
Pearson eText Genetic Analysis: An Integrated Approach -- Instant Access (Pearson+)
3rd Edition
ISBN: 9780135564172
Author: Mark Sanders, John Bowman
Publisher: PEARSON+
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 6, Problem 2P
The flow diagram identifies relationships between bacterial strains in various F factor states. For each of the four arrows in the diagram, provide a description of the events involved in the transition.
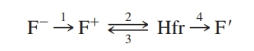
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
DNA sequencing of the entire H. influenzae genomewas completed in 1995. When DNA from the nonpathogenic strain H. influenzae Rd was compared tothat of the pathogenic b strain, eight genes of the fimbrial gene cluster (located between the purE andpepN genes) involved in adhesion of bacteria to hostcells were completely missing from the nonpathogenic strain. What effect would this deletion have oncotransformation of purE and pepN genes using DNAisolated from the nonpathogenic versus the pathogenic strain?
Given what we've discussed in class, what will be most likely outcome if
you conjugate an streptomycin resistant ampicillin sensitive methionine auxotroph E.
coli strain (engineered to be pir+) that is F- with a streptomycin sensitive non-HFR
methionine prototroph strain that is F- and RP4+ but contains pUC18?
Colonies on minimal media + ampicillin +streptomycin plates
No colonies on minimal media +ampicillin +streptomycin plates
In the Avery, McLeod, McCarty Experiment where supernatant from heat killed, virulent S Strain pneumonia solutions were added to non-virulent R Strain pneumonia cell cultures and allowed to grow in liquid media (i.e., broth). In tubes where Protease was added to the supernatant prior to cell culture, what was the observed effect when plating and growing the S. pneumonia cells to solid media?
Chapter 6 Solutions
Pearson eText Genetic Analysis: An Integrated Approach -- Instant Access (Pearson+)
Ch. 6 - For bacteria that are F+, Hfr, F', and F-, perform...Ch. 6 - The flow diagram identifies relationships between...Ch. 6 - Conjugation between an Hfr cell and an F-cell does...Ch. 6 - Bacteria transfer genes by conjugation,...Ch. 6 - Explain the importance of the following features...Ch. 6 - Prob. 6PCh. 6 - Describe what is meant by the term site-specific...Ch. 6 - What is a prophage, and how is a prophage formed?Ch. 6 - How is the frequency of cotransduction related to...Ch. 6 - Describe the differences between genetic...
Ch. 6 - Among the mechanisms of gene transfer in bacteria,...Ch. 6 - What is lateral gene transfer? How might it take...Ch. 6 - Lateral gene transfer is thought to have played a...Ch. 6 - Prob. 14PCh. 6 - A 2013 CDC report identified the practice of...Ch. 6 - Hfr strains that differ in integrated F factor...Ch. 6 - Five Hfr strains from the same bacterial species...Ch. 6 - An interrupted mating study is carried out on Hfr...Ch. 6 - An Hfr strain with the genotype cys+leu+met+strS...Ch. 6 - A triple-auxotrophic strain of E. coli having the...Ch. 6 - Penicillin was first used in the 1940 s to treat...Ch. 6 - An attribute of growth behavior of eight...Ch. 6 - Synthesis of the amino acid histidine is a...Ch. 6 - The phage P1 is used as a generalized transducing...Ch. 6 - Prob. 25PCh. 6 - Prob. 26PCh. 6 - Look closely at the consolidated Hfr map and the...Ch. 6 - Fifty bacterial colonies are on a complete-medium...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Determine the appropriate bacterial strain that can be allowed to mate with the strain that is positive for Hfr and given substrates. Also described Whether the Pyr+ Arg+ exconjugates are also Xyl+ and Mal+.arrow_forwardThe synthesis of arginine by Nuerospora was determined by examining a number of mutant strains that were unable to synthesize the compound. Use the table of bacterial growth below to 1) determine the correct sequence of the synthesis pathway and 2) where in the synthesis pathway each mutation interrupts the synthesis. A “+" indicates growth. Nothing added to Succinate Ornithine added Strain Cirtulline Arginine Added added added growth medium Wild Mutant 1 Mutant 2 Mutant 3 Mutant 4arrow_forward. a. You want to perform an interrupted-mating mappingwith an E. coli Hfr strain that is Pyr+, Met+, Xyl+,Tyr+, Arg+, His+, Mal+, and Strs. Describe anappropriate bacterial strain to be used as theother partner in this mating.b. In an Hfr × F− cross, the pyrE gene enters therecipient in 5 minutes, but at this time point thereare no exconjugants that are Met+, Xyl+, Tyr+,Arg+, His+, or Mal+. The mating is now allowed toproceed for 30 minutes and Pyr+ exconjugants areselected. Of the Pyr+ cells, 32% are Met+, 94% areXyl+, 7% are Tyr+, 59% are Arg+, 0% are His+, and71% are Mal+. What can you conclude about theorder of the genes?arrow_forward
- Describe the experiment done by Frederick Griffith in 1928, where the non-lethal (rough) strain of Streptococcus pneumoniae bacteria killed experimental mice when mixed with the heat-killed smooth strain. Discuss what may have happened to the rough strain after the heat-killed smooth strain was introduced and why you think the rough strain killed the mice. What would be the conclusion and significance of this study and its finding?arrow_forwardWhen present on the leaves of plants, the bacterium Pseudomonas syringae can promote frost damage to plants. Mutant strains, lacking the “ice” gene, have been applied to plants to try and protect the plants from wild-type P. syringae-induced frost. Assume that I am telling the truth that wild-type P. syringae nucleates ice formation at -2 °C, and provide an explanation as to why application of this altered (mutant) bacterium to plants might be a beneficial agricultural strategy in areas where morning lows occasionally dip down to 28-30 °F.arrow_forwardDescribe the differences between the F factor and Hfr transfer. Include how the donor and recipient are affected in each type of conjugation.arrow_forward
- A shuttle vector is a vector constructed so that it can propagate in two different host species. One of the most common types of shuttle vectors is the yeast shuttle vector. (1) Name the THREE (3) most common yeast shuttle vectors. (ii) Based on the answer above, differentiate these THREE (3) vectors.arrow_forwardWhat is the role of the origin of transfer during F+- and Hfr-mediated conjugation? What is the significance of the directionof transfer in Hfr-mediated conjugation?arrow_forwardFor bacteria that are F+, Hfr, F', and F- answer the following. a. Describe the state of the F factor. b. Which of these cells are donors? Which is the recipient? c. Which of these donors can convert exconjugants to a donor state? d. Which of these donors can transfer a donor gene to exconjugants? e. Describe the results of conjugation (i.e., changes in the recipient and the exconjugant) that allow detection of the state of the F factor in a donor strain. f. Describe a "partial diploid" and how it originates.arrow_forward
- The questions are on the image.arrow_forwardIn Escherichia coli, four different Hfr strains, derived from the same F* strains, were mated with F strains auxotrophic for a number of nutritional requirements (Arg Bio Cys Trp Gal His Lac Mal Xyl Leu Met). Matings were interrupted at various intervals and cells were plated on minimal medium supplemented with particular nutrients to test for gene transfer. The following results show the time of entry for each of the genes in four different Hfr strains. Table 1: Time-of-Entry Mapping Data* Hfr strains Genes lac" his" arg' bio 9. Cys gal" trp' 5 11.5 2.5 mal" xyl" leu met Hfr 1 Hfr 2 Hfr 3 Hfr 4 6.5 3.5 11 15 15 4 6. 17.5 5 14.5 3 20 * The numbers denote the number of minutes elapsed before a gene enters the F cells. Draw a circular map of E. coli chromosome.arrow_forwardA pure culture of an unknown bacterium was streaked onto plates of a variety of media. You notice that the colony morphologyis strikingly different on plates of minimal media with glucose compared to that seen on trypticase soy agar plates. How can you explain these differences in colony morphology? Also, describe what happens when a nonsense mutation is introduced into the gene encoding transposase within a transposon and why is it more likely that insertions or deletions will be more detrimental to a cell than point mutations?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning
Biology: The Dynamic Science (MindTap Course List)BiologyISBN:9781305389892Author:Peter J. Russell, Paul E. Hertz, Beverly McMillanPublisher:Cengage Learning

Biology: The Dynamic Science (MindTap Course List)
Biology
ISBN:9781305389892
Author:Peter J. Russell, Paul E. Hertz, Beverly McMillan
Publisher:Cengage Learning
genetic recombination strategies of bacteria CONJUGATION, TRANSDUCTION AND TRANSFORMATION; Author: Scientist Cindy;https://www.youtube.com/watch?v=_Va8FZJEl9A;License: Standard youtube license