
Concept explainers
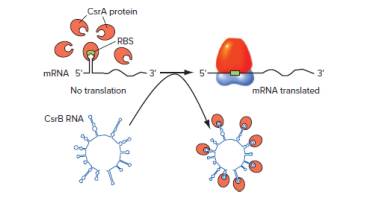
Great variation exists in the mechanisms by which RNAs can mediate gene regulation. In one recently discovered example shown in the following diagram, the genes CsrA and CsrB are global regulators of suites of target genes that are involved in the use of carbon atoms. The product of CsrA is the CsrA protein, which binds to the ribosome binding site (RBS) of target gene mRNA, preventing target gene expression. The product of the CsrB
gene is the CsrB RNA, which contains 22 binding sites for CsrA protein. CsrB RNA can thus compete with target mRNAs for CsrA protein binding. In the presence of high CsrB RNA concentrations, CsrA protein cannot bind to mRNA binding sites, so expression of the target genes is turned on.
| a. For the CsrA and CsrB genes, indicate what kind of molecule the gene product is (DNA, RNA, protein, small molecule), whether it acts as a positive or negative regulator, what stage of gene expression it affects, and whether it acts in cis or in trans. (It will be interesting to compare your answers to those for Problem 30.) |
| b. CsrA/CsrB regulate glycogen biosynthesis and breakdown; glycogen is a To what environmental factor do you think that the CsrA/CsrB system is most likely to respond? Suggest a possible way that this system might be repressible, and then suggest a different hypothesis for how this system might be inducible. (Assume in both cases that CsrB expression is modulated.) Which of your hypotheses would be most consistent with the target genes being involved in glycogen biosynthesis (an anabolic pathway), and which is most consistent with glycogen breakdown (a catabolic pathway)? Explain. |
Want to see the full answer?
Check out a sample textbook solution
Chapter 16 Solutions
ND STONY BROOK UNIVERSITY LOOSELEAF GENETICS: FROM GENES TO GENOMES
- Compare the control of gene regulation in eukaryotes and prokaryotes at the level of initiation of transcription. How do the regulatory mechanisms work? What are the similarities and differences in these two types of organisms in terms of the specific components of the regulatory mechanisms? Address how the differences or similarities relate to the biological context of the control of gene expression.arrow_forwardDiscuss the following argument: “if the expression of every gene depends on a set of transcription regulators, then the expression of these regulators must also depend on the expression of other regulators, and their expression must depend on the expression of still other regulators, and so on. cells would therefore need an infinite number of genes, most of which would code for transcription regulators.” how does the cell get by without having to achieve the impossible?arrow_forwardTranscriptional repressor proteins (e.g., lac repressor), antisense RNA, and feedback inhibition are three different mechanisms that turn off the expression of genes and gene products. Which of these three mechanisms will be most effective in each of the following situations? A. Shutting down the synthesis of a polypeptide B. Shutting down the synthesis of mRNA C. Shutting off the function of a protein For your answers to parts A–C that list more than one mechanism, which mechanism will be the fastest or the most efficient?arrow_forward
- A strain of Arabidopsis thaliana possesses a mutation in the APETALA2 gene. As a result of this mutation, much of the 3′ UTR of the mRNA transcribed from the gene is deleted. What is the most likely effect of this mutation on the expression of the APETALA2 gene?arrow_forward(c) By binding one L-tryptophan molecule/monomer, the trp repressor binds to DNA to suppress syn- thesis of L-tryptophan in E. coli. Below is the amino acid sequence of the helix – (reverse) turn – helix region of the trp repressor that binds to DNA compared to the sequence of the corresponding DNA binding motif of the Prl protein, a different type of repressor protein. A diagram of the trp repressor dimer is also shown. reverse turn trp helix 4 70 Trp -Gly-Glu-Met-Ser-Gln-Arg-Glu-Leu-Lys-Asn-Glu-Leu-Gly-Ala-Gly- Ile- Prl -Ser-Glu-Glu-Ala-Lys-Glu-Glu-Leu-Ala-Lys-Lys-Cys-Gly-Ile-Thr- Val- Pri heilix trp helix 5 80 90 Trp Ala-Thr-Ile-Thr-Arg-Gly-Ser sgn-Ser-Leu-Lys-Ala-Ala- Prl Ser-Gln-Val-Ser-Asn-Trp-Phe-Gly-Asn-Lys-Arg-Ile-Arg- Prl helixarrow_forwardGene X codes for a protein in eukaryotes. A mutated eukaryotic cell contains an altered base-pair in an intron of gene X. Which would be the most likely effect of this mutation on the biomolecules in the cell? The amount of pre-mRNA transcribed from gene X would be less than normal. The amount of functional protein corresponding to gene X would be less than normal. The ability of snRNAs to form a spliceosome would be diminished. The breakdown of mature mRNA corresponding to gene X would be fasterarrow_forward
- Gal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forwardennar region of gene X, which determines the length of the tail in mice, is mutated so that transcription factors bind it at a much higher affinity compared to the wild-type sequence. What is the most likely phenotypic outcome? Tail length will not change because the enhancer is a non-coding sequence Tail length will increased due to increased activity of the gene's promoter Tail length will decreased because any mutation will cause a loss-of-function of these regulatory regions Not just the tail will be enlarged because increased activity of the enhancer will impact many genesarrow_forwardThe following DNA sequence has been determined from DNA isolated from a bit of prehistoric amber material (picture). It corresponds to a complete transcriptional unit without introns. Use the Genetic Code to predict the primary sequence of the polypeptide encoded by this preserved DNA. (Show relevant molecular intermediates, and provide detailed and appropriate labels)arrow_forward
- The lac genotypes are as shown below: P+OcZ-Y+A+// P¯O+Z+Y+A+ (i) The lac operon consists of three structural genes, lacZ, lacY and lacA. Which structural genes are involved in lactose metabolism? Explain. (ii) Draw and explain how lactose repress the gene expression in lac IS/I- heterozygote. (iii) What is the function of the promoter in the bacterial operon?arrow_forwardWhile investigating the function of a specific growth factor receptor gene from humans, researchers found that two types of proteins are synthesized from this gene. A larger protein containing a membrane spanning domain recognizes growth factors at the cell surface, stimulating a specific downstream signaling pathway. In contrast, a related, smaller protein is secreted from the cell and binds available growth factor circulating in the blood, thus inhibiting the downstream signaling pathway. Speculate on how the cell synthesizes these disparate proteins.arrow_forwardATM is a kinase that phosphorylates histone H2AX in response to double-stranded DNA breaks. Which of the following scenarios would most quickly regulate ATM activity in the cell? a) Adding silencing methyl groups to cytosines in the Atm gene b) Modifying the histone code for the Atm gene c) Increasing expression of a miRNA specific for the Atm mRNA d) Activating an E3 ubiquitin ligase specific for the ATM proteinarrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





