
Concept explainers
Gene Transfer Between Bacteria
Core Skill: Connections Look back at Figures 11.1 and 11.2. How did the phenomenon of transformation allow researchers to demonstrate that DNA is the genetic material?
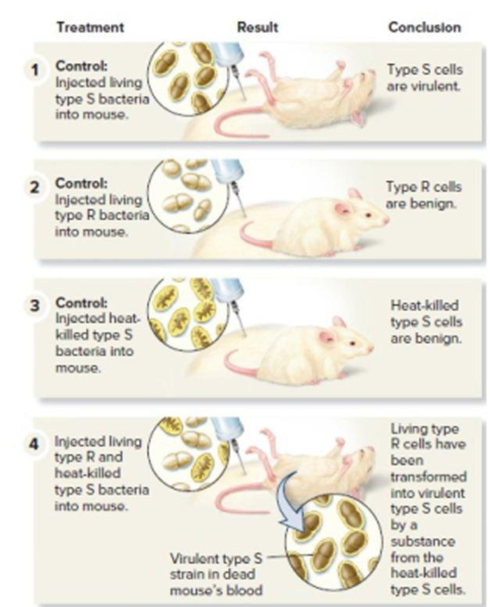
Figure 11.1 Griffith’s experiments showing that genetic material can be transferred from one bacterium to another. Note: To determine if a mouse’s blood contained live bacteria, a sample of blood was also applied to solid growth media. (This part of the procedure is not shown.) For steps 1 and 4, smooth bacterial colonies were observed. For step 2, no bacterial colonies were observed because the type R cells were killed by the immune system of the mouse.
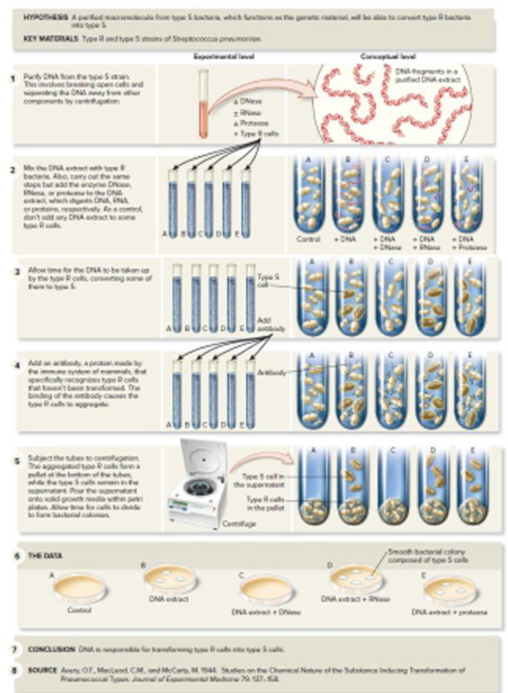
Figure 11.2 The Avery, MacLeod, and McCarty experiments that identified DNA as Griffith’s transformation principle—the genetic material.
Want to see the full answer?
Check out a sample textbook solution
Chapter 18 Solutions
Gen Combo Ll Biology; Connect W/learnsmart Labs Access Card
- AGAROSE GEL ELECTROPHORESIS PRELAB ANSWER ALL QUESTIONS 1. Type of nucleotide is one of the factors that influence electrophoretic mobility. a) True b) False 2. Electrophoresis is used for DNA separation and not proteins. a) True b) False 3. DNA Polymerase is an enzyme to cut DNA into fragments for electrophoresis. a)True b)False 4. Electrophoresis apparatus consists of Gel buffer, chamber and DNA separator. a) True b) False 5. During electrophoresis, DNA fragments collect at the Cathode. a) True b) Falsearrow_forwardExpand PCR? Describe the different Steps involved in this technique?arrow_forwardQUESTION: what difficulty will you experience if you do genetic manipulation to streptomyces spp. and how can this difficulty overcome ?? how could you modulate the gene expression for improving the productivity of an antibiotic produced by the streptomyces strain ? discuss with diagramarrow_forward
- Objective: Get a sense of how genomics, the study of the genome in its entirety,needs to think about how to go about its research. Geonomic DNA is broken up into fragments. The 5’ and 3’ ends of each fragment(a “read”) are sequenced. The sequenced reads are assembled together intocontiguous sequences (“contigs”) based on sequence similarity. The idea is to sequence enough random fragments so that every nucleotide in thegenome is represented on some read. The number of such fragments needed iscalled the coverage, c. The coverage c can be calculated by the formula RL/G, where R is the number ofreads sequenced, L is the average length of a read and G is the total length of thegenome. The units of length are bases (b) or base pairs (bp). Consider a genome whose length is 1000 bp. “Shotgun” sequencing techniquesare applied to the genome, resulting in 20 reads, with an average length of 50 bp.A very important point is that, even though 20 x 50 = 1000, there is no guaranteethat ALL…arrow_forwardObjective: Get a sense of how genomics, the study of the genome in its entirety,needs to think about how to go about its research. Geonomic DNA is broken up into fragments. The 5’ and 3’ ends of each fragment(a “read”) are sequenced. The sequenced reads are assembled together intocontiguous sequences (“contigs”) based on sequence similarity. The idea is to sequence enough random fragments so that every nucleotide in thegenome is represented on some read. The number of such fragments needed iscalled the coverage, c. The coverage c can be calculated by the formula RL/G, where R is the number ofreads sequenced, L is the average length of a read and G is the total length of thegenome. The units of length are bases (b) or base pairs (bp). Consider a genome whose length is 1000 bp. “Shotgun” sequencing techniquesare applied to the genome, resulting in 20 reads, with an average length of 50 bp.A very important point is that, even though 20 x 50 = 1000, there is no guaranteethat ALL…arrow_forwardAnswer: Task 2: Polymerase Chain Reaction After purifying the gDNA extract, you want to isolate and amplify the polystyrenase gene. You perform PCR using the appropriate gene-targeted primers. What is the purpose of the following PCR components? Answer briefly but completely. DNA polymerase isolated from Thermus aquaticus Answer: a. b. Deoxynucleotide triphosphates (dNTPs) Answer: Forward and reverse primers Answer: с. Task 3: Agarose Gel Electrophoresis Your amplicon from PCR was subjected to AGE for analysis. You are shown the image of the gel loaded with the following lanes: (A) negative control, (B) size ladder (1, 2, 3, 4, 6, and 10 kb), (C) GDNA extract, (D) PCR amplicon. However, due to mishaps while loading the gel with the samples, you are not sure which lane is which. You are shown a diagram of the obtained gel below. a. Label each lane of the gel. Write only the corresponding letters in the wells above. b. Above each band in the size ladder, write its size (in kb). c.…arrow_forward
- Lab 9 PCR Homework Introduction: Define the following concepts and processes 1) Compare and contrast the two DNA fingerprinting technologies learned: RFLP vs PCR. What are different and common between two technologies? When would you use PRC but not RFLP?arrow_forwardExperiment: Bradford protein assaygive the answers (4-5 lines) of review questions in the end. the answer should be logical and understandable and without plagiarism. avoid copy-pasting. PLEASE GIVE THE ANSWER OF 3rd AND 4th QUESTION. ITS COMPULSORY.Background information:The Bradford assay is very fast and uses about the same amount of protein as the Lowry assay. It is fairly accurate and samples that are out of range can be retested within minutes. The Bradford is recommended for general use, especially for determining protein content of cell fractions and assessing protein concentrations for gel electrophoresis. The method described below is for a 100 µl sample volume using 5 ml color reagent. It is sensitive to about 5 to 200 micrograms protein, depending on the dye quality. In assays using 5 ml color reagent prepared in lab, the sensitive range is closer to 5 to 100 µg protein. Scale down the volume for the "micro assay procedure," which uses 1 ml cuvettes. Protocols, including…arrow_forwardVISUAL SKILLS Consider the microarray in Figure 20.12.If a sample from normal tissue is labeled with a green fluorescent dye and a sample from cancerous tissue is labeledred, what color spots would represent genes you would beinterested in if you were studying cancer? Explain.arrow_forward
- VISUAL SKILLS Griffith was trying to develop a vaccinefor S. pneumonia when he was surprised to discover thephenomenon of bacterial transformation. Look at the second and third panels of Figure 16.2. Based on these results,what result was he expecting in the fourth panel? Explain.arrow_forwardRemember that although there are many interesting ideas about genetic engineering of plants and animals, this is specifically about GE bacteria. Please be sure you are answering the following questions 3) What are the benefits (or potential benefits) of the engineered bacterium? 4) What are the risks (or potential risks) of the engineered bacterium?arrow_forwardCompare the possible differences between a eukaryotic protein-encoding gene cloned by PCR and the same gene cloned by reverse transcriptase PCR (RTPCR).arrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
