
Concept explainers
The electrophoresis gel shown in part (a) is from a DNase footprint analysis of an operon transcription control region. DNA sequence analysis of a
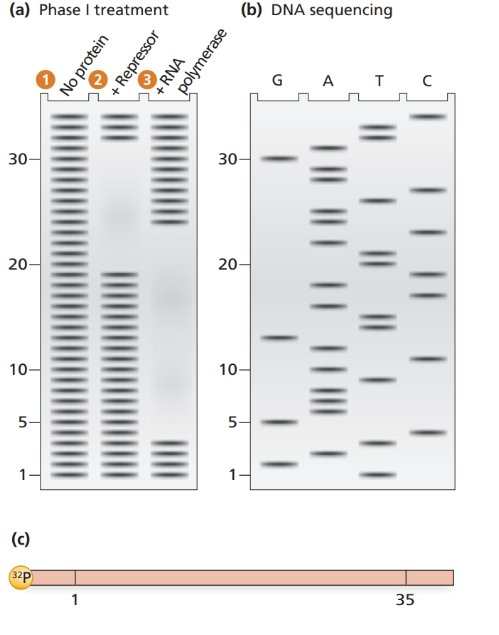
a. Determine the DNA sequence of the region examined.
b. Locate the regions of the sequence protected by repressor protein and by RNA polymerase.
Want to see the full answer?
Check out a sample textbook solution
Chapter 12 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- You then make a screen to identify potential mutants (shown as * in the diagram) that are able to constitutively activate Up Late operon in the absence of Red Bull and those that are not able to facilitate E. Coli growth even when fed Red Bull. You find that each class of mutations localize separately to two separate regions. For those mutations that prevent growth even when fed Red Bull are all clustered upstream of the core promoter around -50 bp. For those mutations that are able to constitutively activate the operon in the absence of Red Bull are all located between the coding region of sleep and wings. Further analysis of each DNA sequence shows that the sequence upstream of the promoter binds the protein wings and the region between the coding sequence of sleep and wings binds the protein sleep. When the DNA sequence of each is mutated, the ability to bind DNA is lost. Propose a final method of gene regulation of the Up Lateoperon using an updated drawn figure of the Up Late…arrow_forwardExplain why (a) inactivation of the O2 or O3 sequence of the lac operon causes only a twofold loss in repression, and (b) inactivation of both O2 and O3 reduces repression ∼70-fold.arrow_forwardIn the lac operon system, do the inducers act at the transcription or translation level? Explain why it acts in that level in a short but concise way.arrow_forward
- You have isolated different mutants (reg1 and reg2) causing constitutive expression of the emu operon (which has genes emu1 and emu2). One mutant contains a defect in a DNA-binding site, and the other has a loss-of-function defect in the gene encoding a protein that binds to the site Say you don’t know which mutant has a defect in the site and which one has a mutation in the binding protein. To figure it out, you construct the two partial diploid strains (i and ii below), and you then assay the levels of the Emu1 and Emu2 proteins in these two strains. F’ (reg1- reg2+ emu1- emu2+) / reg1+ reg2+ emu1+ emu2- F’ (reg1+ reg2- emu1- emu2+) / reg1+ reg2+ emu1+ emu2- What proteins do you predict will be expressed for strains i and ii if reg2 encodes the regulatory protein and reg1 is the regulatory site?arrow_forwardSuppose you have six strains of E. coli. One is wildtype, and each of the other five has a single one of thefollowing mutations: lacZ−, lacY−, lacI−, oc, andlacIS. For each of these six strains, describe thephenotype you would observe using the following assays. [Notes: (1) IPTG is a colorless synthetic molecule that acts as an inducer of lac operon expressionbut cannot serve as a carbon source for bacterialgrowth because it cannot be cleaved byβ-galactosidase; (2) X-gal cannot serve as a carbonsource for growth; (3) E. coli requires active lactosepermease (the product of lacY) to allow lactose,X-gal, or IPTG into the cells.] Colony color in medium containing glycerol as theonly carbon source and X-gal, but no IPTG.d. Colony color in medium containing high levels ofglucose as the only carbon source, X-gal, andIPTG.e. Colony color in medium containing high levels ofglucose as the only carbon source and X-gal, butno IPTGarrow_forward. Suppose you have six strains of E. coli. One is wildtype, and each of the other five has a single one of thefollowing mutations: lacZ−, lacY−, lacI−, oc, andlacIS. For each of these six strains, describe thephenotype you would observe using the following assays. [Notes: (1) IPTG is a colorless synthetic molecule that acts as an inducer of lac operon expressionbut cannot serve as a carbon source for bacterialgrowth because it cannot be cleaved byβ-galactosidase; (2) X-gal cannot serve as a carbonsource for growth; (3) E. coli requires active lactosepermease (the product of lacY) to allow lactose,X-gal, or IPTG into the cells.]a. Growth on medium in which the only carbonsource was lactose.b. Colony color in medium containing glycerol as theonly carbon source, X-gal, and IPTGarrow_forward
- There is Hyaluronic acid synthesis occuring in Group X Strep and it is controlled by an operon with 3 genes, called hasXYZ. Based on the 3-line diagram model, a. How many ribosome binding sites are there for the protein? b. How many promoters are there for the genes? c. How many start codons are there for the protein? d. How many RNA Polymerase binding locations are there for the genes? e. How many proteins will be fully functional? f. How many mRNA strands are made?arrow_forwardHow long would it take for the E. coli RNA polymerase to synthesize the primary transcript for the E. coli genes encoding the enzymes for lactose metabolism, the 5,300 bp5,300 bp lac operon? Assume an average elongation rate of 7070 nucleotides per second. a)How far along the DNA would the transcription "bubble" formed by RNA polymerase move in 10 seconds10 seconds? b)Assuming that human Pol II transcribes at a similar rate, how long does it take to transcribe the 2,000,000 bp2,000,000 bp dystrophin gene?arrow_forwardThe symbols A, B, C, D, E represent all the components of the lac operon, excluding LacA. The symbols are not in the order of the lac operon. Minus and plus signs represent activity of lacZ or LacY in the absence or presence of IPTG. LacI+ or LacI-, and LacO+ or LacOC alleles were used. Which of the symbols represents each of the lac operon components?arrow_forward
- You have isolated two different mutants (reg1 andreg2) causing constitutive expression of the emu operon (emu1 emu2). One mutant contains a defect in aDNA-binding site, and the other has a loss-of-functiondefect in the gene encoding a protein that binds tothe site.a. Is the DNA-binding protein a positive or negativeregulator of gene expression?b. To determine which mutant has a defect in the siteand which one has a mutation in the binding protein, you decide to do an analysis using F′ plasmids. Assuming you can assay levels of the Emu1and Emu2 proteins, what results do you predict forthe two strains (i and ii; see descriptions below) ifreg2 encodes the regulatory protein and reg1 is theregulatory site?i. F′ (reg1−reg2+emu1−emu2+)/reg1+reg2+emu1+emu2−ii. F′ (reg1+reg2−emu1−emu2+)/reg1+reg2+emu1+emu2−c. What results do you predict for the two strains(i and ii) if reg1 encodes the regulatory proteinand reg2 is the regulatory site?arrow_forward. a. How many ribosomes are required (at a minimum)for the translation of trpE and trpC from a singletranscript of the trp operon?b. How would you expect deletion of the two tryptophan codons in the RNA leader to affect theexpression of the trpE and trpC genes?arrow_forwardA marine bacterium is isolated and shown to contain an inducible operon whose genetic products metabolize oil when it is encountered in the environment. Investigation demonstrates that the operon is under positive control and that there is a reg gene whose product interacts with an operator region (o) to regulate the structural genes, designated sg. In an attempt to understand how the operon functions, a constitutive mutant strain and several partial diploid strains were isolated and tested with the results shown in the following table. Host Chromosome F= Factor Phenotype Wild type None Inducible Wild type reg gene from mutant strain Inducible Wild type Operon from mutant strain Constitutive Mutant strain reg gene from wild type Constitutive Draw all possible conclusions about the mutation as well as the nature of regulation of the operon. Is the constitutive mutation in the trans-acting reg element or in the cis-acting o operator element?arrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
