
Concept explainers
(a)
To draw:
The diagram that consist of 5 genes and the enhancers. Suppose that the blue, orange, green, red, purple and black are the activator proteins that binds to the appropriate color-coded control elements of the enhancers.
(a)
To draw an X above the enhancer elements that will transcribe only gene 5 and also, identify the colored activators present.
(b)
To draw a dot above the enhancer elements where blue, orange and green activators are present. Also, identify the transcribed genes.
(c)
Suppose, the genes 1, 2 and 4 codes for the proteins that are nerve-specific and genes 3 and 5 are the proteins that are skin-specific. To identify the activators that will ensure transcription of the appropriate genes.
Concept introduction:
A gene (eukaryotic) and the elements of DNA (deoxyribonucleic acid) are organized properly. The transcription initiation complex (cluster of proteins) assembles on the promoter sequence and the processing or transcription process is initiated.
Pictorial representation:
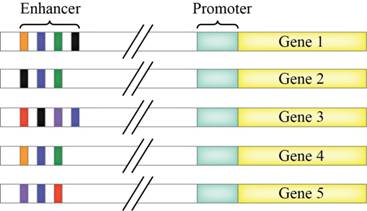
Figure 1: Transcription of genes with the help of enhancer elements.
To draw:
An X above the enhancer elements that will transcribe only gene 5 and also, identify the colored activators present.
(b)
To draw:
A dot above the enhancer elements where blue, orange and green activators are present. Also, identify the associated transcribed genes.
(c)
To suppose:
The genes 1, 2 and 4 codes for the proteins that are nerve-specific and genes 3 and 5 codes for the proteins that are skin-specific. And, identify the activators that will ensure transcription of the appropriate genes.
Want to see the full answer?
Check out a sample textbook solution
Chapter 15 Solutions
Campbell Biology in Focus; Modified Mastering Biology with Pearson eText -- ValuePack Access Card -- for Campbell Biology in Focus (2nd Edition)
- 3). Consider the four mutations (i-iv) described below: i. One of the mutations causing cystic fibrosis in humans is a deletion of three nucleotides that eliminates a phenylalanine at position 508 of the CFTR protein (D508). Normally, CFTR protein is localized to the plasma membrane, where it functions as a chloride ion channel. D508 CFTR is misfolded and all of it is degraded without ever reaching the cell surface. ii. The yeast transcription factor Gal4p contains a DNA-binding domain and a transcriptional activation domain. An allele with a deletion the gene portion encoding the activation domain encodes a truncated Gal4p containing only the DNA-binding domain. It binds to Gal4p target genes at appropriate binding sites in their upstream regulatory regions, but does not activate their transcription. In cells with both wild type and mutant forms of Gal4p, the truncated Gal4p binds more efficiently to target DNA sequences than wild type. iii. Mutations in the acid maltase gene in…arrow_forwardThe figure below shows 6 genes that are differentially expressed based on their specific enhancers. The different activators (A-F) are proteins that can bind to the appropriate control element in the enhancers of these genes. Use the diagram to answer the following question: What gene(s) would be transcribed if you A, C, E and F enhancers had activators bound to them? No need to explain your answer – just provide the gene(s). ABCD Promoter Brain specific gene ABC F Promoter Liver specific gene C Promoter Heart specific gene A CD Promoter Skin specific gene AB D Promoter Gut specific gene EF Promoter Nose specific genearrow_forward13. As you recall, a process called catabolite repression prevents high level of transcription of lacZYA genes when glucose is present in the growth medium. Low levels of glucose lead to production of CAMP, which enables CRP to bind to the lacZYA promoter. If the gene encoding CRP were to experience a loss-of-function mutation, what would be the expected outcome? a. Reduced level of lacZYA transcription in a medium containing only lactose (no glucose) b. No change in lacZYA transcription in a medium containing only lactose (no glucose) c. Increased level of lacZYA transcription in a medium containing only lactose (no glucose) e. Increased level of lacZYA transcription in a medium containing both lactose and glucosearrow_forward
- 3. Figure 2 illustrates how Pitx1 transcription is regulated in different tissues. The center image is that of a stickleback embryo. The drawings in the surrounding boxes show the Pitx1 gene region and activator proteins present in the jaw, pelvis, eye, or pituitary tissues. a. List all the tissues shown in Figure 2 that express the Pitx1 gene. b. If a fish does not produce activator 1 proteins because of a mutation in the gene that encodes those proteins, Pitx1 will be expressed in which of the following tissues? c. If a fish does not produce activator 3 proteins, Pitx1 will be expressed in which of the following tissues? d. A fish inherits a mutation in the Pitx1 coding region. This is a nonsense mutation that introduces a premature stop codon, resulting in a nonfunctional protein. Where would you expect Pitx1 to be expressed in this scenario?arrow_forward2. Predict and explain the effect on GAL1 transcription, in the presence of galactose alone, of the following mutations (image from book pasted for your convenience): a. Deletion of one Gal4-binding site in the GAL1 UAS element. b. Deletion of all four Gal4-binding sites in the GAL1 UAS element. c. Deletion of the Mig1-binding site upstream of GAL1. Deletion of the Gal4 transactivation domain. d. e. Deletion of the GAL80 gene. f. Deletion of the GAL1 promoter. g. Deletion of the GAL3 gene. (a) (b) Glucose Galactose Gal3p Glucose Galactose + Glucose Galactose (c) + UAS GAL Figure 21-10 Molecular Biology: Principles and Practice, Second Edition 2015 Macmillan Education Gal80p -Gal4p Mig1- binding site -ATP Tup1 -ATP Mig1 OFF (basal) GAL ON GAL OFF GALarrow_forward1. Draw a simplified model of a cis-regulatory element with multiple trans-acting factors up-regulating gene expression. Your simplified model must include: a double-stranded DNA molecule labeled with its appropriate orientation (3' and 5', etc.), an area of DNA highlighting the gene of interest to be transcribed, a highlighted area showing the promoter region, one cis-regulatory element upstream of the promoter with its specific transcription factor • RNApol + a general transcription factor + a TATA binding protein all bound to the appropriate area (think about where these bind before transcription starts), an upstream enhancer with its transcription factor bound to it, • and finally the TSS. All parts must be labeled and in the correct area to earn full credit. **I suggest using different colors for this drawing problem**.arrow_forward
- 4e. You also study the expression of 3 different mutants for this gene. For each mutant answer the following: Does this mutation change the sequence of the protein produced? Why or why not? If it does change the sequence of protein be sure to write out the new sequence. If it does not change the protein sequence, what effect (if any) would you expect it to have on expression of the gene? 1 20 ORI 40 60 5'...TTCGAGCTCTCGTCGTCGAGATACGCGATGATATTACTGGTAATATGGGGATGCACTATC...3' 3' ...AAGCTCGAGAGCAGCAGCTCTATGCGCTACTATAATGACCATTATACCCCTACGTGATAG...5' * promoterarrow_forward6) The functioning of the "Ras/MAPK" signal transduction pathway is absolutely essential in order for cells to grow, divide, and migrate. One important protein that is part of this pathway is BRAF. This protein is a kind of enzyme called a "kinase" – an enzyme that transfers a phosphate group onto another protein. - In some melanomas, a mutated form of BRAF called BRAF Val600AGlu drives the progression of the cancer. The drug “vemurafenib" slows the progression of the cancer by slowing the production of the mutant BRAF protein. (National Cancer Institute. 2019. Types of Cancer Treatment. Retrieved from: https://www.cancer.gov/about-cancer/treatment/types/ Is this an example of a traditional cancer therapy or a targeted therapy? Briefly explain your reasoning in the space provided, using information provided in the text to support your answer. Type of therapy (traditional or targeted)?: Brief explanation: 19arrow_forward1. Histone methylation can have many different effects on gene expression. In some cases, histone methylation is associated with activation of transcription, whereas in other cases it can trigger the formation of heterochromatin and a decrease in transcription. If histone methylation has been detected in the region of gene YFG in yeast, describe an experiment that could distinguish whether the methylation is important to activate or repress transcription of gene YFG. al 6 2. An E. coli strain of chromosomal genotype lacl* lacP" lacO* lacz* lacY*. You wish to transform this strain into a wild-type lac operon by the addition of an extra piece of DNA (plasmid). a. What regulatory region(s), gene or genes would you add on this extra DNA that would make this strain express the lac operon genes as wild type? b. How would you design this plasmid if you want the strain of E. coli to express the lac operon genes even if lactose is absent in the medium (constitutively)?arrow_forward
- 1. Describe which enzymes are required for lactose and tryptophan metabolism in bacteria when lactose and tryptophan, respectively, are (a) present and (b) absent. 2. Contrast positive versus negative regulation of gene expression. Describe the role of the repressor in an inducible system and in a repressible system.arrow_forward12. a. You want to create a genetic construct that will express GFP in Drosophila. In addition to the GFPcoding sequence, what DNA element(s) must youinclude in order to express this protein in flies if theconstruct were integrated into the Drosophila genome? Where should such DNA element(s) be located? How would you ensure that GFP is expressedonly in certain tissues of the fly, such as the wing?b. Suppose you insert the GFP coding region plus allof the DNA elements required by the answer to part(a)—except the enhancer—between inverted repeatsfound at the ends of a particular transposable element.Because all of the DNA sequences located betweenthese inverted repeats can move from place to placein the Drosophila genome, you can generate manydifferent fly strains, each with the construct integrated at a different genomic location. You now examine animals from each strain for GFPfluorescence. Animals from different strains showdifferent patterns: some glow green in the eyes,others in…arrow_forwardThe following diagram illustrates four genes from the genome of a certain insect. Different binding sites are labeled in the enhancer region of each gene. enhancer promoter ABC D E Gene 1 A D Gene 2 АС DE Gene 3 ABD Gene 4 In order for a specific gene to be expressed in the insect's cells, all of the gene's binding sites must be bound by transcriptional activators. • The insect's brain cells contain activators that bind to sites A, C, D, and E • The insect's salivary gland cells contain activators that bind to sites A, B, and D Which of the following is the most likely pattern of gene expression for the insect described above? Gene 2 will be expressed in both brain and salivary gland cells Both gene 1 and gene 4 will be expressed in salivary glands, but neither will be expressed in brain cells Gene 1 will be expressed in both brain and salivary glands Both gene 2 and gene 3 will be expressed in brain cells, but neither will be expressed in salivary gland cellsarrow_forward
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning Biology: The Unity and Diversity of Life (MindTap...BiologyISBN:9781305073951Author:Cecie Starr, Ralph Taggart, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology: The Unity and Diversity of Life (MindTap...BiologyISBN:9781305073951Author:Cecie Starr, Ralph Taggart, Christine Evers, Lisa StarrPublisher:Cengage Learning

