
Concept explainers
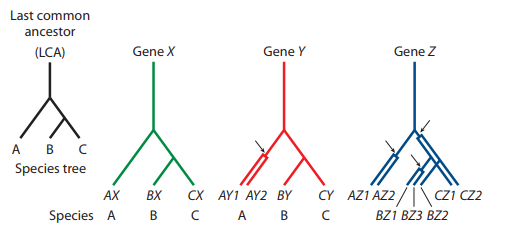
Consider the phylogenetic tree below with three related species (A, B, C) that share a common ancestor (last common ancestor, or LCA). The lineage leading to species A diverges before the divergence of species B and C.
For gene X, no gene duplications have occurred in any lineage, and each gene X is derived from the ancestral gene X via
For gene Y, a gene duplication occurred in the lineage leading to A after it diverged from that leading to B and C. Are genes
For gene Z, gene duplications have occurred in all species. Define orthology and paralogy relationships for the different Z genes.
Learn your wayIncludes step-by-step video

Chapter 18 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
Additional Science Textbook Solutions
Campbell Essential Biology with Physiology (5th Edition)
Campbell Biology (10th Edition)
Campbell Essential Biology (7th Edition)
Biology: Life on Earth with Physiology (11th Edition)
Campbell Biology: Concepts & Connections (8th Edition)
- Referring to the phylogenetic tree shown above, answer the following questions: 1. How many OTUs are included in the phylogenetic analysis? 2. How many clades are there? 3. What is an autapomorphic trait of the domestic cat? Explain why? 4. What is the shared derived trait (synapomorphy) in the Family Felidae? Explain why?arrow_forwardIf mutations such as those of the Ubx gene can drastically change morphology in a single step, why do most evolutionary biologists maintain that modification of existingtraits and the evolution of novel characters have generally proceeded by successive small steps?arrow_forwardYou want to make a phylogenetic tree of a group of three related species of lizards that live on an island. Their genome sequences are highly similar except for a gene that controls body size. In that region of the genome, one of the lizard species has one copy of the growth control gene (L1), the second species has a duplication of the growth control gene (L2) and the third species has three copies of the same gene (L3). The lizard species show an increase in size depending on how many copies of the growth control gene they have (L1 is smallest, L2 is medium-sized and L3 is largest). Is this enough information to determine the phylogenetic relationships between the species, and predict which of the species arrived on the island first (and is the ancestral species)? Yes, because the ancestral lizard genome probably had a single copy of the growth control gene and after arriving on the island it was duplicated, resulting in species L2, and then another duplication occurred resulting in…arrow_forward
- A cladogram used in a comparison of morphology among taxa had equal length branches, but when looked at in a blast webpage using the given gene sequence, the branches all had different lengths. Why is that?arrow_forwardA researcher studying two species (species 1 and species 2) sequences a short stretch of eight codons from the same gene, gene B, in each and compares them. Species 1 and species 2 had a most recent common ancestor 50 million years ago. Species 1: ATC GGG CGG GAC TTA CTA TAT GCC Species 2: ATT GGG CGG GAC TTG CTA TAT GCC Given the differences between the sequences of the two species' genes shown here, what evolutionary force can you predict is most likely in operation on gene B?arrow_forwardSuppose we are sure, because of previous studies, that species 1, 2, and 3 are more closely related to each other than to species 4 (outgroup). We sequence a gene and find ten nucleotide sites that differ among the four species. Draw the most parsimonious tree and label each evolutionary change on the tree (Position – new nucleotide; Example = 8T or 6C). *The answer is below but I do not understand where the numbers or tick marks came from? Could someone explain. For example, why is the 1A on the 2?arrow_forward
- Phylogenetic trees are a type of model that can be used to show how organisms are related through common ancestry. The phylogenetic tree model represents nodes numbered 1 through 8. Using evidence from the phylogenetic tree determine which species would be MOST closely related to the species on branch C? Question options: The species on Branch A is most closely related to the species on branch C because they share the most recent common ancestor at node 1. The species on Branch B is most closely related to the species on branch C because they share the greatest number of common +ancestors. The species on Branch A & B are both most closely related to the species on branch C because they share the most most recent common ancestor at node 2. The species on Branches F, G, H, and I are all equally related to the species on branch C because they all split from a common ancestor at the same time which is illustrated by having nodes 2 and 7 at the…arrow_forwardWould a protein encoded on the core genome or one encoded only on the pan-genome be best to use in constructing a phylogenetic tree? Explain your answerarrow_forwardFor the first phylogenetic tree, if we assume absolute time is NOT represented, can we say that the species in circle B are more closely related than the species in Circle A? For the second phylogenetic tree (if we hold the same assumptions), can we say that B and C are more closely related than A and C?arrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
