
Concept explainers
(a)
Interpretation:
The
Concept introduction:
In mass spectroscopy, compounds can be identified on the basis of the mass of the compound. When the compound breaks into fragment then they can be distinguished from the other compounds. This technique is also used to differentiate the isotopes of compounds. In amino acids, three types of fragments are observed in low energy collisions are a, b and y ions. It is known as tandem mass spectrometry.
Answer to Problem 27.27P
The
Where N is asparagine, F is phenylalanine, E is glutamic acid, S is serine, G is glycine, K is lysine amino acid.
Explanation of Solution
In amino acids, b-type fragments appear due to an amino group or in other words charge is being carried by N-terminal. That is why it is also known as the N-terminus amino acid fragment. The b-type fragment is shown below.
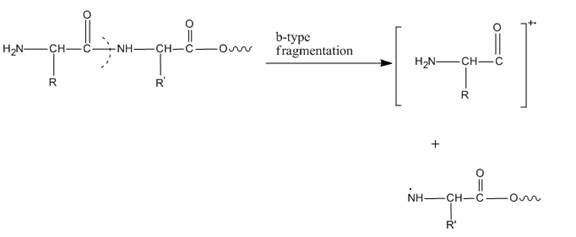
Figure 1
The given peptide is
The
(b)
Interpretation:
The
Concept introduction:
In mass spectroscopy, compounds can be identified on the basis of the mass of the compound. When the compound breaks into fragment then they can be distinguished from the other compounds. This technique is also used to differentiate the isotopes of compounds. In amino acids, three types of fragments are observed in low energy collisions are a, b and y ions. It is known as tandem mass spectrometry.
Answer to Problem 27.27P
The
Where N is asparagine, F is phenylalanine, E is glutamic acid, S is serine, G is glycine, K is lysine amino acid.
Explanation of Solution
In amino acids, y-type fragments appear due to a carboxyl group or in other words charge is being carried by C-terminal. That is why it is also known as the C-terminus amino acid fragment. The y-type fragment is shown below.
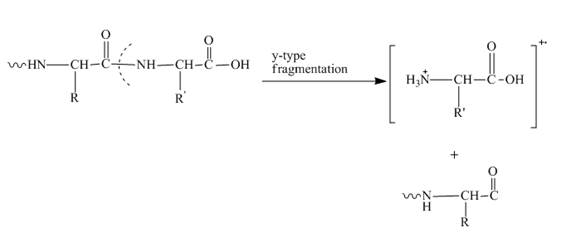
Figure 2
The given peptide is
The
Want to see more full solutions like this?
Chapter 27 Solutions
EBK ORGANIC CHEMISTRY STUDY GUIDE AND S
- 22-61 Polyglutamic acid (a polypeptide chain made only of glutamic acid residues) has an a-helix conformation below pH 6.0 and a random-coil conformation above pH 6.0. What is the reason for this conformational change?arrow_forward22-49 Based on your knowledge of the chemical properties of amino acid side chains, suggest a substitution for leucine in the primary structure of a protein that would probably not change the character of the protein very much.arrow_forwardComplete the following reaction: Naz-peptide -Peptide CA> ران CH CH3 CH3 NP₂ CH H' (hydrolysit) NA₂ V=0arrow_forward
- About a decade ago, it was more common to use peptide mass mapping to identify a protein cut from a gel. Peptide mass mapping involves digesting the protein with trypsin and then measuring the peptide masses using MALDI-TOF mass spectrometry. The more superior method of identifying a protein from a gel is to digest the protein with trypsin and then use liquid chromatography to separate the peptides. The separated peptides are then electrosprayed into a tandem mass spectrometer. The MS-MS data is uploaded into a database such as MASCOT for protein identification. Explain why the LC-MS-MS method is superior to the peptide mass mapping method.arrow_forwardThe amino acid histidine has ionizable groups with pKą values of 1.8, 6.0, and 9.2, as shown. COOH T H₂N-CH | CH₂ H -O pH UT = ZI Ionizable -COOH group CH H₂N-CH T 1.8 COO- pk₁ CH2 UI COO™ H IZ CH -HisH+ COO™ + H₂N-CH 6.0 pK₂ CH ₂ UI -His H CH -NH COO™ H₂N-CH 9.2 pk3 CH ₂ UI —NH₂ H A biochemist makes up 80 mL of a 0.11 M solution of histidine at a pH of 5.1. She then adds 60 mL of 0.10 M HCl. What is the pH of the resulting solution? CHarrow_forwardA protein consists of two types of peptide chains (A and B) with an unknown stoichiometry (AxBy). When you ran this protein directly on reversed phase-HPLC with UV monitor set at 280 nm, two peaks were resolved. Mass spec determined that the peaks represented Chain A and Chain B, respectively. The peak area is 500,000 for Peak A (Chain A) and 100,000 for Peak B (Chain B). The molecular masses of Chain A and Chain B are 25,000 and 5000, respectively. The extinction coefficients for Chain A and Chain B are 1 mL/mg.cm and 0.5 mL/mg.cm, respectively. Please calculate x/y.arrow_forward
- Give the numbers of peptide bond present in given structure, structure is given at pH 10.5, find the error in if pH is 7.4arrow_forwardMost of the ultraviolet absorption of proteins at 280 nm is due to their content of a. Glutamate b. Tryptophan c. Alanine d. Aspartatearrow_forwardFor the following peptide chain WHD (Tryptophan-Histidine-Aspartic acid) : pH = 8 Given pKa value for W : Alpha carboxy = 2.38 + Alpha Amino = 9.39 Given pKa value for H: Alpha carboxy = 1.82 + Alpha Amino = 9.17 Side chain = 6.04 Given pKa value for D : Alpha carboxy = 2.09 + Alpha Amino = 9.82 Side chain = 3.86 a. what is the charge of each amino acid residue b. what is the net charge on the whole protein at pH=8arrow_forward
- 22-35 Why is histidine considered a basic amino acid when the pKa of its side chain is 6.0?arrow_forward22-42 (a) How many atoms of the peptide bond lie in the same plane? (b) Which atoms are they?arrow_forward22-47 How many different tetrapeptides can be made (a) if the peptides contain the residues of asparagine, proline, serine, and metbionine and (b) if all 20 amino acids can be used?arrow_forward
 Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning
Introduction to General, Organic and BiochemistryChemistryISBN:9781285869759Author:Frederick A. Bettelheim, William H. Brown, Mary K. Campbell, Shawn O. Farrell, Omar TorresPublisher:Cengage Learning Chemistry: Principles and ReactionsChemistryISBN:9781305079373Author:William L. Masterton, Cecile N. HurleyPublisher:Cengage Learning
Chemistry: Principles and ReactionsChemistryISBN:9781305079373Author:William L. Masterton, Cecile N. HurleyPublisher:Cengage Learning Chemistry for Today: General, Organic, and Bioche...ChemistryISBN:9781305960060Author:Spencer L. Seager, Michael R. Slabaugh, Maren S. HansenPublisher:Cengage Learning
Chemistry for Today: General, Organic, and Bioche...ChemistryISBN:9781305960060Author:Spencer L. Seager, Michael R. Slabaugh, Maren S. HansenPublisher:Cengage Learning Principles of Instrumental AnalysisChemistryISBN:9781305577213Author:Douglas A. Skoog, F. James Holler, Stanley R. CrouchPublisher:Cengage Learning
Principles of Instrumental AnalysisChemistryISBN:9781305577213Author:Douglas A. Skoog, F. James Holler, Stanley R. CrouchPublisher:Cengage Learning



