wild-type Yellow Body White-eyes Eyeless Body Colour Gray Yellow Eye Colour Red Eye white Eyes absent Eyes absent Eye shape Normal Normal, Extends past tip of Small, Club- Wing shape and size shaped abdomen Antenna shape and size Normal Tan stripes at the end White end with Bristle shape and size stipes very unclear. Discuss the changes in chromosomes that contribute to the mutations tabulated in Table abovearrow_forwardIn yeasts and molds, both parents contribute to the cytoplasm of the offspring. Their slow growth can be due to a nuclear DNA mutation or a mitochondrial DNA mutation. A recessive nuclear mutation (nn) gives rise to a segregational petite. If most of the mtDNA is missing, a neutral petite (Mn) is formed. If some of the mtDNA is missing, a suppressive petite (Ms) is formed. A complete mtDNA (Mc) allows normal growth to occur, even if the other mtDNAs have missing segments. Mc > Ms > Mn Determine the genotype and phenotype of the F1 in following crosses: a. maternal parent is a heterozygous suppressive petite and the paternal parent is a segregational petite with normal mitochondria. b. maternal parent is heterozygous suppressive petite and the paternal parent is a heterozygous neutral petite.arrow_forward
- During the resolution of polyploidy, the subgenomes must function as a single unit. This often results in at least partial dominance of one subgenome over the other. Sequencing of polyploid genomes has demonstrated that TE density is often negatively associated with dominance. If you were presented with data suggesting that a subgenome with higher TE density associated with gene space was dominant, how could you explain the results? Describe al follow up experiment.arrow_forwardD) Two dragonfly populations that have accumulated several differences in their DNA sequences How do you know that is the answer? I thought we were not allowed to use DNA sequences in the lineage species concept.arrow_forwardWhich of the following best explains why coalescent-based phylogenetic inference is important in the age of phylogenomics? A) Coalescent-based methods directly model gene tree histories independently to infer the species tree in a summary-based manner, which is important for phylogenomic analysis were hundreds to thousands of gene histories are analyzed. B) Coalescent-based methods have the most advanced evolutionary models of molecular evolution, which is important for phylogenomic analysis were hundreds to thousands of gene histories are analyzed. C) Coalescent-based methods are no more important than other types of phylogenetic inference, even for phylogenomic analyses. D) None of the above.arrow_forward
- Suppose species 1, 2, and 3 are endemic to a group of islands (such as the Galápagos) and are all descended from species 4, an outgroup. We sequence a gene and find ten nucleotide sites that differ among the four species (among many other loci that do not vary). The nucleotide bases at these sites are Species 1: GCTGATGAGT Species 2: ATCAATGAGT Species 3: GTTGCAACGT Species 4: GTCAATGACA Estimate the phylogeny of these taxa by plotting the changes on each of the three possible trees and determine which tree requires the fewest evolutionary changes. (Please answer including what are 3 possible trees.? )arrow_forwardConsider figure 22.5b. The following statements are true. Choose all applicable options. a) Hair in this case is a developmental homology. b)The hair shown in both species can be considered a homology. c) The common ancestor of both species likely had hair. d) Both species have hair likely because they share a common ancestor.arrow_forwardImagine that you have the DNA sequences from the intron of a gene in three species called A, B, and C. Species A and B are most closely related, while C is more distantly related. The sequences of A and B differ by 18 base pairs, A and C differ by 26 base pairs, and B and C differ by 28 base pairs. Fossils show that species A and B diverged about 1.2 Mya, but there is no fossil evidence as to when the most recent common ancestor of all three species lived. (Draw a simple tree to help you think about the problem) Use the genetic data to estimate that date (most recent common ancestor). HINT = use Eqn 7.1, several times- first to estimate mutation rate. Then to estimate the unknown time since divergencearrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





