
Concept explainers
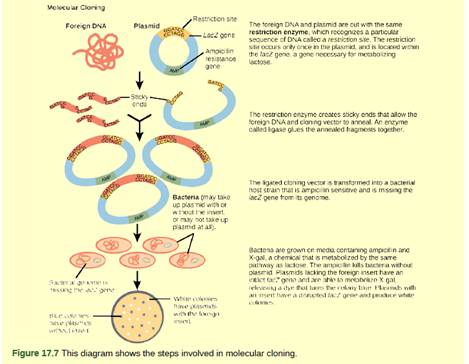
Figure 17.7 You are working in a molecular biology lab and, unbeknownst to you, your lab partner left the foreign genomic DNA that you are planning to clone on the lab bench overnight instead of storing it in the freezer. As a result, it was degraded by nucleases, but still used in the experiment. The plasmid, on the other hand, is fine. What results would you expect from your molecular cloning experiment?
- There will be no colonies on the bacterial plate.
Introduction:
Molecular cloning is a process to reproduce desired fragments of the genome. In cloning the plasmid molecules can be used to provide a folder in which desired genomic fragment is inserted. Then plasmids are introduced into a bacterial host. Plasmids with foreign DNA into them are called recombinant DNA molecules.
Answer to Problem 1VCQ
Correct answer:
The correct answer is option (b)- there will be blue colony only.
Explanation of Solution
Explanation for the correct answer:
Option (b) is given as ‘there will be blue colony only’. In this experiment, when the cloning vector plasmid with degraded foreign DNA is transformed into a bacterial host strain, releases a dye which turns the colony blue. Therefore, option (b) is correct.
Explanation for incorrect answers:
Option (a) is given as ‘there will be no colonies on the bacterial plate’. The colonies will be present on the plate. So, it is a wrong answer.
Option (c) is given as ‘there will be blue and white colonies no colonies’. Both blue and white colonies are only possible when plasmids are with foreign DNA insert and without insert.
So, it is a wrong answer.
Option (d) given as ‘there will be white colony only’. White colonies are not possible as plasmids with insert are not capable to express their characters. So, it is a wrong answer.
Hence, options (a ), (c), and (d) are incorrect.
The molecular cloning is a process to reproduce the desired fragments of the genome is reproduced by molecular cloning with the help of plasmid which provides a folder in which desired genomic fragment is inserted. Hence, the correct answer is option (b).
Want to see more full solutions like this?
Chapter 17 Solutions
EBK BIOLOGY
Additional Science Textbook Solutions
College Physics
Biological Science
Human Physiology: An Integrated Approach (8th Edition)
Anatomy & Physiology (6th Edition)
Concepts of Genetics (11th Edition)
Essentials of Genetics (9th Edition) - Standalone book
- Cloning Genes Is a Multistep Process In cloning human DNA, why is it necessary to insert the DNA into a vector such as a bacterial plasmid?arrow_forwardYou would like to amplify the gene shown below using PCR. What is the sequence of the primer you would generate for the left side of the sequence? a) 5’ GCGGTTAA 3’ b) 5’ CGCCAATT 3’ c) 5’ TTAACCGC 3’ d) 5’ AATTGGCG 3’arrow_forwardYou want to clone a 6,000 bp DNA fragment in E. coli. Which cloning vectors would be appropriate? How will you select transformants?arrow_forward
- In relation to the use of restriction enzymes in recombinant DNA technology, answer the following: You have accidentally torn the labels off two tubes (tube A and tube B), each containing a different plasmid, now you do not know which plasmid is in which tube. Fortunately, you have restriction maps for both plasmids, shown in Figure below. You have the opportunity to test just one sample from one of your tubes. By utilizing agarose gel electrophoresis technique, which restriction enzyme OR combination of restriction enzymes would you use in this experiment to determine which plasmid is found in which tube?. (Hint: if you use Hind III restriction enzyme you are going to get ONE single fragment with a molecular size of → 0.5+0.3+0.2+0.4+1+1 = 3.4 kb).arrow_forwardWith the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following:a. Isolation of target DNA fragments (often referred to as inserts)b. Ligation of inserts into the plasmid, creating recombinant molecules c. Transformation of recombinant plasmids into bacteria or other suitable host for propagationd. Screening/selection of hosts containing the intended recombinant plasmid. For this stage(d), discuss the importance of a second marker that can be used for screening of genomic DNA for colonies containing the pka-1 under the principle of insertional inactivation. This should be properly explained using all the attributes of the plasmid described above.arrow_forwardIf you were doing a cloning experiment where a plasmid was treated with PstI but the insert DNA did not have a PstI restriction site, what restriction enzyme(s) would be suitable to treat the DNA to be inserted?arrow_forward
- A DNA library is a collection of clones, each containing a different fragment of DNA, inserted into a cloning vector. What is the difference between a cDNA library and a genomic DNA library?arrow_forwardUsing nucleotide letters, show the kind of cut that could be made on a DNA molecule to circularize it into a plasmid.What are restriction length polymorphisms, and how are they used?arrow_forwardOne of your colleagues in a molecular laboratory did not put proper labels to some of the vessels he used in his experiments. Because of that, you accidentally mixed the engineered plasmid DNA to a solution contained in a tube labelled with just "A". Apparently, that tube ("A") contains chromosomal genetic material, some RNA fragments, some glucose sugar molecules, some peptides, and long chain proteins from cell debris. You remember that the engineered plasmid will give you the chance to express a very important protein that can help cure Covid-19 patients. How do you get the plasmid from the mixed solution and bring it to its pure plasmid form (devoid of any unwanted materials for your downstream processes). Outline a step by step procedure on this and please make sure to include a step where you can prove that the content of the final vessel is indeed the engineered plasmid.arrow_forward
- If you performed PCR to amplify a gene region, and you began with 20 copies of double stranded genome, after 18 cycles, about how many double stranded copies of that region would you have (in theory)?arrow_forwardYou want to amplify the following sequence of DNA from a gene of interest that you are studying. To do this, you will use PCR. Design a forward and reverse primer. Show where the primers anneal to the template sequence. Template DNA: 5’ ATGACGGAATATAAGCTGGTGGTGGTGG---GGCTGCATGAGCTGCAAGTGTGTGCTCTCCTAA 3’ 3’ TACTGCCTTATATTCGACCACCACCACC---CCGACGTACTCGACGTTCACACACGAGAGGATT 5’ (PLEASE EXPLAIN STEP - BY STEP) Answer : Foward primer 5": Reverse primer 3"arrow_forwardYou have two PCR primers: Forward- 5' TGAGCTAGGC 3' and Reverse- 5' GGTTCAGTCAG 3'. Show the binding sites of the primers to their Double strand DNA template. As the primer sizes are 10 and 11 bp, just write a 30 bp double stranded DNA (making sure the 5' and 3' ends of the double strand DNA in all 4 ends) and show where in the 30 bp double stranded DNA, these two primers would bind in correct orientation.arrow_forward
 Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

