
Biology: The Dynamic Science (MindTap Course List)
4th Edition
ISBN: 9781305389892
Author: Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher: Cengage Learning
expand_more
expand_more
format_list_bulleted
Textbook Question
Chapter 18, Problem 1ITD
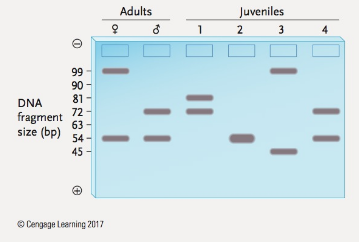
You learned in the chapter that an STR locus is a locus where alleles differ in the number of copies of a short, tandemly repeated DNA sequence. PCR is used to determine the number of alleles present, as shown by the size of the DNA fragment amplified. In the Figure below are the results of PCR analysis for STR alleles at a locus where the repeat unit length is 9 bp, and alleles are known that have 5 to 11 copies of the repeat. Given the STR alleles present in the adults, state whether each of the four juveniles could or could not be an off-spring of those two adults. Explain your answers.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
You are an expert molecular biologist and you just had your regular PCR run and you are now checking your PCR products if you have amplified the correct desired DNA fragment. Upon checking the results of the electrophoresis, you noticed that you have several bands in your gel. You expected that you will only have a band with the same size as with the "X" size of the DNA ladder. Unfortunately, you have also seen bands that fall in the "W", "Y" and "Z" markers of the ladder. Assume you got a very high quality DNA template from your extraction step.
A. What do you think are the possible causes of this incident? Explain briefly.
You are an expert molecular biologist and you just had your regular PCR run and you are now checking your PCR products if you have amplified the correct desired DNA fragment. Upon checking the results of the electrophoresis, you noticed that you have several bands in your gel. You expected that you will only have a band with the same size as with the "X" size of the DNA ladder. Unfortunately, you have also seen bands that fall in the "W", "Y" and "Z" markers of the ladder. Assume you got a very high quality DNA template from your extraction step.
B. What are the steps you will make to be able to avoid this in your next PCR run?
You have two PCR primers: Forward- 5' TGAGCTAGGC 3' and Reverse- 5' GGTTCAGTCAG 3'. Show the binding sites of the primers to their Double strand DNA template. As the primer sizes are 10 and 11 bp, just write a 30 bp double stranded DNA (making sure the 5' and 3' ends of the double strand DNA in all 4 ends) and show where in the 30 bp double stranded DNA, these two primers would bind in correct orientation.
Chapter 18 Solutions
Biology: The Dynamic Science (MindTap Course List)
Ch. 18.1 - What features do restriction enzymes have in...Ch. 18.1 - Prob. 2SBCh. 18.1 - What information and materials are needed to...Ch. 18.2 - What is a transgenic organism?Ch. 18.2 - Prob. 2SBCh. 18.3 - What is a restriction fragment length polymorphism...Ch. 18.3 - Prob. 2SBCh. 18.3 - Prob. 3SBCh. 18 - Prob. 1TYKCh. 18 - Prob. 2TYK
Ch. 18 - Why are antibiotic resistance markers such as ampR...Ch. 18 - After a polymerase chain reaction (PCR), agarose...Ch. 18 - A cDNA and a cloned fragment of genomic DNA share...Ch. 18 - Prob. 6TYKCh. 18 - Which of the following is not true of somatic cell...Ch. 18 - Prob. 8TYKCh. 18 - Prob. 9TYKCh. 18 - Prob. 10TYKCh. 18 - Prob. 11TYKCh. 18 - Discuss Concepts A forensic scientist obtained a...Ch. 18 - 13. Suppose a biotechnology company has developed...Ch. 18 - Prob. 14TYKCh. 18 - You learned in the chapter that an STR locus is a...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- A double stranded DNA molecule is shown below. The same DNA is shown below with the two strands separated. A series of potential PCR primers (labeled 1, 2, 3 and 4) that are complementary to the template DNA are shown. Which set(s) of PCR primers would allow a researcher to amplify the sequences beginning around position 10 and ending around 40? A. Primer 2 and Primer 3 B. Primer 2 and Primer 4 C. Primer 1 and Primer 4 D. Primer 1 and Primer 3arrow_forwardPCR is an exponential copying of the template strands and can be represented by the function: y = a * 2n, where a is the initial number of template copies and n is equal to the number of cycles PCR has gone through. How many DNA fragments would be produced after: 15 cycles? 13 cycles with 13 starting template strands? 29 cycles with 32 starting template strands?arrow_forwardYou would like to amplify the gene shown here using PCR. Assume the primer to the 5' end of the gene sequence shown has already been designed. You need to design a primer to hybridize with the sequence on the right side of the diagram. What will be the sequence of the primer? 5’ —GENE——AATTGGCG 3’ a) 3’ CGCCAATT 5’ b) 5’ GCGGTTAA 3’ c) 5’ CGCCAATT 3’ d) 5’ TTAACCGC 3’ e) 5’ AATTGGCG 3’arrow_forward
- Polymerase chain reaction (PCR) is a technique used to amplify (copy) DNA. Suppose a single, linear molecule of double‑stranded DNA (dsDNA) is amplified by PCR. After one PCR cycle, how many molecules of dsDNA will there be? molecules of dsDNA after one cycle: After three PCR cycles, how many molecules of dsDNA will there be? molecules of dsDNA after three cycles:arrow_forwardThe temperature at which the primers and target DNA hybridize may be changed to influence the stringency of PCR amplification. What effect will changing the hybridization temperature have on the amplification? Let's say you have a certain yeast gene A and want to check whether it has a human equivalent. How might managing the hybridization's rigor benefit you?arrow_forwardAn important feature of restriction enzymes is that each enzyme only recognizes a specific palindrome and cuts the DNA only at that specific sequence of bases. A palindromic sequence can be repeated a number of times on a strand of DNA, and the specific restriction enzyme will cut all those palindromes, no matter what species the DNA comes from. A linear DNA molecule is represented below. The DNA is represented by one line, although in actuality, DNA has two strands. If the DNA molecule has two restriction sites, specifically two repeats of a specific palindrome sequence, A and B, for a specific restriction enzyme: How many fragments would be produced if the DNA is cut by that enzyme? Number each fragment Which fragment would be the largest? Which fragment would be the smallest?arrow_forward
- The answer is option A In the PCR amplification, two primers are used to flank the target region to amplification. The two primers are forward primer and reverse primer. The forward primer binds to the template DNA and the reverse primer binds to the complementary strand. Step 2 In the above question, the two sequences of 21bp primers in the 5' to 3' Orientation that allows amplifying the given gene sequence are : ATGTTTGTTTTTCTTGTTTTA and GTCAAATTACATTACACATAA CAN YOU FURTHER EXPLAIN WHY IT IS "GTCAAATTACATTACACATAA," I don't understand why it's called reverse primer? and where exactly it is binding to on the strand? (A picture would really help)arrow_forwardWhat is expected theoretical number of copies of DNA molecules after 28 cycles in a PCR experiment? What is the percent efficiency of the PCR experiment if the actual number of copies was 500,000,000? Show your calculationsarrow_forwardYou would like to amplify the gene shown below using PCR. What is the sequence of the primer you would generate for the left side of the sequence? a) 5’ GCGGTTAA 3’ b) 5’ CGCCAATT 3’ c) 5’ TTAACCGC 3’ d) 5’ AATTGGCG 3’arrow_forward
- DNA fingerprinting involves using restriction enzymes to cut DNA at a specific sequence, resulting in many fragments of different lengths. Gel electrophoresis then separates the fragments according to size. DNA fingerprints produced from four different individuals is shown below.The DNA for Individuals 3 and 4 could NOT be Select one: a. mitochondrial DNA from two people who have the same maternal grandmother (both their mothers had the same mother) b. mitochondrial DNA from two people who have the same paternal grandmother (both their fathers had the same mother) c. nuclear DNA from identical twins d. nuclear DNA isolated from a hair left at a crime scene and a buccal swab from a suspect who was present at the crimearrow_forwardIf you had the ability to do gene editing with ONE gene for the betterment of human kind, which one would you choose, and why? Assume you could either change an abnormal allele associated with a disease, such as the cystin gene associated with Cystic Fibrosis to its normal wild type, or add a pre-existing human allele to a genome.arrow_forwardBen used the following primer pairs to amplify the COI region of his DNA template: Afterwards, he used Agarose Gel Electrophoresis to visualize the PCR products. Based on the gel: a) Was he able to successfully amplify the desired region in all replicates? b) Are there errors in Ben's gel? Does he need to repeat his PCR?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning
DNA Use In Forensic Science; Author: DeBacco University;https://www.youtube.com/watch?v=2YIG3lUP-74;License: Standard YouTube License, CC-BY
Analysing forensic evidence | The Laboratory; Author: Wellcome Collection;https://www.youtube.com/watch?v=68Y-OamcTJ8;License: Standard YouTube License, CC-BY