
ND STONY BROOK UNIVERSITY LOOSELEAF GENETICS: FROM GENES TO GENOMES
6th Edition
ISBN: 9781260406092
Author: HARTWELL, Leland, HOOD, Leroy, Goldberg, Michael
Publisher: Mcgraw-hill Education/stony Brook University
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 18, Problem 5P
The fly eyes shown in Fig. 18.7 are malformed because they lack a functional copy of a gene called fat facets that is required for eye development. The human genome encodes a protein called Usp9 that is similar in amino acid sequence to the fly Fat facets (Faf) protein. Likewise, the mouse genome encodes a protein called Fam that is similar to Faf and Usp9.
| a. | How could you determine if human Usp9 can substitute for the Drosophila Faf protein? |
| b. | How could you determine if, like Faf in flies, the mouse Fam protein is required for mouse eye development?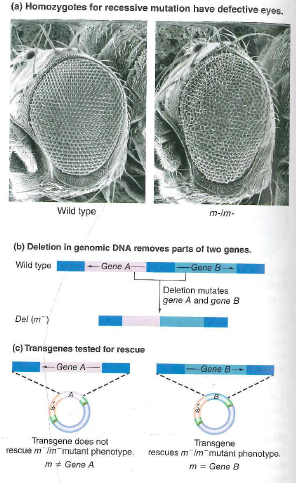 |
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
The following figure shows a screen shot from the UCSC Genome Browser, focusing on a region of the human genome encoding a gene called MFAP3L. (Note hg38 refers to version 38 of the human genome RefSeq)a. Describe in approximate terms the genomic location of MFAP3L.b. Is the gene transcribed in the direction from the centromere-to-telomere or from the telomere-to-centromere?c. How many alternative splice forms of MFAP3L mRNA are indicated by the data?d. How many different promoters for MFAP3L are suggested by the data?
(please do not copy and paste the answer from below. i don't think it is correct.
a. MFAP3L is mostly found in the nucleus in the genome. It is found on chromosome 4 reverse strand. The protein produced by the gene is found in the cell membrane, and it is positioned on the membrane with the carboxyl side of the protein facing the cytosol.
b. The MFAP3L gene is transcribed from the telomere to the centromere.
c. According to the data, there are 11 different splice forms…
You would like to add a nuclear localization sequence (NLS) of Lys-Lys-Lys-Arg-Lys to a
protein that is usually found in the cytoplasm of a yeast cell. To accomplish this, you
introduce the nucleotide sequence encoding the NLS into the gene that encodes the
cytoplasmic protein of interest.
a. What is the size of the nucleotide insert that will encode the NLS? Briefly explain.
5'
3'
b. Below is a diagram of the gene encoding the cytoplasmic protein of interest in
the yeast genome. If your goal is to put the NLS at the carboxyl (C) terminus of
the protein, at which location (A-E) should the NLS be inserted? Briefly explain.
A
TATAA
ATATT
promoter
+1
B
ATG
TAC
D
TAA
ATT
stop
codon
E
3'
5'
Lysine 4 of histone H3 (H3K4) is methylated in thenucleosomes of many transcriptionally active genes.Suppose you want to determine all the places in thehuman genome where nucleosomes contain methylated H3K4.a. Starting with an antibody that specifically bindsonly to the tails of histone H3s that have K4 methylation, what kind of experiment would you perform? Outline the major steps of this experiment.b. Do you think that you would get the same results ifyour starting material was skin cells in one experiment and blood precursor cells in a second experiment? Explain.c. Describe a follow-up experiment that could determine if your data from part (a) are consistent withthe idea that H3K4 methylation marks appear onlyat transcriptionally active genes
Chapter 18 Solutions
ND STONY BROOK UNIVERSITY LOOSELEAF GENETICS: FROM GENES TO GENOMES
Ch. 18 - Match each of the terms in the left column to the...Ch. 18 - Mice are usually gray, but a mouse geneticist has...Ch. 18 - Sometimes, genes transferred into the mouse genome...Ch. 18 - In mice, a group of so-called Hox genes encode...Ch. 18 - The fly eyes shown in Fig. 18.7 are malformed...Ch. 18 - This problem concerns a technique called enhancer...Ch. 18 - Fish and other organisms that live in the Arctic...Ch. 18 - a. Describe two ways you could potentially make a...Ch. 18 - Figure 18.6 shows a picture of Glofish ,...Ch. 18 - Some people are concerned about the possible...
Ch. 18 - The goal of the Knockout Mouse Project is to...Ch. 18 - Prob. 12PCh. 18 - Prob. 13PCh. 18 - a. Which genome manipulation technique would you...Ch. 18 - a. Diagram a knockin construct that could have...Ch. 18 - Prob. 16PCh. 18 - Prob. 17PCh. 18 - The transcription factor Pax6 is required...Ch. 18 - Mouse models for human genetic diseases are...Ch. 18 - One way to determine where inside a cell a protein...Ch. 18 - In Problem 5 in Chapter 17, you saw that a SNP...Ch. 18 - Scientists now routinely use CRISPR/Cas9 to make...Ch. 18 - Geneticists are currently considering using...Ch. 18 - a. Figures 18.9 and 18.12 demonstrated methods to...Ch. 18 - Nonhomologous end-joining NHEJ of a double-strand...Ch. 18 - One problem that researchers sometimes encounter...Ch. 18 - Researchers at the University of California at San...Ch. 18 - Prob. 28PCh. 18 - F. Port and S. Bullock at the University of...Ch. 18 - On Fig 18.14, locate the PAM site and identify the...Ch. 18 - Prob. 31PCh. 18 - Prob. 32PCh. 18 - Recall that Leber congenital amaurosis LCA, a form...Ch. 18 - One potential strategy for gene therapy to correct...Ch. 18 - Recently, scientists have used a mouse model for...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- You have the following DNA coding sequence of a wild-type allele: 5’-ATG TTC CAG CTA GAT GAT ATG CTG GTA ATT GGG GAA CGC GCG CGG TAA-3’ For each of the following mutations: A. State whether the mutation is missense, nonsense, frameshift, or silent. B. Write the codon change that occurs for the missense, nonsense, and silent mutations (ex. GAA -- GAT). C. For frameshift mutations, write out the entire mutant sequence with each codon clearly indicated (if the frameshift creates a new stop codon, end the sequence at the new stop). Using the wild type DNA sequence above as a guide : Write the amino acid sequence of the mutants. Mutant 1: transition at nucleotide 23 Mutant 2: T --> G transversion at nucleotide 29 Mutant 3: an insertion of “A” after nucleotide 14 Mutant 4: transition at nucleotide 7 Mutant 5: An insertion of GG after nucleotide 40 Mutant 6: transition at nucleotide 15 Mutant 7: a deletion of nucleotide 25arrow_forwardDuchenne muscular dystrophy is caused by a mutation in a gene that comprises 2.5 million base pairs and specifies a protein called dystrophin. However, less than 1% of the gene actually encodes the amino acids in the dystrophin protein. On the basis of what you now know about gene structure and RNA processing in eukaryotic cells, provide a possible explanation for the large size of the dystrophin gene.arrow_forwardDifferent mutations in the WDR62 gene that inactivate gene function were found in the genomes of many different people with microcephaly. This information provided strong support for the idea that the WDR62 gene mutation causes microcephaly. A.The human genome sequence identified WDR62 as one of the approximately 27, 000 genes in the human genome. What information about the function of WDR62 do you think was learned originally from the DNA sequence of the normal human genome?arrow_forward
- Drosophila females homozygous for loss-of-functionmutations in the gene aubergine are sterile. RNA-Seqexperiments show that in the ovaries of these females,the levels of RNAs for many kinds of transposable elements are more than 10× higher than in wild-type ovaries. The aubergine gene encodes a Piwi-family protein.a. Why do you think these females are sterile?b. Piwi proteins interact with piRNAs that are transcribed from piRNA gene clusters. Given that thelevels of many kinds of TEs are elevated in mutantovaries, what kinds of DNA sequences do youthink are located in these clusters?c. Many investigators think of piRNAs as a kind ofdefensive mechanism that protects organisms fromthe effects of new transposable elements that mightbe introduced into genomes, for example fromother species. Explainarrow_forwardDuring construction of a knockout mouse, a targeting vector is introduced into mouse embryonic cells, where it integrates into the genome at a ["targeted site", "random location"] by ["homologous recombination", "nonhomologous end joining "] . Pick answers within quotation marks to fill in the blanks.arrow_forwardE. coli strain BW25113 can grow on (D) Arg as the sole carbon source. A scientist working with you on this project has discovered that mice encode a protease that cancleave between (D)Arg‐(D)Arg. a. In a set of discrete steps outline how to construct this genomic library so that recombinant expression can be performed in E. coli b. Once the genomic library has been constructed, describe in DISCRETE steps how you would isolate the DNA sequence that encodes for the protease that cleaves between (D)Ag-(D)Argarrow_forward
- You are studying a large eukaryotic gene that is 439,515 base pairs long. You find the polypeptide that this gene produces in liver cells is 46,771 amino acids long. Your colleague studies the function of this gene in brain cells, and finds the polypeptide produced in the brain is much larger – 61,438 amino acids long. How do you explain this difference? Possible Answers: A. The cell cycle of liver cells is much longer than that of brain cells. B. This is due to alternative splicing. in the brain C. There was a different complement of sequence-specific transcription factor binding sites in the CRM of the brain cells. D. There is no 5' cap added to the gene product from the liver cells.arrow_forwardProteins A,B,C, and D in the diagram are encoded by different genes and interact with each other. Imagine that a mutation in the gene for protein A changes one of the charged amino acid in the red circle area from positive to negative charge (blue arrow). this mutation results in a mutant phenotype. Assume a mutation in the gene for protein B occurs and the double mutants have a phenotype that is almost wild type. How would you best describe the mutation in gene B? Protein C Proten A Protein B Protein B Wild type Mutation in the gene for protein Aarrow_forwardIn the table below, there are four versions of gene A, one of which is normal, and the other three which contain mutations that make the gene product nonfunctional. Focus on the shaded region of the sequence. Use the genetic code table to answer the question. How would you describe Mutation #2? Partial DNA sequence for gene A ("..." indicates many nucleotides of sequence not shown) 5' ... ATG GTG AGC AAG GAG GAG CTG TTC ACC TGT AAA TAG ... Normal Mutation #1 5' ... ATG GTG AGC AAG GAG AAG CTG TTC ACC TGT AAA TAG ... Mutation #2 5' ... ATG GTG AGC AAG TAG GAG CTG TTC ACC TGT AAA TAG ... Mutation #3 5' ... ATG GTG AGC AAG GAG CTG TTC ACC TGT AAA TAG ... Silent mutation Nonsense mutation Frameshift mutations Missense mútationarrow_forward
- Propose an explanation for the small amount of enzyme E produced by genotype 3.arrow_forwardResearchers have identified a gene (FR) responsible for watermelon resistance to infection by Dacus curcurbitae (a close relative of Drosophila melanogaster). They isolate RNA from resistant (FR+) and sensitive (fr-) watermelons and use a probe that will recognize both FR+ and fr- transcripts. They also isolate protein from resistant and sensitive watermelons and perform a Western blot using an antibody that can recognize the fr- and FR+ protein. Describe the results illustrated below and give a plausible molecular explanation for these observations.arrow_forwardThe b1 allele encodes a transcription factor that stimulates production of anthocyanin, a purple pigment in plants. What would be the effect of deleting the seven tandem repeats that are located 100,000 bp upstream of the b1 locus in corn?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Mitochondrial mutations; Author: Useful Genetics;https://www.youtube.com/watch?v=GvgXe-3RJeU;License: CC-BY