
Two inbred lines of sunflowers
a. Use the information above to determine
b. Determine
Trending nowThis is a popular solution!

Chapter 19 Solutions
GENETIC ANALYSIS: INTEGRATED - ACCESS
- In a wild strain of tomato plants, the phenotypic variance fortomato weight is 3.2 g2. In another strain of highly inbred tomatoesraised under the same environmental conditions, the phenotypicvariance is 2.2 g2. With regard to the wild strain,A. Estimate VG.B. What is hB2?C. Assuming that all of the genetic variance is additive, what is hN2?arrow_forwardThe mean, standard deviation, variances, and coefficient of variance of plant height from two rice plants (P1 and P2) and their progeny (F1 and F2) and a backcross generation (P1 x F1) are shown below. Explain the possible reasons for the observed differences in the sample means. Account for the differences in the sample means of P1 and P2. Similarly, account for the differences in the sample means of the F1 and F2. Compare the difference in the parental generations with that in the filial generations.arrow_forwardThe mean and standard deviation of plant height from two rice plants (P1 and P2) and their progeny (F1 and F2) and a backcross generation (P1 x F1) are shown below. Interpret the CV results from each population.arrow_forward
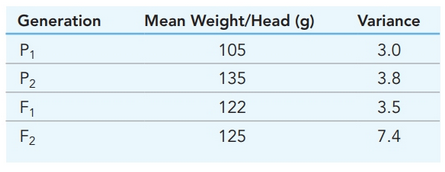
- The mean and standard deviation of plant height from two rice plants (P1 and P2) and their progeny (F1 and F2) and a backcross generation (P1 x F1) are shown below. Complete the table by calculating the variances and coefficient of variation for each population and answer the questionswhich follow (See image) 1. Explain the possible reasons for the observed differences in the sample means. Account forthe differences in the sample means of P1 and P2. Similarly, account for the differences in thesample means of the F1 and F2. Compare the difference in the parental generations with thatin the filial generations.2. Interpret the CV values from each population.3. Compare the sample variances of P1 and P2. Account for any differences. Similarly, comparethe sample variances of the F1 and F2 generations, and account for any differences. Give thepossible causes of variation in each generation.4. Calculate the broad-sense heritability of plant height in this species. Interpret your results.arrow_forwardWhich of the following applies to the Hardy-Weinberg expression:p2 + 2pq + q2?a. Knowing either p2 or q2, you can calculate all the otherfrequencies.b. It applies to Mendelian traits that are controlled by one pairof alleles.c. 2pq = heterozygous individualsd. It can be used to determine the genotype and allelefrequencies of the previous and the next generations.e. All of these are correct.arrow_forwardTwo inbred lines of sunflowers (P1 and P2) produce different total weights of seeds per flower head…. Show more Two inbred lines of sunflowers (P1 and P2) produce different total weights of seeds per flower head. The mean weight (in grams) and the variance of seed weights in different generations are listed in the table below. 1. Use the information to determine VG, VE and VP for this trait. 2. Determine H 2 for this trait. Generation Mean Weight/Head (g) Variance P1 105 3.0 P2 135 3.8 F1 122 3.5 F2 125 7.4 •arrow_forward
- Imagine that you are performing a cross involving seed texture in garden pea plants. You cross true-breeding round and wrinkled parents to obtain F1 offspring. Which of the following experimental results in terms of numbers of plants are closest to what you expect in the F2 progeny? a. 8lOroundseeds b. 8lOwrinkledseeds c. 405:395 round seeds:wrinkled seeds d. 610:190 round seeds:wrinkled seedsarrow_forwarda. 1 dominant allele will contribute 120/10 = 12 cm to the base height of the plant.b. The height of the parent plant 1 Genotype of the parent plant 1 – D1D1D2D2D3D3d4d4d5d5 The height of the parent plant 2 Genotype of the parent plant 2 – d1d1d2d2d3d3D4D4D5D5Contributing alleles – D4D4D5D5. The height of the plant without any contributing alleles would be 80 cm. The plant with genotype d1d1d2d2d3d3D4D4D5D5 has 4 contributing allele each of which contributes 12 cm to the base. Hence, the height of the plant with genotype d1d1d2d2d3d3D4D4D5D5 would be 80 + 12 + 12 + 12 + 12 = 128 cm. c. Parents – D1D1D2D2D3D3d4d4d5d5 × d1d1d2d2d3d3D4D4D5D5 Gametes – D1D2D3d4d5 × d1d2d3D4D5 F1 generation – D1d1D2d2D3d3D4d4D5d5 The height of the plants of F1 generation = 80 + 12 + 12 + 12 + 12 + 12 = 140 cm Hence, Genotype of the F1 = D1d1D2d2D3d3D4d4D5d5 Phenotype of…arrow_forwardThe mean and variance of plant height of two highly inbred strains (P1 and P2) and their progeny (F1 and F2) are shown here. Strain Mean (cm) Variance P1 34.2 4.2 P2 55.3 3.8 F1 44.2 5.6 F2 46.3 10.3 Calculate the broad-sense heritability (H2) of plant height in this species.arrow_forward
- A wide-ranging survey of Nicotonia growing in its natural environment recorded a variation in corolla length ranging from 12mm to 47mm with a variance of 36.5. Subsequently, collected seeds were grown in a greenhouse and it was found that the range was now very much lower with most plants having similar corolla lengths and the variance was now only 8.4. After the plants had grown to maturity and formed seed, seeds were collected from plants with either the shortest and or the longest corollas in the population and planted separately in the greenhouse. When flowers were formed it was found that the variance of the plants with the shortest flowers was now 4.2 while that of the flowers from the longest seeds had become 13.7 Calculate the values for heritability in the different groups of plants and explain why this difference may arise.arrow_forwardA wide-ranging survey of Nicotonia growing in its natural environment recorded a variation in corolla length ranging from 12mm to 47mm with a variance of 36.5. Subsequently, collected seeds were grown in a greenhouse and it was found that the range was now very much lower with most plants having similar corolla lengths and the variance was now only 8.4. After the plants had grown to maturity and formed seed, seeds were collected from plants with either the shortest and or the longest corollas in the population and planted separately in the greenhouse. When flowers were formed it was found that the variance of the plants with the shortest flowers was now 4.2 while that of the flowers from the longest seeds had become 13.7 So,Calculate the new values for heritability in the different groups of plants and explain why this difference may arise.arrow_forwardA wide-ranging survey of Nicotonia growing in its natural environment recorded a variation in corolla length ranging from 12mm to 47mm with a variance of 36.5. Subsequently, collected seeds were grown in a greenhouse and it was found that the range was now very much lower with most plants having similar corolla lengths and the variance was now only 8.4. After the plants had grown to maturity and formed seed, seeds were collected from plants with either the shortest and or the longest corollas in the population and planted separately in the greenhouse. When flowers were formed it was found that the variance of the plants with the shortest flowers was now 4.2 while that of the flowers from the longest seeds had become 13.7 Thus, calculate the new values for heritability in the different groups of plants and explain why this difference may arise.arrow_forward
 Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
Concepts of BiologyBiologyISBN:9781938168116Author:Samantha Fowler, Rebecca Roush, James WisePublisher:OpenStax College
