
Concept explainers
Core Skill: Modeling The goal of this modeling challenge is to revise the model shown in Figure 26.22 to account for recent data indicating that some modern Africans have a small amount of Neanderthal DNA.
Modeling Challenge: As discussed in this section, between 1% and 4% of the DNA of modern humans of European descent is derived from that of Neanderthals. Though most modern humans of African descent do not carry Neanderthal DNA, recent evidence has shown that some of them carry a very small amount. Assuming that the presence of this Neanderthal DNA is not due to recent interbreeding between modern humans of European and African descent, revise the model shown in Figure 26.22 to account for the observation.
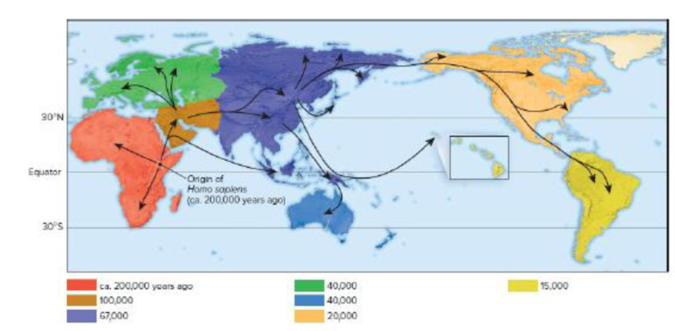
Figure 26.22 A simplified model for the origin and spread of Homo sapiens throughout the world. This map, based on differences of mtDNA throughout current members of the world’s population, suggests Homo sapiens originated in East Africa. About 100,000 years ago, the species spread into the Middle East and from there to Europe, Asia, Australia, and the Americas.
Want to see the full answer?
Check out a sample textbook solution
Chapter 26 Solutions
BIOLOGY (LL)
- Using the skills you learnt in the DNA Analysis tutorial, correctly identify the gene and the species of the following DNA sequence: AACCAGTGTGCTGCAGGCTGCACAGGCCCCCGGGAGAGCGACTGCCTGGTCTGCCGCAAA Using the skills you learnt in the DNA Analysis tutorial, correctly identify the gene and the species of the following DNA sequence: AACCAGTGTGCTGCAGGCTGCACAGGCCCCCGGGAGAGCGACTGCCTGGTCTGCCGCAAA Gene: RNA Polymerase I. Species: Homo Sapiens Gene: Epidermal Growth Factor Receptor (EGFR). Species: Homo Sapiens Gene: DNA Polymerase II. Species: Mus Musculus Gene: Insulin Receptor (INSR). Species: Homo Sapiensarrow_forwardGenome duplication in the animal lineage leading to chordates has produced the largest genome sizes in which of the following vertebrate groups? the cephalochordates (like the lancelet) the chondrichthyes (like the shark) the agnathan fishes (like the hagfish) the teleost fishes (like the trout) the urochordates (like the sea squirt)arrow_forwardEssay: Answer the following questions in paragraph form. Create atlest 3-5 sentences each to support your answer. Base on your own opinion, what does genetics mean? What is its relationship to all living things? What is the connection of Genetics and DNA with each other? Are they related or not? Give your own opinion and you can also cite an example if needed. What is DNA, rDNA and mtDNA? Explain each function. What are the advantages and disadvantages of genetic engineering? If you were given the chance to genetically modify an organism, what would it be and what would you like to modify from that organism?arrow_forward
- a. How does the genetic variation among humans compare to variation in other hominids and what does it indicate about population histories? b. What is the biggest structural difference between our genome and the other hominid genomes? c. What is the hominoid slowdown?arrow_forwardPlease answer fast What parts of the genome and genomic data are not well understood and still need more work? Realities and opportunities for the future?arrow_forwardHow does molecular biology paved way for the development of the DNA fingerprinting technique? and what is its molecular basis?arrow_forward
- Objective: Get a sense of how genomics, the study of the genome in its entirety,needs to think about how to go about its research. Geonomic DNA is broken up into fragments. The 5’ and 3’ ends of each fragment(a “read”) are sequenced. The sequenced reads are assembled together intocontiguous sequences (“contigs”) based on sequence similarity. The idea is to sequence enough random fragments so that every nucleotide in thegenome is represented on some read. The number of such fragments needed iscalled the coverage, c. The coverage c can be calculated by the formula RL/G, where R is the number ofreads sequenced, L is the average length of a read and G is the total length of thegenome. The units of length are bases (b) or base pairs (bp). Consider a genome whose length is 1000 bp. “Shotgun” sequencing techniquesare applied to the genome, resulting in 20 reads, with an average length of 50 bp.A very important point is that, even though 20 x 50 = 1000, there is no guaranteethat ALL…arrow_forwardObjective: Get a sense of how genomics, the study of the genome in its entirety,needs to think about how to go about its research. Geonomic DNA is broken up into fragments. The 5’ and 3’ ends of each fragment(a “read”) are sequenced. The sequenced reads are assembled together intocontiguous sequences (“contigs”) based on sequence similarity. The idea is to sequence enough random fragments so that every nucleotide in thegenome is represented on some read. The number of such fragments needed iscalled the coverage, c. The coverage c can be calculated by the formula RL/G, where R is the number ofreads sequenced, L is the average length of a read and G is the total length of thegenome. The units of length are bases (b) or base pairs (bp). Consider a genome whose length is 1000 bp. “Shotgun” sequencing techniquesare applied to the genome, resulting in 20 reads, with an average length of 50 bp.A very important point is that, even though 20 x 50 = 1000, there is no guaranteethat ALL…arrow_forwardProvide five advantages of Next Generation Sequencing? and explain each of these advantages.arrow_forward
- Assumptions: -There are approximately 3,000,000,000 base paris in the mammalian genome (genes constitute only a portion only a portion of this total. -There are approximately 10,000 genes in the mammalian genome. - A single gene averages 10,000 base pairs in size. -Only 1 out of 3 mutation that occur in gene result in a chnage to the protein structure. In the mammalian genome: How many total base-pairs are in all the mammalian genes? what proportion (%) of the total genome does this represent? What is the probability that a random mutation will occur in any given gene? What is the probability mutation will change the structure of a protein?arrow_forwardTopic: Recombinant pharmaceuticals (for the production of insulin, human growth hormone or blood clotting factors) Question Describe the molecular genetics process using proper scientific terminology. Describe the steps that are involved. How is it performed?arrow_forwardPractice... Lancelet 1. What trait separates Lampreys from tuna on (outgroup) Lamprey Tuna Salamander Turtie Leopard this cladogram? Hair 2. What separates a salamander from a Amniotic egg turtle? Four walking legs 3. Which organism is most related to the Jows leopard? Vertebral column 4. What 4 traits do these two organisms share? 5. Which organism will have DNA most similar to the turtle? 6. Which organism's DNA will differ the most from the leopard?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





