
To review:
The effectiveness of the enhancers associated with the dark or blond hair in driving the expression of the Kitl mRNA (messenger ribonucleic acid).
Introduction:
KITLG gene is controlled by an enhancer and it also expresses itself for hair color in humans apart from other
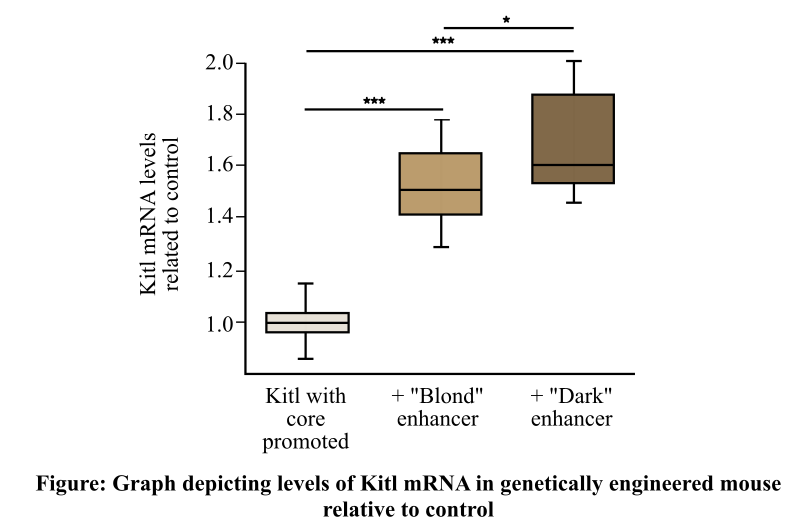
In the given experiment, the Kitl gene (mouse equivalent of KITLG gene) and its core promoter were bonded together by genetic engineering. The amount of Kitl mRNA was measured in the skin after its introduction in the embryo of a mouse. The results are shown in the graph provided below. Probability is given as *P < 0.05 and &*P **lt; 0.001.
Want to see the full answer?
Check out a sample textbook solution
Chapter 19 Solutions
Biological Science (7th Edition)
- PREDICT Compare the types of bacterial genes associated with inducible operons, those associated with repressible operons, and those that are constitutive. Predict the category into which each of the following would most likely fit: (a) a gene that codes for RNA polymerase, (b) a gene that codes for an enzyme required to break down maltose, and (c) a gene that codes for an enzyme used in the synthesis of adenine.arrow_forwardINTERPRET DATA Develop a simple hypothesis that would explain the behavior of each of the following types of mutants in E. coli. Mutant a: The map position of this mutation is in the trp operon. The mutant cells are constitutive; that is, they produce all the enzymes coded for by the trp operon, even if large amounts of tryptophan are present in the growth medium. Mutant b: The map position of this mutation is in the trp operon. The mutant cells do not produce any enzymes coded for by the trp operon under any conditions. Mutant c: The map position of this mutation is some distance from the trp operon. The mutant cells are constitutive; that is, they produce all the enzymes coded for by the trp operon, even if the growth medium contains large amounts of tryptophan.arrow_forwardExplain in detail the role of super-enhancers. Give some examples. How the super-enhancers can be identified?arrow_forward
- Based on the given scenario, tell whether structural genes are expressed (YES) or not expressed (NO) 1. LacY is deleted (YES/NO) 2. LacI is deleted (YES/NO) 3. TrpR is deleted (YES/NO) 4. Promoter sequence is deleted (YES/NO) 5. lactose is present in the environment; glucose is absent (YES/NO) 6. tryptophan is absent in the environment (YES/NO) 7. operator is deleted (YES/NO)arrow_forwardDescribe in detail the role of super-enhancers. Give some examples. How the super-enhancers can be identified?arrow_forwardQ1: As illustrated here, at what control point is transcription regulated? Q2: What is a possible advantage of regulating gene expression before transcription, versus after? Q3: If you wanted to up-regulate production of the hemagglutinin protein in a tobacco plant carrying the hemagglutinin gene, at which control point(s) would that be possible? Justify your reasoning.arrow_forward
- . What is an enhanceosome? Why could a mutation in anyone of the enhanceosome proteins severely reduce thetranscription rate?arrow_forwardFor each of the ff. scenario, state whether the gene is up- or down-regulated and briefly explain the reason behind. 5’-------enhancer------insulator-----gene of interest----3’arrow_forwardpls answer every question, no need for explanation as to why 1. In eukaryotic gene regulation, transcription is regulated by the following EXCEPT a. the binding of a repressor to a promoter.b. the binding of an activator to a promoter.c. the binding of a repressor to an enhancer gene.d. the binding of an activator to an enhancer gene. 2. The following regulatory mechanisms occur before eukaryotic transcription EXCEPT a. the formation of the transcription factor complex. b. the phosphorylation of eukaryotic initiation factors. c. the regulation of nuclear localization of transcription factors. d. the alteration of the DNA-binding domain of the transcription factor. 3. Gene expression may be regulated after translation using the following mechanisms EXCEPT a. by protein phosphorylation. b. by differential splicing. c. by microRNA binding. d. by ubiquitination. 4. If nutritional requirements are not met, gene expression will most likely be…arrow_forward
- Please describe the regulatory role of enhancers that binds to responsible regulators to activate the target gene at a given time and place through comparing different organisms according to the presence of regulatory sequencesarrow_forward. One mechanism by which antisense RNAs act as negative regulators of gene expression is by base pairingwith the ribosome binding site on the sense mRNA toblock translation. In a second, alternative mechanism,the act of transcribing an antisense RNA can somehow prevent RNA polymerase from recognizing thesense promoter for the same gene. Design an experimental approach that would enable you to distinguishbetween these two modes of action at a specific gene.(Hint: What would be the outcome in each case ifhigh levels of the antisense RNA were transcribedfrom a gene on a plasmid?)arrow_forwardDebes et al recently described how aging in humans, mice, nematodes, and other eukaryotes is associated with an increase in the average speed of transcriptional elongation by RNA polymerase II. Overexpression of some proteins that decreased PolII elongation speed extended lifespan in the fly Drosophila. For each of the following proteins, predict whether overexpression of that protein (assuming all other cellular components are normal) would likely reduce transcriptional speed, and briefly justify your answer. a) Mediator proteinsb) Histone proteinsc) Insulator binding proteinsarrow_forward
 Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
