
Concept explainers
The gel electrophoresis pattern in Figure 4.23 was determined by soaking the gel in a solution of ethidium bromide (EtBr). This is a fluorescent molecule with a planar structure:
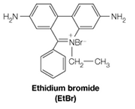
The flat molecule intercalates, or fits directly between two adjacent base pairs in a double helix. In doing so, it unwinds the double helix by 26o for each ethidium molecule bound.
a. If EtBr was added to relaxed, closed circular DNA, would you expect positive or negative supercoiling to occur? Explain.
b. If the circular DNA were nicked (had a single-stranded break) on one strand, what would be the effect on supercoiling?
c. If negatively supercoiled DNA is titrated with EtBr, the electrophoretic mobility decreases at first but then increases at higher EtBr concentrations. Explain.

Want to see the full answer?
Check out a sample textbook solution
Chapter 4 Solutions
Pearson eText for Biochemistry: Concepts and Connections -- Instant Access (Pearson+)
- To answer the prompts below, you will need to draw the chemical structure of the trinucleotide 5' - ACG - 3', labeling the 5' and 3' ends. Opposite this structure, draw the complementary trinucleotide to make a double-stranded DNA molecule. A. What is the complementary trinucleotide sequence from 5' to 3' (enter answer as e.g. CGA)? TCG B. How many non-covalent hydrogen bonds stabilize this structure? 8 C. How many covalent phosphate linkages stabilize this structure? 10 D. Which type of bond takes less energy to break? Phosphate linkage O Non-covalent hydrogen bondsarrow_forwardA closed circular duplex DNA has a 100-bp segment of alternating C and G residues. On transfer to a high salt solution, the segment undergoes a transition from the B conformation to the Z conformation. What is the change in its linking number, writhing number, and twist?arrow_forwardA circular, double-stranded DNA contains 2100 base pairs. The solution conditions are such that DNA has 10.5 bp/turn. (a) What is Lo for this DNA? (b) The DNA is found to have 12 left-handed superhelical turns. What is the superhelix density o?arrow_forward
- The oligonucleotide d-ATGCCTGACT was subjected to sequencing by Sanger’s dideoxy method, and the products were analyzed by electrophoresis on a polyacrylamide. Draw a diagram of the gel banding pattern obtained.arrow_forwardGiven the sequence shown below, write the complementary DNA sequence, using the base-pairing rules, as well as the directionality of the strands: 5'- CGAGGCTAGGTTAACCTG-3'arrow_forwardWhen the helix axis of a closed circular duplex DNA of 2310 bp is constrained to lie in a plane, the DNA has a twist (T) of 207. When released, the DNA takes up its normal twist of 10.5 bp per turn. Indicate the values of the linking number (L), writhing number (W), and twist for both the constrained and unconstrained conformational states of this DNA circle. What is the superhelix density, σ = W/T, of both the constrained and unconstrained DNA circles?arrow_forward
- A-DNA has a helix diameter of 25.5 Angstrom, a pitch per turn of 25.3 Angstrom, and a rise per base pair of 2.3 Angstrom. From this, how many bases are there per turn in A-DNA?arrow_forwardBase analysis of DNA from maize (corn) shows it to have 23 mole percent cytosine (moles per 100 moles total nucleotide). What are the percentages of the other three bases?arrow_forwardCompare the melting temperature of a 1-kb segment of DNA containing20% A residues to that of a 1-kb segment containing 30% A residues under the same conditions.arrow_forward
- a) It is known that double stranded DNA is denatured at low pH. pKa values should allow the determination of whether this is due to perturbation of the hydrogen bonding in A-T and/or G-C base pairs. The table gives values for the pKas of different protonated groups in the nucleobases.Nucleobase Position & pKa A N1, 3.5 G N7, 1.6; N1, 9.2 C N3, 4.2 T N3, 9.7a) Draw the A-T and G-C base pairs. - Label the bases with the one-letter code. – - Number the atoms in the rings and label the atom that attaches to the sugar. - Mark the groups that interact in normal…arrow_forwardAssume that a poly(A) tract five base pairs long produces a 20° bend in aDNA strand. Calculate the total (net) bend produced in a DNA if the center base pairs (the third of five) of two successive (dA)5 tracts are located (a) 10 base pairs apart; (b) 15 base pairs apart. Assume 10 base pairs per turn in the DNA double helixarrow_forwardIn the Watson-Crick model for the DNA double helix (B form) the A-T and G-C base pairs share all but one of the following properties. Which is the exception? None of the proton-binding groups in the purine and pyrimidine bases is in its charged or ionized form. The plane of the base pair is roughly perpendicular to the axis of the helix in each case. The number of hydrogen bonds formed between the two bases of the base pair is the same. O The distance between the two glycosidic (base-sugar) bonds is the same in both base pairs, within a few tenths of an angstrom.arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





