
Concept explainers
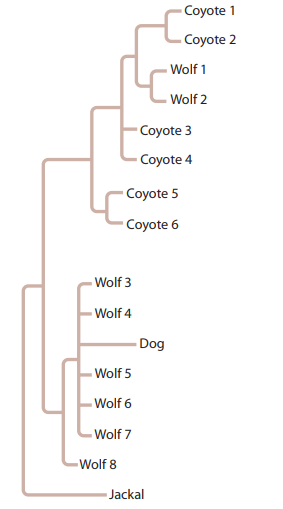
Wolves and coyotes can interbreed in captivity; and now, because of changes in their habitat distribution, they may have the opportunity to interbreed in the wild. To examine this possibility, mitochondrial DNA from wolf and coyote populations throughout North America
Want to see the full answer?
Check out a sample textbook solution
Chapter 17 Solutions
Pearson eText Genetic Analysis: An Integrated Approach -- Instant Access (Pearson+)
- A group of researchers are trying to determine the relationship between various groups of organisms. The researchers analyze the amino acid sequence of the protein cytochrome c in various groups of organisms. Once they sequence cytochrome c in all organisms, they determine the number of amino acid substitutions in each group. The results can be seen in the graph below, which plots the data with respect to the time since the divergence of the members of paired groups from a common ancestor. A B mammals and reptiles с Number of Amino Acid Substitutions (per 100 amino acids) in Cytochrome, c D 60- 50+ 40+ 30- 20+ 10+ 0 0 O birds and reptiles A graph showing when various groups of organisms diverged from a common ancestor Based on the graph above, can you conclude which of the following organisms are most distantly related? 200 400 600 800 1,000 Time Since Divergence from a Common Ancestor (millions of years) fish and land vertebrates mammals and reptiles birds and mammals insects and…arrow_forwardSuppose species 1, 2, and 3 are endemic to a group of islands (such as the Galápagos) and are all descended from species 4 on the mainland (which will serve as an outgroup; its large population size means that no new mutations have become fixed in its population in the time since the islands were colonized). You sequence a gene and find ten nucleotide sites that differ among the four species (among many other loci that do not vary). The nucleotide bases at these sites are: Species 1: GCTGATGAGT Species 2: ATCAATGAGT Species 3: GTTGCAACGT Species 4: GTCAATGACA Estimate the phylogeny of these taxa by plotting the changes on each of the three possible unrooted trees and determining which tree requires the fewest evolutionary changes.arrow_forwardA fossil believed to be the most recent common ancestor between housecats and lions is radio-dated to about 11 million years ago (mya). An analysis of a neutral section of mitochondrial DNA in both species finds 15 DNA base pair differences between the two cats today. Meanwhile, an analysis of housecats and pumas shows 4 DNA base pair differences between them, which are believed to have diverged around 3 mya. Using this data, what is the best estimate of the time since divergence of housecats and lynx, which have 11 DNA base pair differences in this mitochondrial DNA region? 5 mya O6 mya O 7 mya O8 mya 9 myaarrow_forward
- Which of the following best explains the number of similarities between the amino acid sequences of the Drosophila Hedgehog protein and the Chicken Indian Hedgehog protein? O A. The Drosophila hedgehog gene evolved from hedgehogs, which are distantly related to birds. O B. Both genes evolved from a gene present in the last common ancestor of Drosphila and chickens, and the number of differences reflects the amount of time that has elapsed during the evolution of these two lineages. a During the evolution of Drosophila and chickens, a hedgehog like gene arose independently in each lineage, then the gene that arose in chickens diversified. A These genes are unrelated, and the fact that they are similar is only because the proteins need to have similar biochemical properties. They are unrelated because chickens don't have segments and Drosophila larvae don't have limb buds.arrow_forwardHumans carry a variety of non-functional genetic sequences, called processed pseudogenes, in their DNA. we can estimate how long ago these sequences first appeared in the genomes of our ancestors. In humans, processed pseudogenes include the three options below which would be the least widespread among other primate species a. alpha-enolase psi1 (11 million years old) b. AS PSI 7 (16 million years old) c. CALM II PSI3 (36 million years old)arrow_forwardThe platypus is one of a very small number of mammals that are venomous. Researchers compared the substances in the platypus venom to that of venomous reptiles. They found that the venom in platypus was derived from a protein called defensin, while that in many reptiles was based on a protein called crotamine. The protein structure of these molecules is remarkably similar, though they are controlled by unrelated genes in each taxa. Both molecules originated as non-toxic antimicrobial compounds. Part A: Venom in both platypus and reptiles is an example of: A. Homologous traits B. Analogous traits C. Adaptive radiation D. Genetic drift Part B: A trait matrix for the amniotes is below. Using the trait matrix, match the taxa to the letters at the branch tips. (see attached image) Using the image, match the taxa to the letters at the branch tips 1. Chicken 2. Kangaroo 3. Viviparity 4. Placenta Part C: In which location(s) did each trait evolve on the phylogeny? If more than one location…arrow_forward
- DNA-DNA hybridization studies in the 1960s demonstrated that common chimpanzees (Pan troglodytes), and pygmy chimpanzees (Pan paniscus), are most closely related to which of the following non-chimpanzee primate species (sharing about 98% of protein-coding nucleotides)? the agile gibbon (Hylobates agilis) the human (Homo sapiens) the orangutan (Pongo pygmaeus) the gorilla (Gorilla gorilla) the olive baboon (Papio anubis) 7. Which of the following mobile genetic elements is a retrotransposon, making up about 10% of the human genome, but no longer codes for a functional gene product (and is therefore a “pseudogene”)? the LINE-1 element (repeated up to 50,000 times in the human genome) the DNA-only transposons known as “jumping genes” (in corn) the avocado sunblotch viroid the typical transposon found in a bacterial chromosome the Alu sequence (repeated up to 1,000,000 times in the human genome) . Which of the following self-replicating biological agents carries only one…arrow_forwardYour descendants have fortunately inherited your fascination with rodent genetics. Fifty thousand years after the original volcanic eruption, your descendants return to the island where the two populations are still separated. They repeat the same experiments that you carried out. Their data indicate there is no reproductive isolation between members of A1 and A2. DNA sequences from randomly chosen individuals from the two populations are shown here – A1 ATAAGA A1 ACAAGA A1 ACATGA A2 TCATTT A2 TCATTG A2 TCATTT A)Name one mechanism by which differences between A1 and A2 became fixed in the two populations (not how the differences arose in the first place).arrow_forward50. The polar bear (Ursus arctos maritimus) is thought to have evolved from the Siberian brown bear (Ursus arctos beringianus), in the Kamchatka peninsula of Russia during the Riss glacial period, about 150,000 years ago, when advancing glaciers isolated a subpopulation of brown bears in Kamchatka, preventing them from interbreeding with the ancestral brown bear population in mainland Asia. As the Kamchatka bears adapted to a tundra habitat (evolving into polar bears), they became distinct from the ancestral brown bears, who remained adapted to a boreal forest habitat. With the boundary between tundra and boreal forest persisting to the present, isolating modern brown bears from polar bears, the descendant species (tundra-dwelling polar bears) probably originated from the ancestral species (boreal forest-dwelling brown bears) by means of which of the following pre-zygotic mechanisms? A. speciation by F2 hybrid breakdown B. allopatric speciation by means of vicariance C. speciation by…arrow_forward
- Your descendants have fortunately inherited your fascination with rodent genetics. Fifty thousand years after the original volcanic eruption, your descendants return to the island where the two populations are still separated. They repeat the same experiments that you carried out. Their data indicate there is no reproductive isolation between members of A1 and A2. DNA sequences from randomly chosen individuals from the two populations are shown here – A1 ATAAGA A1 ACAAGA A1 ACATGA A2 TCATTT A2 TCATTG A2 TCATTT A)Describe relative levels of genetic variation (simply state low or high) among individuals in population A1. B)Describe relative levels of genetic variation (simply state low or high) among individuals in population A2. C)Describe relative levels of genetic variation (simply state low or high) between individuals in populations A1 and individuals in population A2.arrow_forwardYou are studying two populations of pika. You find that the population inhabit in talus (broken rocks) are greyish. Another population live in coniferous forests have black fur. When individuals from each population are brought together in the lab, they produce offspring whose appearance is intermediate between the two parents. The offspring can breed and reproduce successfully with each other or pika of either parent population. You sequence three genes, Pigmentosa, Sparkly, and Zippy. For each of the three genes, the two populations differ from each other at a few nucleotides (<1%). Why is it hard to determine if these two pika populations belong to the same species or are two different species? In your answer/argument, refer to at least two pieces of evidence from the above scenario. Note, we are NOT asking you to determine if these are the same species or not.arrow_forwardA. What is the wild progenitor of maize and where is it found? B. George Beadle concluded that this plant was the likely ancestor of maize (corn) even though the two plants appear very different. What evidence did Dr. Beadle collect that led to his conclusion? C. How long ago was maize domesticated and what evidence was utilized to determine this? D. Dr. Doebley and his team compared the DNA sequence of maize to that of a number of teosinte varieties from throughout Mexico. What did their analysis reveal?arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
