
Concept explainers
Northern blot analysis is performed on cellular mRNA isolated from E. coli. The probe used in the northern blot analysis hybridizes to a portion of the lacY sequence. Below is an example of the gel from northern blot analysis for a wild
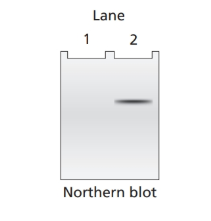
a. Lac+ bacteria with the genotype
b. Lac- bacteria with the genotype
c. Lac- bacteria with the genotype
d. Lac+ bacteria with the genotype
e. Lac- bacteria with the genotype
f. Lac- bacteria with the genotype
g. Lac- bacteria with the genotype
Want to see the full answer?
Check out a sample textbook solution
Chapter 14 Solutions
Genetic Analysis: An Integrated Approach (2nd Edition)
- Which of the following pieces of information about our recombinant DHFR protein do you predict to obtain from Western Blot analysis? A) Protein size B) Protein sequence C) Protein functionarrow_forwardIf a student uses a monoclonal antibody to detect a protein using a direct Western-blotting protocol after running an SDS PAGE, the student can detect a 50kd protein. However, if the student tries to use this antibody to immunoprecipitate the 50 kd protein and then Western -blot the immunoprecipitated samples, the student found that there is no band at all at 50kd region in the Western blot. How do you explain these results based on what you know about protein structure and antigen-antibody recognition? (Hint: Western blot analysis explores an SDS-containing buffer, whereas the immunoprecipitation buffer uses a NP40-containing buffer to maintain proteins in their native form).arrow_forwardFor Pet41 (choose Pet41 a, b, or c as provided in the image) how would you design the primers (forward and reverse) for the following gene of interest and what restriction enzymes would be used (as shown in the image)? Be sure to explain and elaborate on why selected and how. Gene of Interest: atgggc gacaaaggga 241 cccgagtgtt caagaaggcc agtccaaatg gaaagctcac cgtctacctg ggaaagcggg 301 actttgtgga ccacatcgac ctcgtggacc ctgtggatgg tgtggtcctg gtggatcctg 361 agtatctcaa agagcggaga gtctatgtga cgctgacctg cgccttccgc tatggccggg 421 aggacctgga tgtcctgggc ctgacctttc gcaaggacct gtttgtggcc aacgtacagt 481 cgttcccacc ggcccccgag gacaagaagc ccctgacgcg gctgcaggaa cgcctcatca 541 agaagctggg cgagcacgct taccctttca cctttgagat ccctccaaac cttccatgtt 601 ctgtgacact gcagccgggg cccgaagaca cggggaaggc ttgcggtgtg gactatgaag 661 tcaaagcctt ctgcgcggag aatttggagg agaagatcca caagcggaat tctgtgcgtc 721 tggtcatccg gaaggttcag tatgccccag agaggcctgg cccccagccc acagccgaga 781 ccaccaggca gttcctcatg tcggacaagc ccttgcacct…arrow_forward
- Why are restriction endonucleases considered a bacteria’s “innate immune system”? Why is CRISPR-Cas9 considered a bacteria’s “adaptive immune system”? What does CRISPR stand for? What is the difference between crRNA and tracrRNA? Why are both needed for Cas9 to function? What does PAM stand for? Where is it found? What is the difference between Non-homologous End Joining (NHEJ) and Homology Directed Repair (HDR)? What is the Guide RNA (gRNA) a chimera of? Why use a gRNA? What new things are researchers doing with CRISPR-Cas9? Reflecting on what you now know about CRISPR-Cas9, what are your thoughts on it’s use in humans and other organisms? What should we be allowed to do? Not do? Are viruses living? Why or why not? What does obligate intracellular parasite mean? What does every virus have? What is the difference between capsomers and…arrow_forwardAnalyzing Cloned Sequences A base change (A to T) is the mutational event that created the mutant sickle cell anemia allele of beta globin. This mutation destroys an MstII restriction site normally present in the beta globin gene. This difference between the normal allele and the mutant allele can be detected with Southern blotting. Using a labeled beta globin gene as a probe, what differences would you expect to see for a Southern blot of the normal beta globin gene and the mutant sickle cell gene?arrow_forwardAttached is a cartoon model of the CRISPR system, with 3 critical components labeled. A. What is the identity of each component? B. What is the function of component a? C. What is the function of component c?arrow_forward
- What are the major components of the CRISPR-Cas9 system? What mechanism does it employ to combine DNA? Explain the process of how the CRISP-Cas9 system is able to create recombinant DNA. Relate the idea of gene modification to the fields of vaccines and applied microbiology as well.arrow_forwardBelow is a map of the E. coli cloning vector after the insertion of pka-1. Ori denotes the origin of replication; amp denotes the ampicillin resistance gene. HindIII, BamHI, SalI, and EcoRI designate restriction enzyme sites. There are no other restriction enzyme sites found on this vector. The numerals denote the number of base pairs between different locations on the plasmid. For instance, there are 400 base pairs between the HindIII and BamHI site, and there are 3000 base pairs in the entire cloning vector (following the integration of pka-1). With the use of well-illustrated diagrams, reconstruct the entire cloning process by explaining different stages of the cloning process that involves the following: Isolation of target DNA fragments (often referred to as inserts) Ligation of inserts into the plasmid, creating recombinant molecules Transformation of recombinant plasmids into bacteria or other suitable host for propagation Screening/selection of hosts containing the intended…arrow_forwardWhich vectors can be used to clone a continuous fragment of DNA with the following lengths? a. 4 kb b. 20 kb c. 35 kb d. 100 kbarrow_forward
- You are examining the processing of rRNA in E. Coli using a Northern blot. Normally, a 30S rRNA transcript is transcribed and then processed (cleaved) into 23S, 16S, and 5S mature rRNAs. You suspect the gene RPO7 is involved, so you make a mutant strain of bacteria and compare it to a Wild-type (Wt) strain. You run RNA extracts from the two strains on a Northern blot and probe it with a radioactive probe that binds to all rRNA. You get the following results.arrow_forwardYou are working for a pharmaceutical company are are tasked with creating E. Coli that express an influenza antigen (a protein that is recognized by our immune system) fortreatment of the influenza virus. The cDNA of the antigen is available in a carrier vector(pCARRY) that contains a kanamycin resistance gene as shown on the left below. Inorder to get E. Coli to express the antigen, you need to clone the gene for the influenzaantigen into an expression vector that contains an E. Coli-specific promoter. You obtain such an expression vector, which contains an ampicillin resistance gene as shown on the right below (pBACTERIA) a. An end generated by digestion with BamH1 can be ligated to an end generated by digestion with BlgII. Why is this possible? b. To clone the gene for the influenza antigen from pCARRY into pBACTERIA plasmid:i. What restriction enzyme should you use to digest pCARRY? ii. What restriction enzyme should you use to digest pBACTERIA? c. After you ligate the products of…arrow_forwardYou are employed in a gene therapy laboratory to test the effectiveness of a vector for correcting the DF508 –CFTR mutation. The vector uses CRISPR –CAS9 genome editing to correct the mutated gene in situ (CFTR-CRISPR). An Ussing chamber experiment was performed using nasal airway cells obtained from a patient with the DF508 CFTR mutation which were transformed with either a control vector consisting of an inert CRISPR-CAS-9 construct (CONTROL) or the active, gene correcting CRISPR CAS-9 construct (CFTR-CRISPR). The following transepithelial voltage values were obtained for the drug regimen shown under open circuit and current clamp (injected current of 10mA) conditions: CONTROL Baseline Amiloride Forskolin Inh172 Vte (mV) -78 -2 -4 -3 DVRt (mV) 8 16 7.5 13 R (W.cm-2) 800 1600 750 1300 I (mA.cm-2) 98 1 5 2 CFTR-CRISPR Vte (mV) -38 -6 -23 -9 DVRt (mV) 9 27 6 11 R (W.cm-2) 900 2700 600…arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
