
Concept explainers
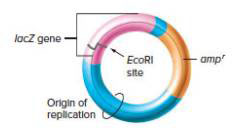
The lacZ gene from E. coli encodes the enzyme β-galactosidase, which can catalyze the conversion of a colorless compound called X-gal into a blue product. Molecular biologists have taken advantage of this property by constructing plasmid vectors that contain the lacZ gene with an EcoRI site in its middle (see figure that follows). After cutting this vector with the EcoRI enzyme, scientists ligate it together with EcoRI-digested human genomic DNA, transform the resultant molecules into ampicillin-sensitive E. coli cells, and plate these cells on petri plates containing ampicillin and X-gal. Some of the colonies growing on this plate are white in color, while others are blue. Why?
Trending nowThis is a popular solution!

Chapter 9 Solutions
Genetics: From Genes to Genomes
- You are attempting to clone a 3 kb gene from the bacterial sp Microbacterium foliarum into the EcoRI site of the 6.0 kb plasmid shown. If you restrict the plasmid with EcoR1, how many bands will you obtain on agarose gel electrophoresis? Compare it with a plasmid in which no gene has been inserted. Draw a representative agarose gel showing the bands, and the direction of current flow as well point out the positive and negative electrodesarrow_forwardAs an onion and burger lover, your friend would like to incorporate the cheems gene (“cheM”) from the rare Shiba Inu into sweet onions, in order to perfect the vegetable’s viability as a hamburger ingredient. To do so, she plans to clone the cheM gene (with an eukaryotic promoter) into the dOge plasmid. The dOge plasmid is capable of replicating in E. coli and also contains an ampicillin resistance gene, under a prokaryotic promoter, as a selective marker. Then, she plans to perform a trans-domain conjugation between E. coli (containing the plasmid with the cheM gene) and onion cells. However, you are not confident that this project will be successful, and point out some issues with the overall methodology. Identify two issues that will not allow the onion to express the cheM gene, based on the overall methodology described above.arrow_forwardBelow is a portion of an exon from a gene that encodes protein X in the genome of the plant Arabidopsis. Wildtype DNA3’ TTC AAT GCT CCG AAT ACC 5’ template strand5’ AAG TTA CGA GGC TTA TGG 3’ non-template strand A new strain (Strain B) of Arabidopsis is identified with the same region of the gene coding for protein X: 3’ TTC AAT GCT CCC AAT ACC 5’ template strand5’ AAG TTA CGA GGG TTA TGG 3’ non-template strand Compare the two DNA sequences and look for any differences. Based on what you find a. There is no mutation in Strain B compared to Strain A. b. After the point of the mutation, all the amino acids encoded by the Strain B template will be different than the Strain A protein X. c. Protein X made from the Strain B template will be much shorter than protein X made from the Strain A template d. Protein X from Strain B will have one amino acid difference that would not affect protein function. e. There is a mutation but there will not be any difference in the…arrow_forward
- If you wanted to express a library of human proteins in yeast, there are several good reasons why it would be better to use a cDNA library instead of a genomic library from humans. Which one of the following options is not such a reason? a. yeast may not initiate transcription or splicing of a human gene at the correct locations b. some genes will have very high representation (many plasmids contain the gene), while others will have very low representation (few or no plasmids contain the gene) in the cDNA library c. most genomic library clones would be useless, because only ~1.5% of the human genome actually encodes proteins d. a cDNA library can contain multiple splice variants, which are common for human genes e. introns are often very large in the human genome, making it impossible in many cases to contain a genomic version of an entire gene in a single plasmidarrow_forwardIf you use the pUC18 vector to clone in the MCS region, predict the following: a) Do bacteria that are blue in color have a cloned insert? b)Do bacteria that are white in color have a cloned insert? c) If you were to grow these cells on Chloramphenicol (an antibiotic), would the bacteria with the pUC plasmid grow? Why or Why not?arrow_forwardWith reference to the image below, discuss the process and principle involved for screening/selection of hosts (last stage of cloning) containing the intended recombinant plasmid. LacZ' = Gene for alpha-peptide of β-galactosidase.arrow_forward
- Below is a map of the E. coli cloning vector after the insertion of pka-1. Ori denotes the origin of replication; amp denotes the ampicillin resistance gene. HindIII, BamHI, SalI, and EcoRI designate restriction enzyme sites. There are no other restriction enzyme sites found on this vector. The numerals denote the number of base pairs between different locations on the plasmid. For instance, there are 400 base pairs between the HindIII and BamHI site, and there are 3000 base pairs in the entire cloning vector (following the integration of pka-1). With the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following: Isolation of target DNA fragments (often referred to as inserts) Ligation of inserts into the plasmid, creating recombinant molecules Transformation of recombinant plasmids into bacteria or other suitable host for propagation Screening/selection of hosts containing the intended…arrow_forwardA plasmid that is both ampicillin and tetracyclineresistant is cleaved with PstI, which cleaves within theampicillin resistance gene. The cut plasmid is ligated withPstI-digested Drosophila DNA to prepare a genomic library,and the mixture is used to transform E. coli K12. Question: If recombinant cells were plated on medium containingampicillin or tetracycline and medium withboth antibiotics, on which plates would you expectto see growth of bacteria containing plasmids withDrosophila DNA inserts?arrow_forwardA PCR reaction was performed to amplify the XULA3 gene, which is bp 882-5,364 on a plasmid that is 11,719 bp. After the PCR, the product was digested with XhoI. There are XhoI sites on the plasmid at bp 1,434, 4,655, and 7,368. Calculate the size(s) that would result when the product is digested with XhoI. Then enter the size of the largest fragment (in bp).arrow_forward
- When using a conventional plasmid cloning vector containing a b-galactosidase gene, it is possible to perform a "blue-white screen" to determine which bacteria have taken up a plasmid into which a DNA fragment as been inserted, as opposed to those that have taken up just reclosed plasmid vector, by growing the transformed cells on nutrient agar plates containing the artificial b-gal substrate X-gal. Will bacteria that have taken up a plasmid into which a DNA fragment has been inserted form a blue colony or a white colony when grown on this medium? Briefly explain why these bacteria would form a colony of the color you chose.arrow_forwardIn the formation of recombinant DNA, a restriction endonuclease cuts a bacterial plasmid to give sticky ends. The DNA segments that are to be added to the plasmid are cleaved with the same restriction endonuclease. What aresticky ends and why is it important that the target DNA and the plasmid it will be incorporated into have complementary sticky ends?arrow_forwardFind a plasmid map for pET11a and create a basic procedure for cloning a gene into this vector. Which selection method and substance would you use for this plasmid after transformation?arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





