
Concept explainers
Answers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book.
Predicting a Sanger Sequencing Pattern The oligonucleotide d-AGATGCCTGACT as subjected to sequencing by Sanger’s dideoxy method, using fluorescent-tagged dideoxynucleotides and capillary electrophoresis, essentially as shown in Figure 11.3. Draw a diagram of the gel-banding pattern within the capillary.
Interpretation: A diagram of the gel-banding pattern within the capillary is to be drawn.
Concept introduction: A laboratory technique that is used for the separation of charged molecules such as proteins, DNA and RNA on the basis of their size is known as gel electrophoresis. This technique is useful to distinguish DNA fragments of various lengths.
Answer to Problem 1P
A diagram of the gel-banding pattern within the capillary is,
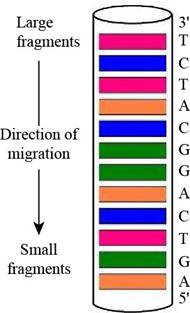
Explanation of Solution
In gel electrophoresis, using dyes such as radioactive labels or fluorescent tags makes it possible to see the DNA on the gel after separation. They are going to appear on the gel as bands. Therefore, the fluorescently labeled dideoxynucleotides result in the formation of the gel banding pattern. The labeled dideoxynucleotides are then added to the growing chain of DNA and capillary electrophoresis is applied to the resulting fragments.
The given oligonucleotide is d-AGATGCCTGACT that was subjected to sequencing by Sanger’s dideoxy method. In gel-banding pattern within the capillary, the top of the column consists of larger fragments and the bottom of the column has smaller fragments. The
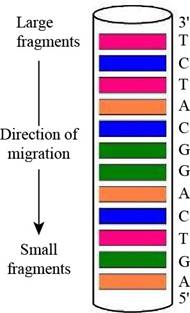
Figure 1
Want to see more full solutions like this?
Chapter 11 Solutions
BIOCHEMISTRY II >CUSTOM<
- Answers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Abundance of the Different Bases in the Human Genome Results on the human genome published in Science (Science 291 :1304—1350 [2001]) indicate that the haploid human genome consists of 2.91 gigabase pairs (2.91 X ]09 base pairs} and that 27% of the bases in human DNA are A. Calculate the number of A. T, G, and C residues in a typical human cell.arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Identify Proteins Using BLAST Searches of Peptide Fragment Sequences Go to the National Center for Biotechnology Information Web site at httlp:llhwww.ncbi.nlm.niih.goyl. From the menu (if Popular Resources on the right-hand side, click on “BLAST. Under the Basic BLAST heading on the new page that comes up, dick on protein blast. lit the Enter Query Sequence box at the top of the page that comes up, enter the following sequence: NQMMK.SR.N- LTKDRCKP. Confirm that the database under ChoOsC Search Set us set (111 nr (nonredundant protein Sequences), then click the BLAST button at the bottom (if the page td see the results of your search. Next, enter this sequence from a different protein: SLQTASAPDVYAlGfcCA. Identify the protein from which this sequence was derived.arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Preparing cDNA Libraries from Different Cells Describe an experimental protocol for the preparation of to cDNA libraries, one from anaerobically grown yeast cells and the second from aerobically grown yeast cell.arrow_forward
- Answers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Calculating Tms and Separating DNA Molecules That Differ in G:C Content At 0.2 M Na+, the melting temperature of double-stranded DNA is given by the formula, Tm = 69.3 + 0 41 (% G + C). The DNAs from mice and rats have (G + C) contents of 44% and 40%, respectively. Calculate the Tms for these DNAs in 0.2 M NaCl. If samples of these DNAs were inadvertently mixed, how might they be separated from one another?arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Solving the Sequence of an Oligopeptide From Sequence Analysis Data Amino acid analysis of a decapeptide revealed the presence of the following products: The following facts were observed: Neither car boxy peptidase A nor B treatment of the- decapeptide had any effect. Trypsin treatment yielded two tetrapcptides and free Lys. Clostripain treatment yielded a tetrapcptide and a hexapeptidc. Cyanogen bromide treatment yielded an octapeptide and a dipeptide of sequence NP (using the one-letter codes). Chymotrypsin treatment yielded two tripeptides and a telrapeptide. The N-terminal chymotryptic peptide had a net charge of — 1 at neutral pi I and a net charge of —3 al pH 12. One cycle of Ed man degradation gave the PTH derivative What is the ammo acid sequence of this decapeptide?arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Consider the following peptide sequences: EANQIDEMLYNVQCS LTTLE DTVPW LG VHLDITVPL SWTWTLYVKL QQNWGGLWILTLVWFLM CNMKHGDSQCDERTYP YTREQSDGHIPKMNCDS AGPFGPDGPTIGPK Which of the preceding sequences would be likely to be found in each of the following: A parallel -sheet An antiparallel -sheet A tropocollagen molecule The helical portions of a protein found in your hairarrow_forward
- Answers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. CRISPR/Cas9: Design of a gRNA to Target the Human PVALB Gene The human PVALB gene, which encodes the Ca2+-binding protein parvalbumin, can be Targeted by CRISPR/Cas9, at the protospacer sequence - ATGCAGGAGGGTGGCGAGAGGGGCCGAGAT- followed by a -TGG-PAM trinucleotide. Give the sequence of the spacer region of a gRNA that will target the complementary DNA strand at this site. Include at the 3'-end of your gRNA sequence a region that will form a stem-loop structure with a 5'-AGCAUAGCUGUAAAAC- sequence downstream in the gRNA to create the dsRNA-binding site for Cas9.arrow_forwardAnswers to all problems are at the end οΓthis book. Detailed solutions are available in the Student Solutions Manual. Study Guide, and Problems Book. Superbug infections are becoming more common around the world. Many of these infections arise from the action of -lactamases, of which there are several types with different mechanisms of action. Consult the end-of-chapter reference by von Nussbaum and Schiffer and write detailed mechanisms for the serine -lactamases and metallo- -lactamases.arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Deducing DNA Sequence from Sanger Sequencing Results The output of an automated DNA sequence determination by the Sanger dideoxy chain termination method, performed as illustrated in Figure 11.3, is disp1ayed at right. What is the sequence of the original oligonucleotide?arrow_forward
- Answers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Designing Primers for PCR Amplification of a DNA Sequence Given the following short DNA duplex of sequence (53)ATGCCGTAGTCGATCATTACGATAGCATAGCACAGGGATCCA- CATGCACACACATGACATAGGACAGATAGCAT what oligonucleotide primers (17-mers) would be required for PCR amplification of this duplex?arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Structural complementarity is the key to molecular recognition, a lesson learned in Chapter 1. The principle of structural complementarity is relevant to answering problems 5, 6, 7,11, 12, and 19. The quintessential example of structural complementarity in all of biology is the DNA double helix. What features of the DNA double helix exemplify structural complementarity?arrow_forwardAnswers to all problems are at the end of this book. Detailed solutions are available in the Student Solutions Manual, Study Guide, and Problems Book. Evaluation of -Helices in Proteins The hem agglutinin protein in influenza virus contains a remarkably long -helix, with 53 residues. How long is this -helix (in nm)? How many turns does this helix have? The typical residue in an -helix is involved in two H bonds. How many H bonds are present in this helix?arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
