
Concept explainers
To estimate: The date when the most recent common ancestor of all three species lived using the given genetic data.
Introduction: A common ancestor of different species is an evolutionary link between the different species. Different organisms have evolved by differentiation and divergence from the common ancestor, which is their origin. The degree of differentiation or the time that it takes for an organism to evolve from a common ancestor is measured by the genetic distance.
Explanation of Solution
Pictorial representation: Fig.1 represents the phylogenetic tree of the common ancestry.
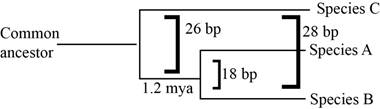
Fig.1: Phylogenetic tree
A species and B species are closely related, but species C is more distant to A and B. The phylogenetic tree shows that the common ancestor of A and B were seen 1.2 million years ago. As species C is even more distant, the common ancestor of all three was seen much before 1.2 million years ago. Since it took 1.2 million years to lead to a difference of 18 base pairs, time when the common ancestor on A, B, and C was seen, can be estimated to be 1.2 to 2 million years before the common ancestor of A and B.
To describe: The assumptions made to get the estimated date.
Introduction: A common ancestor of different species is an evolutionary link between the different species. Different organisms have evolved by differentiation and divergence from the common ancestor, which is their origin. The degree of differentiation or the time that it takes for an organism to evolve from a common ancestor is measured by the genetic distance.
Explanation of Solution
Species C is more distant to species A and B, hence their common ancestor is expected to present much earlier than the common ancestor of A and B. Species C is estimated to have lived 1.2 to 2 million years before the common ancestor of A and B.
To derive this, several assumptions are made such as follows:
- It is assumed that
speciation is a long and gradual process which occurs over a period of time. - Phylogenetic trees can be used to study diversification.
- It is assumed that the three species A, B, and C have some similarities and hence have a common ancestor, which serves as a point of origin for diversification of species.
Want to see more full solutions like this?
Chapter 7 Solutions
Evolution
- The phylogenetic tree for vertebrates depicted below was constructed from sequence data for two rRNA mitochondrial genes (12S and 16S). How do the results of this analysis compare with the phylogenetic trees in Figures 32.10 and 32.24? Identify the major clades of vertebrates on the tree depicted below. Source: R. Zardoya and A. Meyer. 1998. Complete mitochondrial genome suggests diapsid affinities of turtles. Proceedings of the National Academy of Sciences, USA 95:1422614231. Copyright 1998 National Academy of Sciences, U.S.A.arrow_forwardA researcher studying two species (species 1 and species 2) sequences a short stretch of eight codons from the same gene, gene B, in each and compares them. Species 1 and species 2 had a most recent common ancestor 50 million years ago. Species 1: ATC GGG CGG GAC TTA CTA TAT GCC Species 2: ATT GGG CGG GAC TTG CTA TAT GCC Given the differences between the sequences of the two species' genes shown here, what evolutionary force can you predict is most likely in operation on gene B?arrow_forwardImagine that you have the DNA sequences fromthe intron of a gene in three species called A, B,and C. Species A and B are most closely related,while C is more distantly related. The sequencesof A and B differ by 18 base pairs, A and C differby 26 base pairs, and B and C differ by 28 basepairs. Fossils show that species A and B divergedabout 1.2 Mya, but there is no fossil evidence asto when the most recent common ancestor ofall three species lived. Use the genetic data toestimate that date. What assumptions are youmaking to get this estimate?arrow_forward
- Why is molecular data for phylogenetic inference best analyzed with use of an explicit model of molecular evolution? A) This is true of morphological data, not molecular data, it is impossible to model changes in molecular sequence data because it is constantly evolving. B) Because molecular data is known to only experience random changes and is constantly evolving, a chaotic model of evolution can universally be applied to molecular sequence data for phylogenetic analysis. C) Because molecular data is known to experience non-random changes in terms of the likelihood of different types of mutations -- transitions vs. transversions, at different codon positions, which can be used to infer sequence evolution and relationship. D) None of the above.arrow_forwardSuppose we are sure, because of previous studies, that species 1, 2, and 3 are more closely related to each other than to species 4 (outgroup). We sequence a gene and find ten nucleotide sites that differ among the four species. Draw the most parsimonious tree and label each evolutionary change on the tree (Position – new nucleotide; Example = 8T or 6C). *The answer is below but I do not understand where the numbers or tick marks came from? Could someone explain. For example, why is the 1A on the 2?arrow_forwardWhich of the following statements best explains the importance of sister taxa for understanding the evolution of ingroup taxa? A) The sister group informs of the likely state of the common ancestor of the ingroup. B) The sister group is composed only of ancestral characters representing a primitive taxon having ancestral states for all characters able to be studied. C) The sister group allows us to infer character state polarity in order to understand how character states have changed during evolution of the ingroup. D) A & B E) A& C F) All of the above.arrow_forward
- If mutations such as those of the Ubx gene can drastically change morphology in a single step, why do most evolutionary biologists maintain that modification of existingtraits and the evolution of novel characters have generally proceeded by successive small steps?arrow_forwardA cladogram used in a comparison of morphology among taxa had equal length branches, but when looked at in a blast webpage using the given gene sequence, the branches all had different lengths. Why is that?arrow_forwardIn the past, the evolutionary history of whales was represented by cladogram A, shown below. As you can see, whales were believed to be closely related to mesonychids, an extinct group of mammals that looked similar to wolves. Today, that cladogram has been revised, as shown in cladogram B. Which of the following statements best describes the reason for this change? A - Cladograms A and B are hypotheses that changed as new evidence became available.B - Cladogram B was revised to show a water-to-land pattern of evolution in groups of organisms.C - Cladogram B was altered to better include similarities in habitat as new information became available.D - Cladograms are organized today to show a much more simplified pattern like the one shown in cladogram B.arrow_forward
- By comparing DNA sequence of a specific gene, we can determine the evolutionary relationship of two organisms. Based on the sequence listed below, which two species would you expect to be more closely related? Organism No.1 with ATG CAA TAC GCC, organism No. 2 with ATG CAT GAC ACC and organims No. 3 with ATG CAT TAC GCC A. Organims No.2 and No.3 are likely more closely related. b.Organims No.1 and No.2 are likely more closely related. c.Organims No 1 and No.3 are likely more closely related.arrow_forwardAsian tiger mosquito Trace its origin and evolutionary history or changes in the species. Describe its structures and their functions. What is the importance of this species to our environment? Give trivia about this species. Does this organism produce oxygen? Explain. What are the ancestral species of your chosen organism? Has this species been genetically engineered? If yes, in what way? How does this species reproduce? What organisms have similar structures to this species? Do these structures have the same function? Does the species have tissues, organs, and/or organ systems? What is its role in the flow of energy? Give at 15 least a sentence to each question.arrow_forwardIf you found the same gene in all organisms you test, what could this suggest about the evolution of this gene in the history of life on earth? What if a gene is very similar in most species that you study, but quite different in a particular group? What if it’s present in nearly everything, but the specific sequences vary quite a bit?arrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
