
Pearson eText for Biochemistry: Concepts and Connections -- Instant Access (Pearson+)
2nd Edition
ISBN: 9780137533114
Author: Dean Appling, Spencer Anthony-Cahill
Publisher: PEARSON+
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 6, Problem 23P
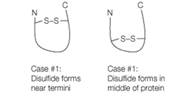
Disulfide bonds have been shown to stabilize proteins (i.e., make them less likely to unfold). Consider the cases shown schematically below for two variants of the same protein. In case #1 the disulfide forms between Cys residues that have been introduced near the protein N- and C-termini, and in case #2 the disulfide forms between Cys residues that have been introduced in the middle of the protein sequence. Which protein is likely to be more stable? (Note: Assume the disulfide bond is intact in both the unfolded and folded states). Explain your reasoning.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
You have discovered a novel protein that has a pI = 5.5. To study the functional properties of this new protein, your research group has made a mutant that contains two amino acid changes—namely, a surface Phe residue in the normal protein has been replaced by His (side chain pKa = 6.1) and asurface Gln has been replaced by Glu (side chain pKa = 6.0). Is the pI of themutant protein predicted to be greater than, less than, or the same as the pIof the normal protein? Support your answer with the appropriate calculation
You have discovered a novel protein that has a pl = 5.5. To study the functional properties of
this new protein your research group has made a mutant that contains two amino acid
changes-namely, a surface Phe residue in the normal protein has been replace by His (side
chain pk = 6.1), and a surface Gln has been replace by Glu (side chain pk. = 6.0). The pl of the
mutant protein is predicted to be:
A. Greater than the pl of the normal protein.
B. Less than the pl of the normal protein.
C. The same as the pl of the normal protein.
Consider the peptide Asp-Lys-Phe-Glu-Asn-Tyr-Gln-Val-Cys. In a single
beaker, you treat this peptide with 2 proteases. One protease cleaves at the
N-terminus of aromatic R groups and the other cleaves at the C-terminus of
polar, non-ionizable R groups.
Following the enzymatic digestion, you want to separate your peptide
fragments so that you can identify them. You choose to separate the
fragments using an anion exchange column. Beginning at pH=6 you apply
your peptide fragments to the column and you gradually decrease the pH of
the column stopping the separation when the pH of the column equals 4.
Omitting chemical structures, write the amino acid sequence of the peptide
fragments that are produced from this digest. Write the order that these
fragments will elute from the column (if at all). (Relevant pKa values are:
2.1, 3.8, 4.3, 8.3, 9.6, 10.1, and 10.5)
Chapter 6 Solutions
Pearson eText for Biochemistry: Concepts and Connections -- Instant Access (Pearson+)
Ch. 6 - Prob. 1PCh. 6 - Bovine pancreatic trypsin inhibitor (BPTI; Figure...Ch. 6 - A schematic structure of the subunit of...Ch. 6 - In the protein adenylate kinase, the C-terminal...Ch. 6 - Give two reasons to explain why a proline residue...Ch. 6 - Consider a small protein containing 101 amino acid...Ch. 6 - a. Based on a more conservative answer to Problem...Ch. 6 - The following sequence is part of a globular...Ch. 6 - a. A protein is found to be a tetramer of...Ch. 6 - Under physiological conditions, the protein...
Ch. 6 - Theoretical and experimental measurements show...Ch. 6 - The peptide hormone vasopressin is used in the...Ch. 6 - A protein gives under conditions of buffer...Ch. 6 - A protein gives a single band on SDS get...Ch. 6 - It has been postulated that the normal...Ch. 6 - Below are shown two views of the backbone...Ch. 6 - Do you expect a Pro Gly mutation in a...Ch. 6 - Rank the following in terms of predicted rates...Ch. 6 - Shown below are two cartoon views of the small...Ch. 6 - Prob. 20PCh. 6 - In most cases, mutations in the core of protein...Ch. 6 - A Leu Ala mutation at a site buried the core of...Ch. 6 - Disulfide bonds have been shown to stabilize...Ch. 6 - Cartoon renderings of the proteins Top 7 and adaH2...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- Imagine a globular protein with a histidine that is involved in a salt bridge to an aspartic acid, in an internal (hydrophobic) region of the protein. What would happen to the pKa of that histidine R group if the protein was mutated, and the aspartic acid was replaced with an alanine? (i.e., would the pKa increase, decrease, or stay the same?). Assume that the location of the histidine stays the same. Explain your answer clearlyarrow_forwardPro-CHEMBIO peptide is a large peptide sequence with three disulfide bonds (indicated by the lines between cysteine residues). CHEMBIO Protease cleaves amide bonds between two basic amino acid residues to give the final peptide that is active in the body. The products of this protease reaction are: 1. Two chains that make up active CHEMBIO peptide, which contain the N- and C-terminus sections of the original Pro-CHEMBIO molecule 2. A separate inactive peptide What if instead of CHEMBIO Protease, this peptide was cleaved by trypsin? Indicate the cleavage locations on the structure below. asaparrow_forwardA recent genome sequencing project for the bacterium Burkholderia mallei has identified a new protein with high similarity to the lysylphosphatidylglycerol flippase enzyme. A short section of the new protein sequence is shown below. TVEVNAPGDVQKALSELQQINDGRLDIRI (a) Are any reverse turns likely to be present? Explain your answer. (b) Are any beta-strands likely to be present? Explain your answer. (c) Are any alpha helices likely to be present? Explain your answer. (d) Is any supersecondary structure likely to be present? Explain your answer. (e) Identify two residues that are likely to be buried in the core of the folded protein. Explain your answer. (f) Identify two residues that are likely to be hydrogen bonded to each other. Explain your answer.arrow_forward
- Assume that the translational error frequency, d, is 1 * 10–4. (a) Calculate the probability of making a perfect protein of 100 residues. (b) Repeat for a 1000-residue protein.arrow_forward. Disulfide bonds have been shown to stabilize proteins (i.e., make them less likely to unfold). Consider the cases shown schematically below for two variants of the same protein. In case #1 the disulfide forms between Cys residues that have been introduced near the protein N- and C-termini, and in case #2 the disulfide forms between Cys residues that have been introduced in the middle of the protein sequence. Which protein is likely to be more stable? (Note: Assume the disulfide bond is intact in both the unfolded and folded states). Explain your reasoning.arrow_forwardGiven the following Wild Type and Mutated DNA sequences: 1.) Identify where the base pair change occurs (what letters changed?) 2.) For BOTH sequences, write the mRNA strands, define the codon regions (with spaces), and amino acid sequences. 3.) Describe what kind of mutation has occurred (missense, nonsense, or silent), and what effect this may have on the protein. Wild Type DNA Sequence: 3' - CCTCGTTATGTG - 5' Mutated DNA Sequence: 3' - CCTCGTTATTTG - 5'arrow_forward
- Which of the following sequences is most likely to form a beta-turn? Explain why? Sequence A: Ser-Pro-Asn-Val Sequence B: Gly-Ser-Asn-Pro Sequence C: Ser-Asn-Gly-Valarrow_forwardAssume that the translational error frequency, 8, is 1 × 10-4. (a) Calculate the probability of making a perfect protein of 100 residues. (b) Repeat for a 1000-residue protein.arrow_forwardLike a helices, B sheets often have one side facing the surface of the protein and one side facing the interior, giving rise to an amphiphilic sheet with one hydrophobic surface. From the sequences listed below, pick the one that could form a strand in an amphiphilic B sheet. Think about the way side chains are arranged in a strand of a ß sheet (See Figure below). a) ALSCDVETYWLI b) DKLVTSIAREFM c) DSETKNAVFLIL d) TLNISFQMELDV e) VLEFMDIASVLD IINIIarrow_forward
- Assume that the translational error frequency, δ, is 1 x 10–4.(a) Calculate the probability of making a perfect protein of 100 residues.(b) Repeat for a 1000-residue protein.arrow_forwardThe genetic information contained in DNA consists of a linear sequence of coding units known as codons. Each codon consists of three adjacent DNA nucleotides that corresponto a single amino acid in a protien. The E.coli DNA molecule contains 4.70 x 10^6 base pairs. Determine the number of codons that can be present. Assuming that the average protein in E.coli consists of a chain of 400 amino acids, calculate the maximum number of protiens that can be coded by an E.coli DNA molecule.arrow_forwardCystic fibrosis (CF) is an inherited disorder caused by different types of mutations, many of which prevent ions from moving across cell membranes. Normally there are channel proteins that allow passage of the ions, but in patients with one kind of CF these proteins seem odd. Closer examination shows that these proteins display the correct amino acid sequence. However, they fail to do their job. A) Given that the primary structure of the protein is correct, what can you infer about the DNA sequence for the gene coding this protein on this patient, is there a mutation? Explain. B) Why is the primary structure insufficient to guarantee the proper function of the protein?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON

Biochemistry
Biochemistry
ISBN:9781319114671
Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.
Publisher:W. H. Freeman

Lehninger Principles of Biochemistry
Biochemistry
ISBN:9781464126116
Author:David L. Nelson, Michael M. Cox
Publisher:W. H. Freeman

Fundamentals of Biochemistry: Life at the Molecul...
Biochemistry
ISBN:9781118918401
Author:Donald Voet, Judith G. Voet, Charlotte W. Pratt
Publisher:WILEY

Biochemistry
Biochemistry
ISBN:9781305961135
Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougal
Publisher:Cengage Learning

Biochemistry
Biochemistry
ISBN:9781305577206
Author:Reginald H. Garrett, Charles M. Grisham
Publisher:Cengage Learning

Fundamentals of General, Organic, and Biological ...
Biochemistry
ISBN:9780134015187
Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. Peterson
Publisher:PEARSON
Biomolecules - Protein - Amino acids; Author: Tutorials Point (India) Ltd.;https://www.youtube.com/watch?v=ySNVPDHJ0ek;License: Standard YouTube License, CC-BY