
Cengage Advantage Books: Biology: The Dynamic Science, Loose-leaf Version
4th Edition
ISBN: 9781305655911
Author: Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher: Brooks Cole
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 19, Problem 1ITD
Below is a sequence of 540 bases from a genome. What information would you use to find the beginnings and ends of open reading frames? How many open reading frames can you find in this sequence? Which open reading frame is likely to represent a protein- coding sequence, and why? Which are probably not functioning protein-coding sequences, and why? Note: for simplicity’s sake, analyze only this one strand of the DNA double helix, reading from left to right, so you will only be analyzing three of the six reading frames shown in Figure 19.4.
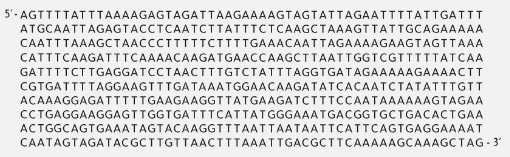
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Shown below is an R loop prepared for electron microscopy by annealing a purified eukaryotic messenger RNA with DNA from a genomic clone containing the full-length gene corresponding to the mRNA.
(a) How many exons does the gene contain? How many introns?
(b) Where in this structure would you expect to find a 5′,5′-internucleotide bond? Where would you expect to find a polyadenylic acid sequence?
Consider the following DNA sequence, which codes for a short polypeptide:
5'-ATGGGCTTAGCGTAGGTTAGT-3'
Determine the mRNA transcript of this sequence. You have to write these sequences from the 5' end to the 3' end and indicate those ends as shown in the original sequence in order to get the full mark.
How many amino acids will make up this polypeptide?
Determine the first four anticodons that will be used in order to translate this sequence.
For each of the five short mRNA nucleotide sequences given in the table below: 3. Translate the original sequence (for these short sequences start translation at the first nucleotide) 4. Identify (and highlight or underline) the one nucleotide difference between the original (left) and altered (right) sequences 5. For each altered nucleotide sequence give the type of mutation (effect at the DNA/nucleotide level; see #1 above) 6. Translate each changed sequence. Does the mutation result in a change in the amino acid sequence? If so, what is the effect of the mutation on protein structure (amino acid sequence; see #2 above)
Chapter 19 Solutions
Cengage Advantage Books: Biology: The Dynamic Science, Loose-leaf Version
Ch. 19.1 - What additional biological questions can be...Ch. 19.2 - What is the principle behind whole-genome shotgun...Ch. 19.2 - Prob. 2SBCh. 19.2 - Prob. 3SBCh. 19.2 - Prob. 4SBCh. 19.3 - Prob. 1SBCh. 19.3 - Prob. 2SBCh. 19.3 - Prob. 3SBCh. 19.4 - Prob. 1SBCh. 19.4 - Prob. 2SB
Ch. 19 - Prob. 1TYKCh. 19 - How do pseudogenes differ from genes? a. They are...Ch. 19 - Prob. 3TYKCh. 19 - Prob. 4TYKCh. 19 - Prob. 5TYKCh. 19 - Prob. 6TYKCh. 19 - About 95% of the average human transcription unit...Ch. 19 - Prob. 8TYKCh. 19 - Prob. 9TYKCh. 19 - When two protein-coding genes have very similar...Ch. 19 - Prob. 11TYKCh. 19 - Prob. 12TYKCh. 19 - Prob. 13TYKCh. 19 - Discuss Concepts The genome of the yeast...Ch. 19 - Prob. 15TYKCh. 19 - Prob. 16TYKCh. 19 - Prob. 17TYKCh. 19 - Below is a sequence of 540 bases from a genome....
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- We have talked about several examples of cis-acting elements that have dyad symmetry (inverted repeat symmetry). Some function on the level of DNA, and others function on the level of RNA. Give one example of one that functions at the DNA level and briefly explain why the sequence requires dyad symmetry to work properly. Note: you don't have to give an exact sequence, just the name of the element. Edit View Incort Format Tools Tabloarrow_forwardWhat is the length in AA’s of the LilP protein? Assume fMet is NOT CLEAVED. Enter just the number, nothing else! Write out the sequence of the polypeptide in AA: use the three letter notation, e.g. Met-Ser-Pro- A lilP mutant called lilPXS is isolated that produces a truncated polypeptide of only 6 AA in length. Describe a single basepair DNA change that would lead to this truncated version of the protein. Multiple options are possible (100 words max.)arrow_forwardRefer to the DNA sequence provided: 3’ -TACTGAAGCGGCAGCCCCGCATGAGTAGACCTTACT-5’ a. What is the mRNA transcript of the anticoding strand of the DNA model? b. What is the amino acid sequence of the polypeptide chain that will be translated from the mRNA in (a)?arrow_forward
- The practical limit for the number of different RNA sequences screened in a SELEX experiment is 10¹5. Suppose you are working with oligonucleotides 36 nucleotides long. How many sequences exist in a randomized pool containing every sequence possible? total number of sequences: What percentage of these can a SELEX experiment screen? percentage of RNA molecules screened in SELEX experiment: Suppose you wish to select an RNA molecule that catalyzes the hydrolysis of a particular ester. From what you know about catalysis, which SELEX strategy might allow you to select the appropriate catalyst? %arrow_forwardBelow is a sequence of DNA. 5'-ttaccgataattctctctcccctcttccatgattctgattaaagaaggcgagaacgaaactatttgttaatacc-3' Using the one letter code for Amino Acids, what is the predicted AA sequence of the shortest ORF (from N to C-terminal end)? Using the one letter code for Amino Acids, what is the predicted AA sequence of the longest ORF (from N to C-terminal end)?arrow_forwardKnowing that the genetic code is almost universal, a scientist uses molecular biological methods to insert the human - globin gene (shown in the figure below (Links to an external site.)) into bacterial cells, hoping the cells will express it and synthesize functional - globin protein. Instead, the protein produced is nonfunctional and is found to contain many fewer amino acids than does -globin made by a eukaryotic cell. Explain why and give thoughts as to how to overcome this.arrow_forward
- Examine the following sashimi plot from a transcriptomics experiment. The red peaks mostly RNA STAR on data 22; data 16; ar 34 2 1 545326 550237 555148 560059 O a. correspond to exons and represent respective coverage of exons O b. correspond to introns and represent respective coverage of intronsarrow_forwardRefer to the DNA sequence provided:3’ -TACTGAAGCGGCAGCCCCGCATGAGTAGACCTTACT-5’ b. What is the amino acid sequence of the polypeptide chain that will be translated from the mRNAin (a)? (The question for A is this: What is the mRNA transcript of the anticoding strand of the DNA model?)arrow_forwardWhen the cDNA was sequenced by the Sanger method utilizing ddCTP, the following products were obtained: Tetranucleotide Hexanucleotide Nonanucleotide Decanucleotide Dodenucleotide Octadecanucleotide Nonadecanucleotide 21-nucleotide 6c. What is the sequence of the bases in the mRNA coding for the peptide above? Thearrow_forward
- A protein has the following amino acid sequence: Met-Tyr-Asn-Val-Arg-Val-Tyr-Lys-Ala-Lys-Trp-Leu-Ile-His-Thr-Pro You wish to make a set of probes to screen a cDNA library for the sequence that encodes this protein. Your probes should be at least 18 nucleotides in length. Q. How many different probes must be synthesized to be certain that you will find the cDNA sequence that specifies the protein?arrow_forwardA protein has the following amino acid sequence: Met-Tyr-Asn-Val-Arg-Val-Tyr-Lys-Ala-Lys-Trp-Leu-Ile-His-Thr-Pro You wish to make a set of probes to screen a cDNA library for the sequence that encodes this protein. Your probes should be at least 18 nucleotides in length. Q. Which amino acids in the protein should be used to construct the probes so that the least degeneracy results?arrow_forwardA 2500 bp region of the human genome encodes two genes. One of the genes encodes a protein of 600 amino acids and the other gene encodes a protein of 280 amino acids. The mRNA sequences of the two genes do not contain any of the same nucleotide sequences (i.e. they do not overlap). How is this possible? Fully explain your answer.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning
Genome Annotation, Sequence Conventions and Reading Frames; Author: Loren Launen;https://www.youtube.com/watch?v=MWvYgGyqVys;License: Standard Youtube License