
Concept explainers
To analyze:
A rII lysis mutation resulted from a point mutation is tested against numerous deletion mutations. A series of seven point mutations are mapped along the rIIA gene, and these are tested for their capability of producing wild-type recombinants when crossed with rII partial-deletion mutants. The results are listed in the table below, the “
Using this data, show the stretch and end-points of each deletions as precisely as you can.
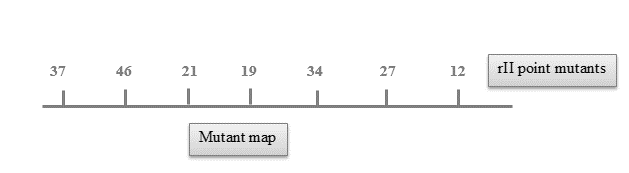
| Deletion Mutation |
Point mutation | ||||||
| 12 | 19 | 21 | 27 | 34 | 37 | 46 | |
| B622 | + | + | - | + | + | + | - |
| CT48 | - | + | + | - | - | + | + |
| MB101 | + | + | + | + | + | - | - |
| VG14 | + | - | + | + | + | + | + |
| N220 | + | - | - | + | - | + | + |
Introduction:
The recombination frequency expresses the distance among genes. Least the recombination frequency, closer the genes are while greater recombination frequencies show that the gene distance is more.
Point mutation is either deletion or addition of a
There are two types of mutants in
Want to see the full answer?
Check out a sample textbook solution
Chapter 6 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- The presence (+) or absence (−) of six sequences in each of five bacterial artificial chromosome (BAC) clones (A–E) is indicated in the following table. Using these markers, put the BAC clones in their correct order and indicate the locations of the numbered sequences within them.arrow_forwardAfter isolating six Neurospora mutations, you mate A and a representatives of each in all pairwise combinations and determine which mutations complement. Table 1gives the results, with a (+) indicating complementation, and a (-) representing lack of complementation. With the exception of mutant 1, how many different genes are represented in this experiment, or can you tell from these experiments?arrow_forwardGenomic DNA from a family where sickle-cell disease is known to be hereditary, is digested with the restriction enzyme MstII and run in a Southern Blot. The blot is hybridised with two different 0.6 kb probes, both probes (indicated in red in the diagram below) are specific for the β-globin gene (indicated as grey arrow on the diagram below). The normal wild-type βA allele contains an MstII restriction site indicated with the asterisk (*) in the diagram below; in the mutated sickle-cell βS allele this restriction site has been lost. What size bands would you expect to see on the Southern blots using probe 1 and probe 2 for an individual with sickle cell disease (have 2 βS alleles)? Probe 1 Probe 2 (a) 0.6kb 0.6kb and 1.2kb (b) 0.6kb and 1.8kb 0.6kb, 1.2kb and 1.8kb (c) 1.2kb 0.6kb (d) 1.8kb 1.8kb a. (a) b. (b) c. (c) d. (d)arrow_forward
- In each of the illustrations below, a segment of a chromosome has two copies of a transposable element. In panel a, they are oriented in the same direction, whereas in panel b they are in opposite directions. A double strand break occurs in element A and is repaired by homologous recombination using element B as a repair template. For each case, what will the chromosome look like after homologous recombination occurs? Choose one of the five options below, 1-5.arrow_forwardTo detect the CAG repeat expansion with a particular gene where 30 repeats in Normal changes to 250 repeats in a certain disease, how can we diagnose the condition. How To identify Y chromosome microdeletion ( which involves the deletion of AZF locus) using conventional karyotyping? If not then why. How will you diagnose a chromosomal translocation event?arrow_forward1) Using the diagram below, sketch in the pattern of bands you would expect to see after digesting the DNA of the TT, Tt and tt genotypes of the TAS2R38 gene. Use the Base Pair Standards on the left of the diagram as a reference in drawing the positions of the bands. Base Pair TASTER GENO TYPE Standards (Base Pairs) TT Tt tt 500 400 300 200 100 50arrow_forward
- The following table lists 4 bacterial strains that are partial diploids for lac operon genes. Given the activity of beta-galactosidase measured for each strain in the absence (-lac) or presence (+lac) of lactose, complete the table by choosing the appropriate symbol (+, -, C, S) to indicate the allele of the gene or site missing from the table (blue numbers). strain A BC 5 C D 7 chromosome I O 1 2 4 1 [Select] 9 3 [Select] [Select] [Select] 9 [Select] + + Z + + 6 + I +5 + 10 plasmid O 3 + 7 C Z + 8 8 2 [Select] 4 [Select] 6 [Select] B-gal act. -lac +lac 0.002 0.003 0.002 0.058 0.063 0.121 0.059 0.062 Select] 1 ✔ [ Select] + is C Sarrow_forwardThree mutations were obtained in a bacterial gene. An antibody is available for the protein product of this gene. Both Northern analysis (RNA separated by electrophoresis, blotted, and probed with DNA) and Western analysis (proteins separated by electrophoresis, blotted, and probed with antibodies) were performed on the mutants. The results are summarized below. Northern Size Western Size 1 2 3 1 2 3 + Long Short Short Long For each mutation, what kind of mutation occurred and how do you know? a) Mutant 1 b) Mutant 2 c) Mutant 3arrow_forwardIn the Holliday model for homologous recombination shown, the resolution steps can produce recombinant or nonrecombinantchromosomes. Explain how this can occur.arrow_forward
- Four E. coli strains of genotype a+ b- are labeled 1, 2, 3, and 4. Four strains of genotype a- b+ are labeled 5, 6, 7, and 8. The two genotypes are mixed in all possible combinations and (after incubation) are plated to determine the frequency of a+ b+ recombinants. The following results are obtained, where M = many recombinants, L = low numbers of recombinants, and 0 = no recombinants:On the basis of these results, assign a sex type (either Hfr, F+, or F-) to each strain.arrow_forwardYou have the following DNA coding sequence of a wild-type allele: 5’-ATG TTC CAG CTA GAT GAT ATG CTG GTA ATT GGG GAA CGC GCG CGG TAA-3’ For each of the following mutations: A. State whether the mutation is missense, nonsense, frameshift, or silent. B. Write the codon change that occurs for the missense, nonsense, and silent mutations (ex. GAA -- GAT). C. For frameshift mutations, write out the entire mutant sequence with each codon clearly indicated (if the frameshift creates a new stop codon, end the sequence at the new stop). Using the wild type DNA sequence above as a guide : Write the amino acid sequence of the mutants. Mutant 1: transition at nucleotide 23 Mutant 2: T --> G transversion at nucleotide 29 Mutant 3: an insertion of “A” after nucleotide 14 Mutant 4: transition at nucleotide 7 Mutant 5: An insertion of GG after nucleotide 40 Mutant 6: transition at nucleotide 15 Mutant 7: a deletion of nucleotide 25arrow_forwardYou have a patient in your clinic presenting symptoms of cystic fibrosis. You screen their CFTR gene for mutations, and find the following list: CFTR 320 L V CFTR 341 S W CFTR 528 E D CFTR 976 F Q CFTR 1235 S R Which mutation(s) are likely causing cystic fibrosis in this patient? You also sequence a newborn family member of this patient. They have all of these same mutations, other than the one at position 976, and no other mutations in CFTR. Do you predict this person will develop cystic fibrosis? Explain why.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





