
Concept explainers
MAKE CONNECTIONS Ø Assign each DNA segment at the top of Figure 18.8 to a sector in the pie chart in Figure 21.6.
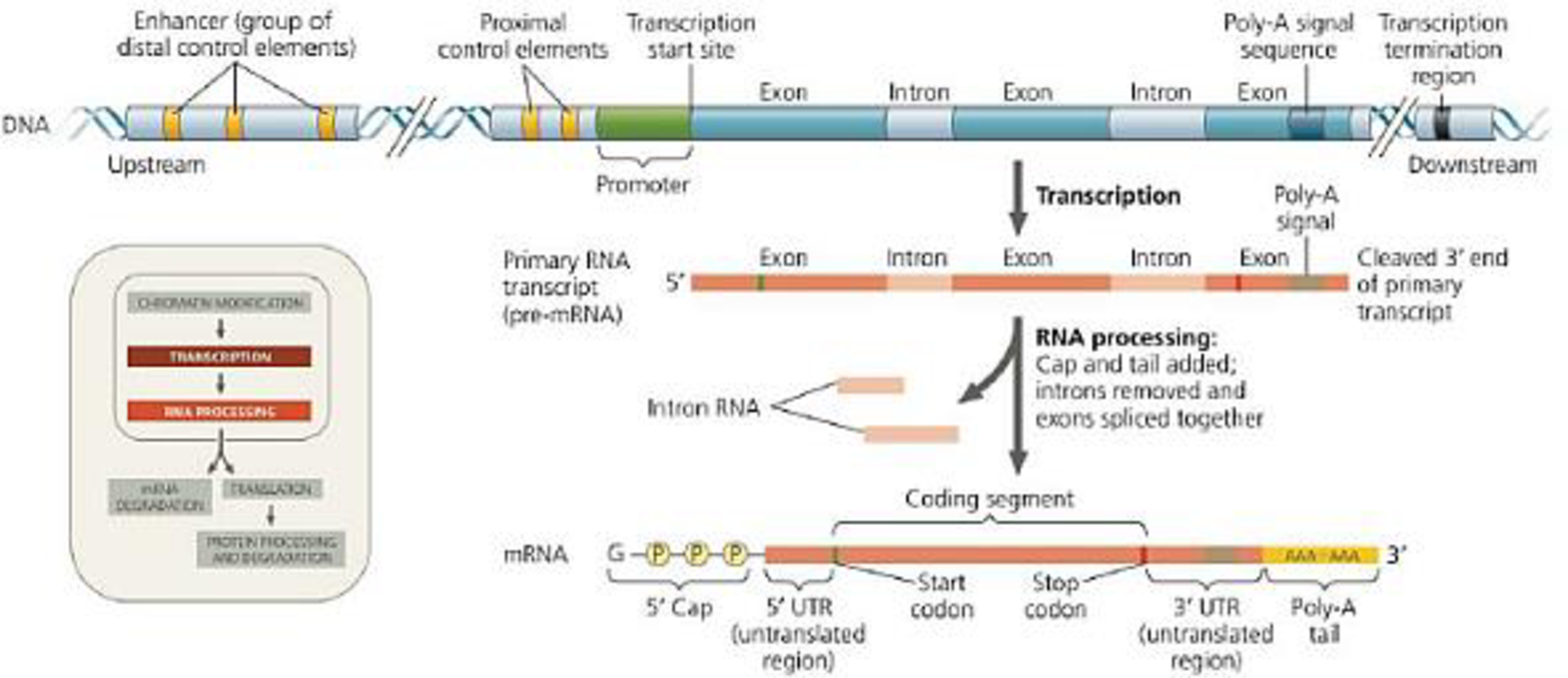
Ú Figure 18.8 A eukaryotic gene and its transcript. Each eukaryotic gene has (distal to) the promoter. Distal control elements can be grouped together as enhancers, one of
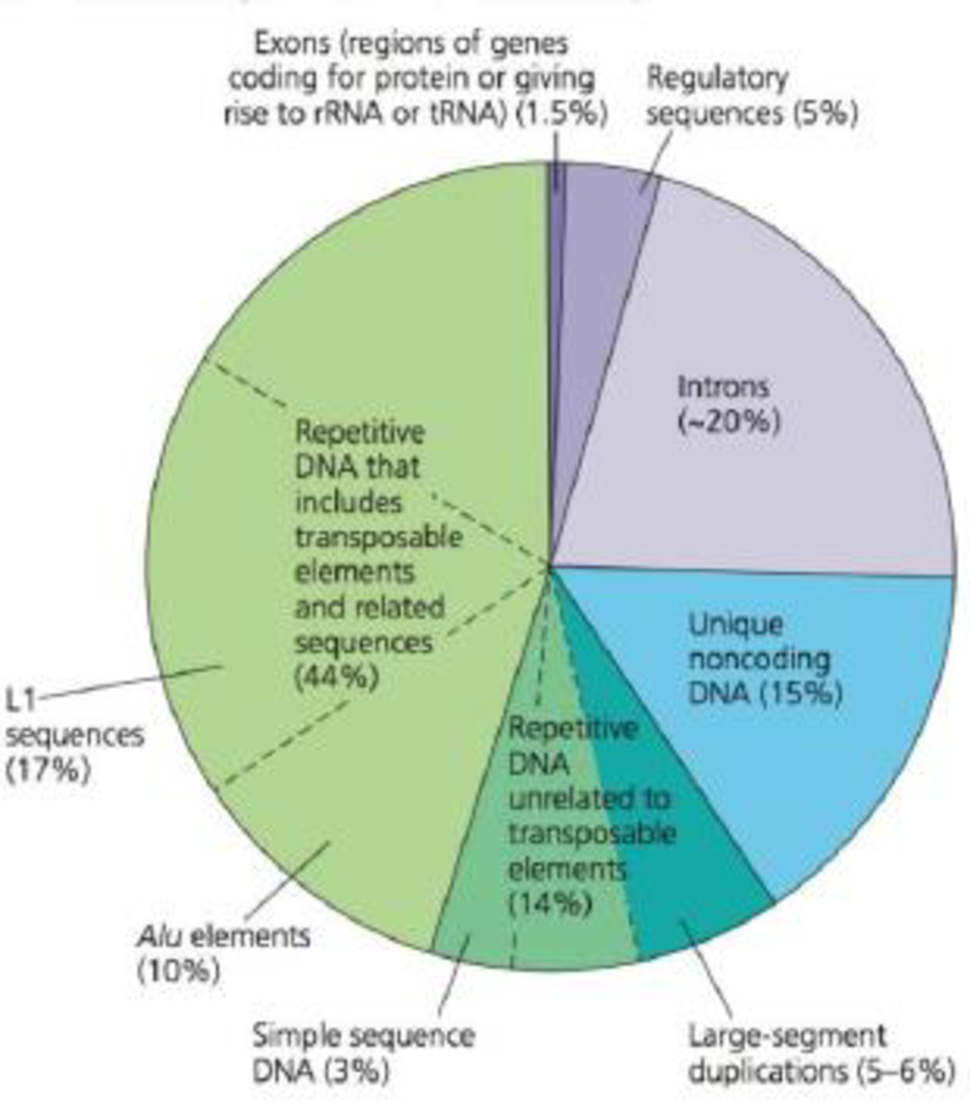
Ú Figure 21.6 Types of DNA sequences in the human genome.
The gene sequences that code for proteins or are transcribed into rRNA or tRNA molecules make up only about 1.5% of the human genome (dark purple in the pie chart). while introns and regulatory sequences associated with genes (light purple) make up about a quarter. The vast majority of the human genome does not code for proteins (although much of it gives rise to RNAs), and a large amount is repetitive DNA (dark and light green and teal).
Want to see the full answer?
Check out a sample textbook solution
Chapter 21 Solutions
CAMPBELL BIOLOGY RUTGERS 115 >C<
Additional Science Textbook Solutions
MARINE BIOLOGY
Campbell Essential Biology (7th Edition)
Human Physiology
Prescott's Microbiology
Anatomy & Physiology
Human Anatomy & Physiology (2nd Edition)
- Q11. Ribosomes are “ribonucleoprotein particles” in that they are composed mostly of rRNA with some associated ribosomal proteins. How are the genes coding for ribosomal RNAs the same as the genes coding for ribosomal proteins? They both have a transcriptional start site. They both have similar open reading frames to facilitate binding. They both have a transcriptional terminator. They both suffer frameshift mutations with the insertion of 2 nucleotides. A. 1, 2 and 3 B. 1 and 3 C. 2 and 4 D. 4 only E. All of 1, 2, 3 and 4 are correctarrow_forwardConsider this short mRNA: 5’ – AUGGCAGUGCAA – 3’. Answer the following questions assuming the code is non-overlapping. How many codons are represented in this oligonucleotide? If the second G were changed to a C, what would be the resulting amino acid?arrow_forwardQ1: How many codons code for isoleucine? For tryptophan? For leucine? Q2: What codons are associated with asparagine? With serine? Q3: From the partial mRNA sequence that you specified in Figure 10.6’s question 3 as being transcribed from the DNA template strand, remove only the first A. What amino acid sequence would be translated as a result of this change? How does that sequence compare to the amino acid sequence you translated from the original mRNA sequence?arrow_forward
- mRNA: 5’ – UGAUCAUGAUCUCGUAAGAUAUC – 3’ -Draw a box around the sequence where protein synthesis will begin. What is this sequence called? Does an amino acid get inserted at this site? If so, which one? -Draw in an arrow to show the direction that a ribosome will move along the mRNA strand. -From the starting point, mark off the codons, and identify the correct amino acid that will be inserted at that codon. -Draw a second box around the sequence where protein synthesis will stop. What is this sequence called? -Label the N-terminus and C-terminus of the polypeptide (amino acid) chain.arrow_forward-If the first G get deleted what kind of mutation will happen? Show the change in amino acid sequence. deletion lead to change in reading frame (triplet grouping) of the mRNA, so that nucleotides are grouped into different codons, lead to significant changes in amino acid sequence downstream of the mutation. TACCTA-CACACATGTAGGTGGGCAAAGTT -Multiple mechanisms regulate gene expression in eukaryotes. What are the mechanisms that control the gene expression in nucleus and what are the mechanisms that control the gene expression in cytoplasm. (names not definitions)arrow_forwardMAKE CONNECTIONS The restriction site for an enzymecalled PvuI is the following sequence:5’-CGATCG-3’3’-GCTAGC-5’arrow_forward
- 17) Which mRNA modification is likely absent if the mRNA is degrading prematurely from the 5’ end of the mRNA? A) addition of the 3’ polyadenylated tail B( splicing together if exons C removal of introns D RNA editing E addition of the 7- methlgguanosine cap to the 5’ end arrow_forwardPart1: Using the DNA nucleotide sequences for the wild-type and mutant genes in the following tables, determine the complementary mRNA sequence for the five portions of the Mc1r gene provided. (Note: You are only transcribing short portions of the DNA sequence for this protein. The actual gene contains 954 base pairs.) Part2: Using completed the mRNA sequence, determine the resulting amino acid sequence of the MC1R protein. (Note: You are translating only a portion of protein. The full protein is 317 amino acids long. The numbers above the columns in the tables indicate amino acid positions in the protein sequence.) You may use the genetic code chart providedarrow_forwardQ1: Why is only one strand of DNA used as a template? Q2: If a mutation occurred within the promoter or terminator region, do you think it would affect the mRNA transcribed? Why or why not? Q3: The template strand of part of a gene has the base sequence TGAGAAGACCAGGGTTGT. What is the sequence of RNA transcribed from this DNA, assuming that RNA polymerase travels from left to right on this strand?arrow_forward
- Select all that is true about transcription in eukaryotes. 1.Introns get spliced out of pre-mRNA 2.Transcription occurs simultaneously and in the same place as translation 3.General transcription factors assist in transcription 4.Poly A tail and a 5' 7-methyl guanosine cap are added for transcript stability 5.The sigma factor positions the RNA polymerase holoenzyme at the promoterarrow_forward15. differntial splicing allows a single gene to produce multiple proteins true or flasevarrow_forwardProject: You want to make a cat that glows in the dark (its nose, ears, and tail should glow). Choose the best answer. 9) To get started on this project, you isolate and cut out the gene in a jellyfish that codes for the green fluorescent protein (GFP). The next thing you need to do is attach this gene to the correct promoter so that you can be sure it is expressed appropriately in the cat. Which of the following promoters should you use? a) any promoter is fine b) the original promoter found in the jellyfish c) a promoter from a gene that is expressed in cells of the cat's nose, ears, and tail 10) You are now ready to transfer the piece of recombinant DNA you have prepared to the cat. You want to be sure that the transgenic cat you create will be able to pass the fluorescence on to its offspring. What is the best type of cell to transfer the recombinant DNA to? a) an unfertilized egg b) a fertilized egg c) somatic cells of an adult catarrow_forward
 Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning

