
Bio 121 Campbell Biology Truman College
17th Edition
ISBN: 9781323670637
Author: Urry, Cain
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 18, Problem 11TYU
draw it The diagram below shows five genes, including their enhancers, from the genome of a certain species. Imagine that yellow, blue, green, black, red, and purple activator proteins exist that can bind to the appropriately color-coded control elements in the enhancers of these genes.
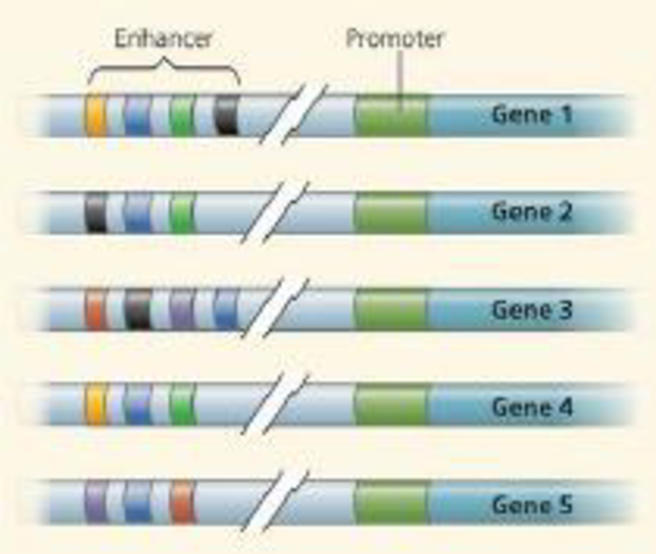
(a) Draw an X above enhancer elements (of all the genes) that would have activators bound in a cell in which only gene 5 is transcribed. Identify which colored activators would be present.
- (b) Draw a dot above all enhancer elements that would have activators bound in a cell in which the green, blue, and yellow activators are present. Identify which gene(s) would be transcribed.
- (c) Imagine that genes 1,2, and 4 code for nerve-specific proteins, and genes 3 and 5 are skin specific. Identify which activators would have to be present in each cell type to ensure transcription of the appropriate genes.
Expert Solution & Answer
Trending nowThis is a popular solution!

Students have asked these similar questions
1. - Which factor(s)/protein(s) would you expect to occur at positions C and G ? (name them)
2. - Would you expect e1, e2, or e3 to be an enhancer for gene t? (explain briefly)
3. - Which genes would you expect to be most likely regulated by enhancer e1? (explain briefly)
4. - In this example genes t - x are expressed but genes y and z are not transcribed. At which position would you expect to see a sharp transition between the chromatin marks H3K27me3 and H3K36me3? (name the position, A - I).
Which of the following statements concerning enhancer DNA sequences is true?
Group of answer choices
Enhancer DNA sequences must be located before (upstream) the genes they control.
Enhancers DNA sequences are close to the genes they activate (via looping) even if they very far away as measured through the linear DNA sequence.
Enhancer DNA sequences will not recruit their respective protein factors if you artificially insert the enhancer backwards.
All these statements are true.
The following diagram illustrates four genes from the genome of a certain insect. Different binding
sites are labeled in the enhancer region of each gene.
enhancer
promoter
ABC D E
Gene 1
A
D
Gene 2
АС
DE
Gene 3
ABD
Gene 4
In order for a specific gene to be expressed in the insect's cells, all of the gene's binding sites must
be bound by transcriptional activators.
• The insect's brain cells contain activators that bind to sites A, C, D, and E
• The insect's salivary gland cells contain activators that bind to sites A, B, and D
Which of the following is the most likely pattern of gene expression for the insect described above?
Gene 2 will be expressed in both brain and salivary gland cells
Both gene 1 and gene 4 will be expressed in salivary glands, but neither will be expressed in brain cells
Gene 1 will be expressed in both brain and salivary glands
Both gene 2 and gene 3 will be expressed in brain cells, but neither will be expressed in salivary gland cells
Chapter 18 Solutions
Bio 121 Campbell Biology Truman College
Ch. 18.1 - How does binding of the trp corepressor to the trp...Ch. 18.1 - Describe the binding of RNA Polymerase,...Ch. 18.1 - WHAT IF? A certain mutation in E. coli changes...Ch. 18.2 - In general, what are the effects of histone...Ch. 18.2 - MAKE CONNECTIONS Speculate about whether the same...Ch. 18.2 - Compare the roles of general and specific...Ch. 18.2 - Once mRNA encoding a particular protein reaches...Ch. 18.2 - WHAT IF? Suppose you compared the nucleotide...Ch. 18.3 - Compare miRNAs and siRNAs, including their...Ch. 18.3 - WH AT IF? Suppose the mRNA being degraded in...
Ch. 18.3 - MAKE CONNECTIONS Inactivation of one of the X...Ch. 18.4 - MAKE CONNECTIONS As you learned in Chapter 12,...Ch. 18.4 - MAKE CONNECTIONS Explain how the signaling...Ch. 18.4 - How do fruit fly maternal effect genes determine...Ch. 18.4 - Prob. 4CCCh. 18.5 - Prob. 1CCCh. 18.5 - Under what circumstances is cancer considered to...Ch. 18.5 - MAKE CONNECTIONS The p53 protein can activate...Ch. 18 - Compare and contrast the roles of a corepressor...Ch. 18 - Describe what must happen in a cell for a gene...Ch. 18 - Why are miRNAs called noncoding RNAs? Explsin how...Ch. 18 - Describe the two main processes that cause...Ch. 18 - Compare the usual functions of proteins encoded by...Ch. 18 - If a particular operon encodes enzymes for making...Ch. 18 - Muscle cells differ from nerve cells mainly...Ch. 18 - The functioning of enhancers is an example of (A)...Ch. 18 - Cell differentiation always involves (A)...Ch. 18 - Which of the following is an example of...Ch. 18 - What would occur if the repressor of an inducible...Ch. 18 - Absence of bicoid in mRNA from a Drosophila egg...Ch. 18 - Which of the following statements about the DNA in...Ch. 18 - Within a cell, the amount of protein made using a...Ch. 18 - Prob. 10TYUCh. 18 - draw it The diagram below shows five genes,...Ch. 18 - Prob. 12TYUCh. 18 - Prob. 13TYUCh. 18 - SCIENCE. TECHNOLOGY, AND SOCIETY Trace amounts of...Ch. 18 - WRITE ABOUT A THEME: INTERACTIONS In a Short essay...Ch. 18 - SYNTHESIZE YOUR KNOWLEDGE The flashlight fish has...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Now read this abstract from a 2013 journal article What is the authors' explanation of how Gal80p works? Note UASG from the question above is the same as UASGAL The DNA-binding transcriptional activator Gal4 and its regulators Gal80 and Gal3 constitute a galactose-responsive switch for the GAL genes of Saccharomyces cerevisiae. Gal4 binds to GAL gene UASGAL. (upstream activation sequence in GAL gene pro- moter) sites as a dimer via its N-terminal domain and activates transcription via a C-terminal transcription activation domain (AD). In the absence of galactose, a Gal80 dimer binds to a dimer of Gal4, masking the Gal4AD. Galactose triggers Gal3-Gal80 interaction to rapidly initiate Gal4-mediated transcription activation. Just how Gal3 alters Gal80 to relieve Gals0 inhibition of Gal4 has been unknown, but previous analyses of Gal80 mutants suggested a possible competition between Gal3-Gal80 and Gal80 self-association interactions. Here we assayed Gal80-Gal80 interactions and tested for…arrow_forwardYou have been analyzing wing development mutants in Drosophila and have collected the genetic/Western data below. Your epistasis analysis (not shown), suggested your current model that DW-2 is upstream of WL-1. What is a plausible mechanism for how DW-2 regulates WL-1? mutant name wingless-1 doublewing-2 Prated with Ants-Wingless MW standards || | | mutanttype ODW-2 inhibits transcription of WL-1 O DW-2 polyubiquitinates WL-1 null null phenotype Model Ac O DW-2 phosphorylates WL-1 and prevents it from entering the nucleus O DW-2 cleaves WL-1 proteolytically Wings Mode of regulation DW-2 Wings Wingsarrow_forward8. Yeast genes have cis-acting elements upstream oftheir promoters, similar to enhancers, called upstreamactivating sequences or UASs. Several target genes involved in galactose utilization are regulated by onetype of UAS called UASG, which has four bindingsites for an activator called GAL4. Two target genesregulated by UASG are GAL7 and GAL10. The GAL80 protein is an indirect repressor of GAL7 and GAL10transcription: At UASG, GAL80 binds to GAL4 protein and blocks GAL4’s activation domain. In thepresence of galactose, GAL80 no longer binds GAL4.In which gene(s) (GAL4 and/or GAL80) shouldyou be able to isolate mutations that allow the constitutive expression of the target genes GAL7 andGAL10 in the absence of galactose? In each case,what characteristics of the protein would the mutationdisrupt?arrow_forward
- a. Would you expect a cell to continue or to stopdividing at a nonpermissive high temperature if itis a temperature-sensitive Ras mutant whose protein product is fixed in the GTP-bound form atnonpermissive temperature?b. What would you expect if you had a temperaturesensitive mutant in which the Ras protein staysin the GDP-bound form at high temperature?arrow_forwardThe diagram below shows a closeup of regulatory proteins binding to one of the UASG elements near the GAL7, GALI0, and GALI genes, which code for the protein products needed for yeast to use the sugar galactose. The red triangle symbolizes an "effector" molecule that binds to Gal80p. In this hypothesis (which has since been shown to be incorrect), what could be happening to Gal80p when it is bound to the effector molecule that causes it to change its position and uncover the Gal4p transcriptional activation domain. Hint: think about what effector molecules do upon binding to proteins such as the the Lac repressor protein or the CAP protein. Galactose absent, glucose absent Gal80p. _Activation domain Gal4p dimer -Binding domain UASG Galactose present, glucose absent Activation domain Gal80p- Binding domain UASG For the toolbar, press ALT+F10 (PC) or ALT+FN+F10 (Mac).arrow_forward1. The following image shows a mechanism in which gene expression activity is regulated by ligand. Arg RS-2 Teu Met RBS Ligand RBS hidden a. What is this kind of regulatory machnism called? b. Does it involve transcription or translation? c. What happens in the presence of the ligand? d. What happens in the absence of theligand? e. What do you think the genes that are regulated here - metabolic (breakdown) or anabolic (buildup) for the ligand? Explainarrow_forward
- What would be the likely phenotypic effect of a mutation in HOTAIR and prevented it from binding to LSD1? It would be be unable to bind target genes H3K27 would not be trimethylated, leading to increased transcription of target genes H3K27 would not be trimethylated, leading to increased transcription of target genes H3K4 would not be demthylated, leading to increased transcription of target genes H3K4 would not be demthylated, leading to decreased transcription of target genes PRC2 would not be bound by HOTAIR leading to increased expression of target genes no functional effects on target genes.arrow_forwardennar region of gene X, which determines the length of the tail in mice, is mutated so that transcription factors bind it at a much higher affinity compared to the wild-type sequence. What is the most likely phenotypic outcome? Tail length will not change because the enhancer is a non-coding sequence Tail length will increased due to increased activity of the gene's promoter Tail length will decreased because any mutation will cause a loss-of-function of these regulatory regions Not just the tail will be enlarged because increased activity of the enhancer will impact many genesarrow_forwardUnderstanding why these receptors are present when they are not supposed to be is important in finding new treatments for Cushing's syndrome. When the sequences for the promoters for the receptor genes were analyzed there was no common sequences found. This led to the theory that the loss of methylation of cytosine in the promoters of these genes was leading to their inappropriate expression. How could the loss of cytosine methylation lead to overexpression of the receptor genes? O a. Increased histone acetyltransferase activity O b. Decreased histone deacetylase activity Oc Decreased histone acetyltransferase activity O d. Increased histone deacetylase activityarrow_forward
- Aurora AAurora A is a protein that acts as a kinase (transfers phosphates to molecules). Many types of cancer cells, including breast cancer cells, have higher than normal levels of this protein.Expressions of Aurora A genes in normal breast tissues (n = 10), normal tissues adjacent to tumors (n = 12) and breast tumors (n = 14).Scientists studying the production of Aurora A protein in normal frog cells observed that the amount of this protein in the cells changed throughout the cell cycle.Scientists tested chemicals that block Aurora 2 to see if they could be used as anti-cancer drugs. They found that some of the candidate drugs did slow the growth of cancer cells in cell culture in the lab. But when they tested these drugs in cancer patients to see if the drugs could slow the growth of solid tumors, they found that the benefit to patients was small when compared to the development of severe side effects such as anemia (low red blood cell count) and leukopenia (low white blood cell…arrow_forward6) The functioning of the "Ras/MAPK" signal transduction pathway is absolutely essential in order for cells to grow, divide, and migrate. One important protein that is part of this pathway is BRAF. This protein is a kind of enzyme called a "kinase" – an enzyme that transfers a phosphate group onto another protein. - In some melanomas, a mutated form of BRAF called BRAF Val600AGlu drives the progression of the cancer. The drug “vemurafenib" slows the progression of the cancer by slowing the production of the mutant BRAF protein. (National Cancer Institute. 2019. Types of Cancer Treatment. Retrieved from: https://www.cancer.gov/about-cancer/treatment/types/ Is this an example of a traditional cancer therapy or a targeted therapy? Briefly explain your reasoning in the space provided, using information provided in the text to support your answer. Type of therapy (traditional or targeted)?: Brief explanation: 19arrow_forwardDiscuss the following argument: “if the expression of every gene depends on a set of transcription regulators, then the expression of these regulators must also depend on the expression of other regulators, and their expression must depend on the expression of still other regulators, and so on. cells would therefore need an infinite number of genes, most of which would code for transcription regulators.” how does the cell get by without having to achieve the impossible?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax

Biology (MindTap Course List)
Biology
ISBN:9781337392938
Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. Berg
Publisher:Cengage Learning

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax
An Introduction to the Human Genome | HMX Genetics; Author: Harvard University;https://www.youtube.com/watch?v=jEJp7B6u_dY;License: Standard Youtube License