
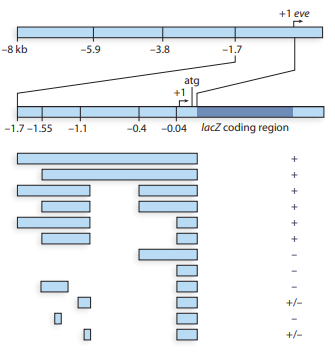
The Drosophila even
Want to see the full answer?
Check out a sample textbook solution
Chapter 14 Solutions
Genetic Analysis: An Integrated Approach Plus Mastering Genetics with Pearson eText -- Access Card Package (3rd Edition) (What's New in Genetics)
- ennar region of gene X, which determines the length of the tail in mice, is mutated so that transcription factors bind it at a much higher affinity compared to the wild-type sequence. What is the most likely phenotypic outcome? Tail length will not change because the enhancer is a non-coding sequence Tail length will increased due to increased activity of the gene's promoter Tail length will decreased because any mutation will cause a loss-of-function of these regulatory regions Not just the tail will be enlarged because increased activity of the enhancer will impact many genesarrow_forwardThe MAT locus allows yeast to switch mating type through a very complex mechanism. However, it has informed us a great deal about what aspects of gene expression typical to all organisms? Options: higher order changes in chromatin affect transcriptional efficiency that general transcription factors must first bind directly to histone tails and only then can they interact with their cognate binding sites that DNA methylation is involved in this silencing mechanism that SIR2 is required for all types of transcriptional repression that expression of Pol III genes provides a means of identifying active chromatinarrow_forwardThe UG4 gene is expressed in the stem and leaf tissue of the plant Arabidopsis thaliana. To identify potential mechanisms regulating UG4 gene expression, six small deletion mutations are made in cloned sequence containing the upstream regulatory region. The full length mutant segments Which mutation(s) affect an enhancer? Why? Which mutations identify the promoter? Why? Speculate about the reason for different transcription rated obtained for fusion constructs E and F.arrow_forward
- Name the lambda promoters whose expression is regulated by the cro protein. For each promoter you named, is cro an activator or a repressor of transcription from that promoter?arrow_forwardIn the sea urchin, early development may occur even in the presence of actinomycin D, which inhibits RNA synthesis. However, if actinomycin D is present early in development but is removed a few hours later, all development stops. In fact, if actinomycin D is present only between the sixth and eleventh hours of development, events that normally occur at the fifteenth hour are arrested. What conclusions can be drawn concerning the role of gene transcription between hours 6 and 15?arrow_forwardThe IMD2 promoter contains three upstream transcription start sites (TSS) that are utilized under high GTP conditions and a single downstream TSS (-106) that is normally only utilized under low GTP conditions. In a wild type cell, expression of IMD2 mRNA only occurs if transcription initiates from the -106 TSS. In 300 words or less, describe: 1.) The normal function of Ssl2, and 2.) why a mutation in Ssl2, that increases its catalytic rate, would allow expression of the IMD2 ORF under high GTP conditions. (Conditions under which the IMD2 ORF is NOT expressed in the wild type.)arrow_forward
- Gal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forwardGR and PPAR are transcription factors that bind to GRE and PPARE sequences respectively and activate transcription of genes. A reporter cell line is created in which the the green fluoresecent protein (GFP) is controlled by a GRE sequence and the pink fluorescent protein mCherry is under control of a PPARE sequence. If the gene for GR is introduced into the reporter cell line, the cells produce a green color. Chimeric proteins are created in which the DNA Binding Domains (DBD) and Activation Domains (AD) of the transcription factors are introduced into various cell lines. Match the following cell-types with the fluorescent color(s) you would expect the cells to produce.arrow_forwardThe extracellular protein factor Decapentaplegic(Dpp) is critical for proper wing development in Drosoph-ila (Figure Q21–3A). It is normally expressed in a narrowstripe in the middle of the wing, along the anterior–pos-terior boundary. Flies that are defective for Dpp formstunted “wings” (Figure Q21–3B). If an additional copyof the gene is placed under control of a promoter that isactive in the anterior part of the wing, or in the posterior part of the wing, a large mass of wing tissue composed ofnormal-looking cells is produced at the site of Dpp expres-sion (Figure Q21–3C and D). Does Dpp stimulate cell divi-sion, cell growth, or both? How can you tell?arrow_forward
- Discuss the morphological differences between the parasegments and segments of Drosophila. Discuss the evidence, providing specific examples, that suggests the parasegments of the embryo are the subdivisions for the organization of gene expression.arrow_forward1) Histone methylation can have many different effects on gene expression. In some cases, histone methylation is associated with activation of transcription, whereas in other cases it can trigger the formation of heterochromatin and a decrease in transcription. If histone methylation has been detected in the region of gene YFG in yeast, describe an experiment that could distinguish whether the methylation is important to activate or repress transcription of gene YFG. 2) An E. coli strain of chromosomal genotype lacI lacP lacO lacZ lacY . You wish to transform this strain into a wild-type lac operon by the addition of an extra piece of DNA (plasmid). What regulatory region(s), gene or genes would you add on this extra DNA that would make this strain express the lac operon genes as wild type? How would you design this plasmid if you want the strain of E. coli to express the lac operon genes even if lactose is absent in the medium (constitutively)?arrow_forwardA protein or RNA that regulates gene expressionin trans—either at the level of transcription, RNAsplicing, or translation—must have specificity for onetarget gene or a group of target genes. Explain howspecificity is achieved in each casearrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





