
Genetic Analysis: An Integrated Approach (2nd Edition)
2nd Edition
ISBN: 9780321948908
Author: Mark F. Sanders, John L. Bowman
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 17, Problem 15P
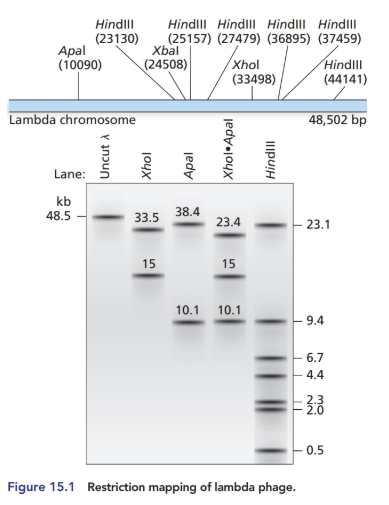
The bacteriophage lambda genome can exist in either a linear form (see Figures
How many fragments will be formed by restriction enzyme digestion with XhoI alone, with XbaI alone and with both XhoI and XbaIin the linear and circular forms of the lambda genome?
Diagram the resulting fragments as they would appear on an agarose gel after electrophoresis.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
How would you expect your results to change if the enzyme did not have time to cut all of the sites? (Hint, remember that there are many pieces of the lambda DNA in the tube – draw yourself a simplified model where there are four restriction sites, and imagine what would happen if the enzyme only cut at one, two, three or all four, randomly, each time).
How would you expect your results to change if you treated them with both EcoRI and HindIII in the same tube?
Why might it be safer for the virus to have its DNA become circular when it enters the host, rather than remain linear?
A 22-kb piece of DNA has the following restriction sites:A batch of this DNA is first fully digested by HpaI alone, then another batch is fully digested by HindIII alone, and finally, a third batch is fully digested by both HpaI and HindIII together. The fragments resulting from each of the three digestions are placed in separate wells of an agarose gel, separated by gel electrophoresis, and stained by ethidium bromide. Draw the bands as they would appear on the gel.
For Pet41 (choose Pet41 a, b, or c as provided in the image) how would you design the primers (forward and reverse) for the following gene of interest and what restriction enzymes would be used (as shown in the image)? Be sure to explain and elaborate on why selected and how.
Gene of Interest:
atgggc gacaaaggga 241 cccgagtgtt caagaaggcc agtccaaatg gaaagctcac cgtctacctg ggaaagcggg 301 actttgtgga ccacatcgac ctcgtggacc ctgtggatgg tgtggtcctg gtggatcctg 361 agtatctcaa agagcggaga gtctatgtga cgctgacctg cgccttccgc tatggccggg 421 aggacctgga tgtcctgggc ctgacctttc gcaaggacct gtttgtggcc aacgtacagt 481 cgttcccacc ggcccccgag gacaagaagc ccctgacgcg gctgcaggaa cgcctcatca 541 agaagctggg cgagcacgct taccctttca cctttgagat ccctccaaac cttccatgtt 601 ctgtgacact gcagccgggg cccgaagaca cggggaaggc ttgcggtgtg gactatgaag 661 tcaaagcctt ctgcgcggag aatttggagg agaagatcca caagcggaat tctgtgcgtc 721 tggtcatccg gaaggttcag tatgccccag agaggcctgg cccccagccc acagccgaga 781 ccaccaggca gttcctcatg tcggacaagc ccttgcacct…
Chapter 17 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
Ch. 17 - 15.1 What purpose do the bla and lacZ genes serve...Ch. 17 - The human genome is 3109 bp in length. How many...Ch. 17 - 15.3 Ligase catalyzes a reaction between the...Ch. 17 - You have constructed four different libraries: a...Ch. 17 - Using the genomic libraries in Problem 4, you wish...Ch. 17 - The human genome is 3109bp. You wish to design a...Ch. 17 - 15.7 Using animal models of human diseases can...Ch. 17 - 15.8 Compare methods for constructing homologous...Ch. 17 - 15.9 Chimeric genefusion products can be used for...Ch. 17 - 15.10 Why are diseases of the blood simpler...
Ch. 17 - Injection of double-stranded RNA can lead to gene...Ch. 17 - Compare and contrast methods for making transgenic...Ch. 17 - 15.13 It is often desirable to insert cDNAs into a...Ch. 17 - 15.14 A major advance in the s was the development...Ch. 17 - 15.15 The bacteriophage lambda genome can exist in...Ch. 17 - 15.16 The restriction enzymes Xho and Sal cut...Ch. 17 - 15.17 The bacteriophage has a single-stranded DNA...Ch. 17 - 15.20 You have identified a cDNA clone that...Ch. 17 - You have isolated a genomic clone with an EcoR I...Ch. 17 - 15.18 To further analyze the CRABS CLAW gene (see...Ch. 17 - 15.21 You have isolated another cDNA clone of the...Ch. 17 - 15.22 You have identified five genes in S....Ch. 17 - You have generated three transgenic lines of maize...Ch. 17 - 15.24 Bacterial Pseudomonas species often possess...Ch. 17 - 15.25 Two complaints about some transgenic plants...Ch. 17 - 15.26 In Drosophila, lossoffunction Ultrabithorax...Ch. 17 - Prob. 27PCh. 17 - The highlighted sequence shown below is the one...Ch. 17 - The RAS gene encodes a signaling protein that...Ch. 17 - Vitamin E is the name for a set of chemically...Ch. 17 - 15.31 You have cloned a gene for an enzyme that...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Consider the ends of the DNA fragments shown below. They have been produced by digestion of a single sequence of DNA using a number of restriction endonucleases. 1. 5'A 3' 3'TTCGA5' 2. 5'G 3' 3'CAGCT5' 3. 5'AATTC3' 3' G5 4. 5'TCGAC3' 3' G5' 5. 5'GGG 3' 3'CCC 5' Which of these ends are capable of annealing and being joined by DNA ligase?arrow_forwardA linear piece of DNA that is 14 kb long is cut first by EcoRI alone, then by SmaI alone, and finally, by both EcoRI and SmaI together. The following results are obtained: Draw a map of the EcoRI and SmaI restriction sites on this 14-kb piece of DNA, indicating the relative positions of the restriction sites and the distances between them.arrow_forwardAssume that a circular plasmid is 3200 base pairs in length and has restriction sites for HindIII restriction enzyme at the following locations: 400, 700, 1400, 2600. Give the expected sizes of the restriction fragments following complete digestion.arrow_forward
- A group of overlapping clones, designated A through F, is isolated from one region of a chromosome. Each of the clones is separately cleaved by a restriction enzyme, and the pieces are resolved by agarose gel lectrophoresis,with the results shown below. There are nine different restriction fragments in this chromosomal region, with a subset appearing in each clone. Using this information, deduce the order of the restriction fragments in the chromosome.arrow_forwardWhen linear DNA is sequenced, the nucleotide base pairs are numbered from the start to finish. a DNA molecule that is 3133 base pairs long is digested with RsaI restriction enzyme recognition sites at base numbers 366, 1534, and 2207. What are the sizes of the DNA fragments that will be produced after the DNA is digested with RsaI?arrow_forwardRefer to the diagram of pUC18 (Fig.) to determine which restriction enzymes you could use to insert a gene that would interfere with production of β-galactosidase by the host cell.arrow_forward
- Below is a map of the E. coli cloning vector after the insertion of pka-1. Ori denotes the origin of replication; amp denotes the ampicillin resistance gene. HindIII, BamHI, SalI, and EcoRI designate restriction enzyme sites. There are no other restriction enzyme sites found on this vector. The numerals denote the number of base pairs between different locations on the plasmid. For instance, there are 400 base pairs between the HindIII and BamHI site, and there are 3000 base pairs in the entire cloning vector (following the integration of pka-1). With the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following: Isolation of target DNA fragments (often referred to as inserts) Ligation of inserts into the plasmid, creating recombinant molecules Transformation of recombinant plasmids into bacteria or other suitable host for propagation Screening/selection of hosts containing the intended…arrow_forwardA linear piece of DNA that is 30 kb long is first cut with BamHI, then with HpaII, and, finally, with both BamHI and HpaII together. Fragments of the following sizes were obtained from this reaction: BamHI: 20-kb, 6-kb, and 4-kb fragments Hpall: 21-kb and 9-kb fragments BamHI and Hpall: 20-kb, 5-kb, 4-kb, and 1-kb fragments Draw a restriction map of the 30-kb piece of DNA, indicating the locations of the BamHI and HpaII restriction sitesarrow_forwardWhen the restriction endonuclease EcoRI is used to digest a 10 kb DNA fragment, it produces 4 kb and 6 kb-sized fragments. Digesting the 10 kb fragment with BamHI yields three fragments, each ranging in size from one to three and a half kilobytes. Four pieces of 0.5, 1, 3 and 5.5 kb are formed after using both enzymes. Create a restriction map for this 10 kb piece of DNA using the information you have collected. Make a note of where the two enzymes cut, as well as the distances between the enzymes.arrow_forward
- Give the recognition sequences for each of the restriction enzymes (a) PagI, (b) AluI, (c) PstI, and (d) RcaI. Show the sequence in double-stranded DNA (dsDNA) format, indicate the cleavage position with a ^, and mention which types of ends are generated in each case (blunt or sticky; 5’ or 3’ overhang)? Recognition sequence for a)PagI cleaves DNA at the recognition sequence 5' ---T ^CATGA--- 3' b) AluI cleaves DNA at the recognition sequence 5' AG^CT 3' c) PstI cleaves DNA at the recog.arrow_forwardKnowing that you are using HindIII and EcoRI to cut your plasmids, and that those two enzymes cut within the MCS, use the map of pUC19 provided below to compute: What will be the sizes of the 2 restriction fragments if NO insert is present in pUC19? What will be the sizes of the 2 restriction fragments if the approximately 317 bp RT-PCR product (insert from WT satC dimer) was ligated successfully into the SmaI site? What will be the sizes of the 2 restriction fragments if TWO approximately 317 bp RT-PCR products (2 ligated inserts from WT satC dimer) were ligated into the SmaI site?arrow_forwardWhen circular DNA is sequenced, the nucleotide base pairs are numbered starting from a fixed position on the DNA, all the way around, usually in a clockwise manner. a DNA molecule that is 3133 base pairs long is digested with RsaI restriction enzyme recognition sites at base numbers 366, 1534, and 2207. What are the sizes of the DNA fragments that will be produced after the DNA is digested with RsaI?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license