
PRESCOTT'S MICROBIO W/PROCTORIO
11th Edition
ISBN: 9781264731060
Author: WILLEY
Publisher: MCG
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 18.3, Problem 2CC
Retrieve, Infer, Apply
Examine figure 18.8. How might metagenomics be used to isolate genes encoding a potentially new antibiotic?
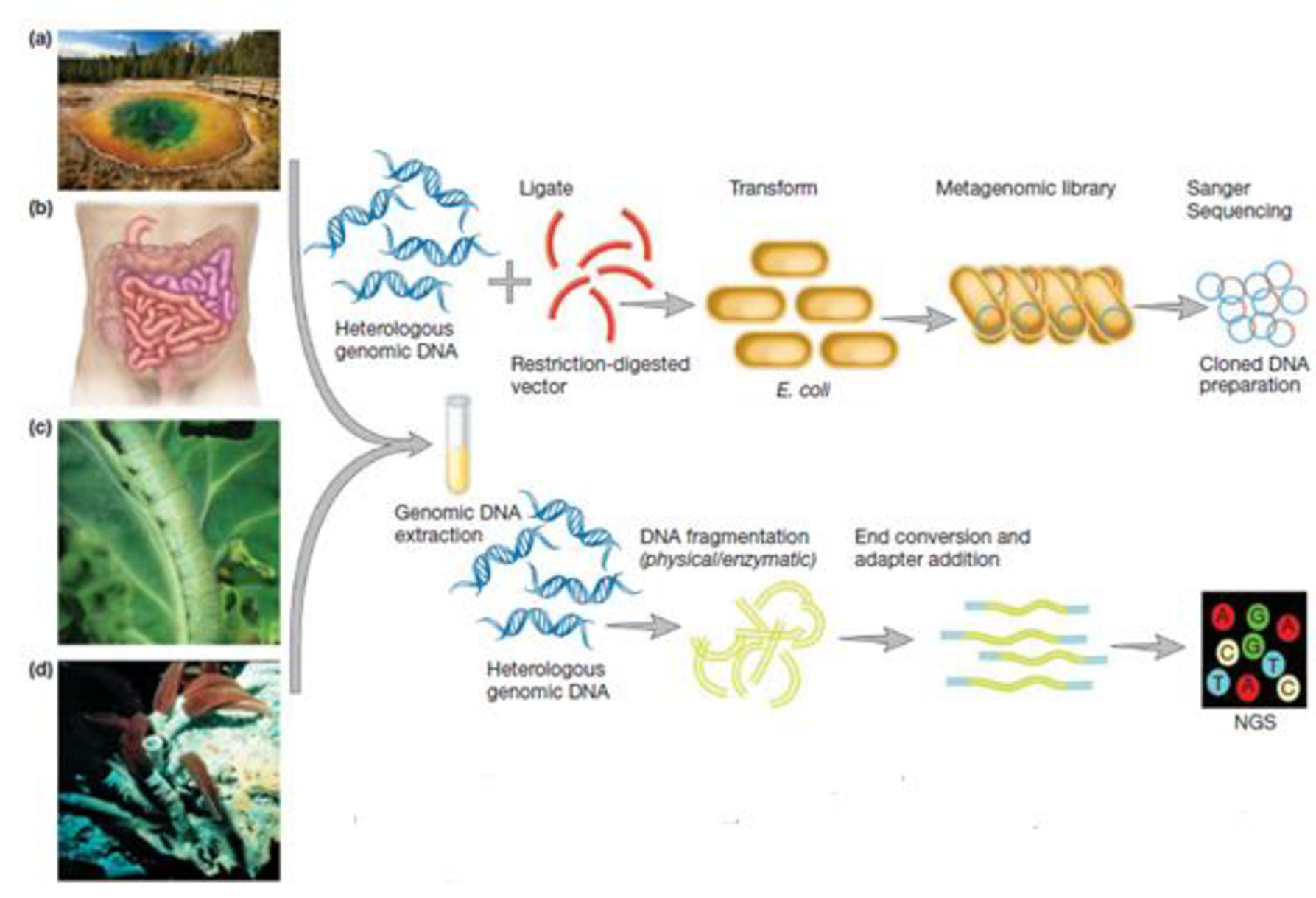
Figure 18.8 Metagenomic analysis. DNA has been extracted directly from environments such as (a) bacterial mats at Yellowstone National Park, (b) the human colon, (c) cabbage white butterfly larvae, and (d) tube worms from hydrothermal vents. The DNA can be used to construct a metagenomic library and analyzed by Sanger sequencing (top); however, NGS is more likely to be used (bottom), thereby avoiding the need for genomic library construction.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Do not use chatgpt.
Give
Please help answer the starred question
Chapter 18 Solutions
PRESCOTT'S MICROBIO W/PROCTORIO
Ch. 18.1 - MICRO INQUIRY What is the function of the 3-OH...Ch. 18.1 - MICRO INQUIRY Why is it important that identical...Ch. 18.2 - MICRO INQUIRY Which step (or steps) in this...Ch. 18.2 - Retrieve, Infer, Apply Why is the Sanger technique...Ch. 18.2 - Retrieve, Infer, Apply Explain the difference...Ch. 18.2 - Retrieve, Infer, Apply Why does reversible chain...Ch. 18.2 - Prob. 4CCCh. 18.2 - Retrieve, Infer, Apply Suggest a medical and an...Ch. 18.3 - Retrieve, Infer, Apply NGS techniques are...Ch. 18.3 - Retrieve, Infer, Apply Examine figure 18.8. How...
Ch. 18.4 - Prob. 1MICh. 18.4 - Prob. 1CCCh. 18.4 - Prob. 2CCCh. 18.4 - Prob. 3CCCh. 18.5 - Figure 18.12 Metabolic Pathways and Transport...Ch. 18.5 - Prob. 2MICh. 18.5 - Prob. 3MICh. 18.5 - Prob. 1CCCh. 18.5 - Retrieve, Infer, Apply How might the following...Ch. 18.5 - Retrieve, Infer, Apply Compare and contrast...Ch. 18.5 - Retrieve, Infer, Apply Why does two-dimensional...Ch. 18.5 - Retrieve, Infer, Apply What is the difference...Ch. 18.5 - Retrieve, Infer, Apply Describe a ChIP-Seq...Ch. 18.7 - Prob. 1MICh. 18.7 - Retrieve, Infer, Apply Cite an infectious disease...Ch. 18.7 - Prob. 2CCCh. 18.7 - Prob. 3CCCh. 18 - Prob. 1RCCh. 18 - Prob. 2RCCh. 18 - Prob. 3RCCh. 18 - Prob. 4RCCh. 18 - Prob. 5RCCh. 18 - Prob. 1ALCh. 18 - Prob. 2ALCh. 18 - You are developing a new vaccine for a pathogen....Ch. 18 - Prob. 4AL
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Discuss the underlying biochemical principle of the nucleic acid sequencing methods known as Semiconductor (Ion Torrent) sequencing.arrow_forwardSort the steps to be followed in order to produce human growth hormone (HGH) in the correct order Extract HGH from the medium Transform bacteria with the vector carrying HGH Digest (cut) HGH gene and plasmid using the same restriction enzyme Isolate the HGH gene from a human cell and a plasmid from a bacterium Clone the transgenic bacteria in the appropriate medium Mix HGH gene and plasmid; and use ligase (enzyme) to link them to form a rDNA Mo Question 7 of 15arrow_forwardwhat is the whole-genome shotgun sequencing? Also briefly explain its strategy to assemble the genome sequence.arrow_forward
- TOPIC: PCR and Gene Cloning Basics Question: What are 2 possible roles of CaCl2 in the transformation process?arrow_forwardWhat type of enzymes are used to “cut” desired DNA sequences for use in recombinant gene technology experiments? Identify those two enzymes used to cut and paste both genes into the plasmid. Identify all three strategies used in this lab to maximize transformation success. Explain what it means for bacteria to be “competent.” Explains why bacterial competency is this important for this investigation.arrow_forwardProvide five advantages of Next Generation Sequencing? and explain each of these advantages.arrow_forward
- Experiment A microarray was hybridized with a mixture of two differentially labeled fluorescent cDNAs, one prepared using retinal RNA of 1-day-old mice (labeled with a green fluorescent dye) and the other prepared using retinal RNA of 28-day-old mice (red fluorescence). The two probes were prepared from identical amounts of retinal tissues and were mixed together for hybridization to the microarray. Unhybridized probes were washed away, and the microarray was photographed in a fluorescence microscope.arrow_forwardwhat is the main purpose of performing the bioinformatics analysis of 16s rRNA genes lab?arrow_forwardBacteriophage lambda is as one of the routinely used molecular cloning vectors, which alsoserved as a model system for the study of bacteriophage morphogenesis, DNA replication, andgene regulation. (a)Blunt-end cloning is proven to be more challenging, compared to sticky-end cloning.Illustrate ONE (1) approach that can be conducted in order to perform blunt-end cloning ofa PCR product to a plasmid vector.d) Evaluate the advantages of mRNA as a source material for generating DNA fragments, inrelation to gene cloning.e) E. coli, in most cases is not a suitable host for cloning genes from eukaryotes. Discuss thisstatement.arrow_forward
- Transcriptome analysis involves two separate methodologies: gene expression and RNA seq analyses. The 10 items below are a scrambled listing of the steps used in the two procedures. Identify the steps involved in RNA seq from the list below. Use the numbers in the list to refer to each step. Once the steps for RNA seq have been identified, write the steps in the order in which they are performed during the experiment. (1) DNA sequencing (2) Allow for hybridization and wash excess cRNA. (3) Mix labeled cRNA with array chip. (4) PCR amplification (5) Measure fluorescence intensity to determine abundance of transcripts. (6) Add labeled cRNA at each microarray location. (7) Map cDNA sequences to the genome of the organism to determine identity and abundance of transcripts. (8) mRNA isolation from cells (9) Prepare fluorescently labeled cRNA probes (10) cDNA synthesisarrow_forwardPrimer Development Pick your favorite gene and develop primers for a target within the gene. Provide the following information: Gene name: Species: Accession number: Primer target: Primer Output: Explain why you selected this primer set.arrow_forwardNeed helparrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning
Molecular Techniques: Basic Concepts; Author: Dr. A's Clinical Lab Videos;https://www.youtube.com/watch?v=7HFHZy8h6z0;License: Standard Youtube License