
Concept explainers
Gene Transfer Between Bacteria
Core Skill: Connections Look back at Figures 11.1 and 11.2. How did the phenomenon of transformation allow researchers to demonstrate that DNA is the genetic material?
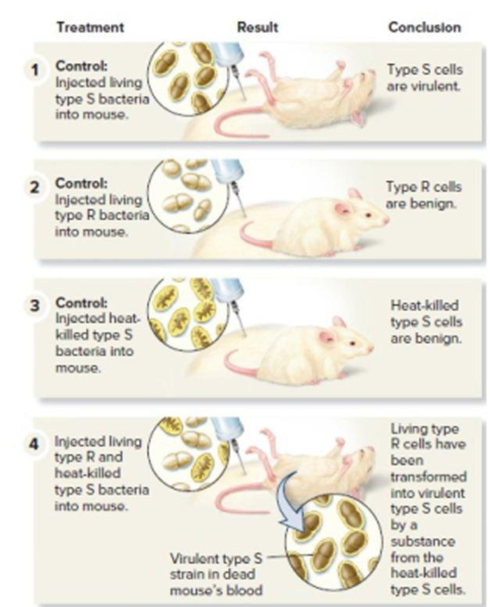
Figure 11.1 Griffith’s experiments showing that genetic material can be transferred from one bacterium to another. Note: To determine if a mouse’s blood contained live bacteria, a sample of blood was also applied to solid growth media. (This part of the procedure is not shown.) For steps 1 and 4, smooth bacterial colonies were observed. For step 2, no bacterial colonies were observed because the type R cells were killed by the immune system of the mouse.
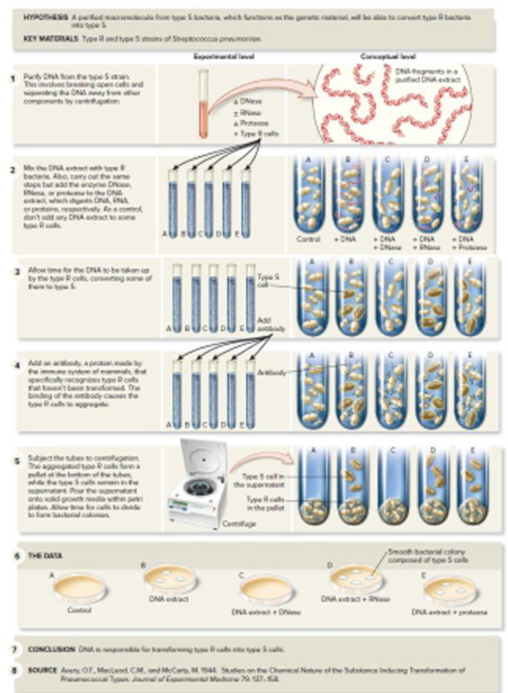
Figure 11.2 The Avery, MacLeod, and McCarty experiments that identified DNA as Griffith’s transformation principle—the genetic material.
Want to see the full answer?
Check out a sample textbook solution
Chapter 19 Solutions
Biology
- Compare the possible differences between a eukaryotic protein-encoding gene cloned by PCR and the same gene cloned by reverse transcriptase PCR (RTPCR).arrow_forwardlink: https://www.aaas.org/news/science-newly-identified-bacteria-break-down-tough-plastic How is the new PETase (2018) different from the 2016 version in function?arrow_forwardQ11) Explain, using the pCR 2.1 vector, what we would see if we try to grow cells on a plate with ampicillin and Xgal in each of these scenarios:a) transformation not successful (meaning the plasmid did not go inside the bacteria)b) transformation was successful, but no insert in LacZ gene (plasmid went in the bacteria but there was no inserted DNA in the LacZ gene of the plasmid)c) transformation and the ligation reactions were successful (got the plasmid in, and the plasmid is carrying extra DNA in the LacZ region, yay)arrow_forward
- Pls Help me. 2. If a selection assay be made to identify cells that have incorporated the recombinant DNA, what type of medium will be used? How will it work?arrow_forwardQ5. In lab 6, you will set up a PCR reaction to yield amplified DNA products of the rpo B(RNA polymerase B) and 16s rRNA genes. In lab 7, you will set up a sequencing reaction using about 40 ng of these PCR products that you will then add a sequencing primer to the sequencing reaction. If a sequencing reaction is set up with a 100-fold excess of primer to the PCR-amplified DNA product you want to sequence, calculate the number of sequencing primers you will need to add to this sequencing reaction. To perform this calculation, assume your 40 ng sample of double stranded DNA is 4,000 base-pairs (bp) long. Using these two variables of your PCR product [length (in bp) and amount (in g, grams)], you can calculate the number of DNA molecules being sequenced with two conversion factors using the formula below: The average weight of 1 bp is approximately ~650 (g/mole)/bp Avogadro’s number which is 6.02 X 1023 molecules/mole The easiest way to solve this problem is to figure out how much one…arrow_forwardEnhancement Questions: What do you think are the crucial steps in Polymerase Chain Reaction technique? Enumerate the advantage and disadvantage of Polymerase Chain Reaction technique?arrow_forward
- VISUAL SKILLS Griffith was trying to develop a vaccinefor S. pneumonia when he was surprised to discover thephenomenon of bacterial transformation. Look at the second and third panels of Figure 16.2. Based on these results,what result was he expecting in the fourth panel? Explain.arrow_forwardQ. Based on your understanding of plasmids’ answer the following questions.1. Why is a daughter cell that fails to inherit a kind of plasmid dies? 2. How transfer of Col E1 plasmid to an F cell occurs? 3. What component you will must include in constructing a plasmid cloning vector that can replicate in both bacteria and yeast? 4. How copy number of a plasmid may affect its segregation? 5. How the yeast 2u circle maintains 80 copies in a cell?arrow_forwardDescribe how PCR is used to amplify a specific gene from total cellular DNA.arrow_forward
- exercise 1: Determine the circular map of plasmid p101 for enzymes A, B and Cusing the size of the fragments (in kb) obtained after digestion with the enzymes.A) 17B) 17C) 10, 7A x B ) 12, 5A x C ) 10, 6, 1B x C ) 7, 6, 4Make sure to explain what each of the digest suggests (number of sites; size ofthe plasmid, etc..) and how you came up with the map you draw (thoughtprocess).arrow_forwardWhat type of enzymes are used to “cut” desired DNA sequences for use in recombinant gene technology experiments? Identify those two enzymes used to cut and paste both genes into the plasmid. Identify all three strategies used in this lab to maximize transformation success. Explain what it means for bacteria to be “competent.” Explains why bacterial competency is this important for this investigation.arrow_forwardExpand PCR? Describe the different Steps involved in this technique?arrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
