
Interpretation:
For the given reaction under given conditions the balanced equation, conjugate acid-base pair and the direction of equilibrium should be identified.
Concept introduction:
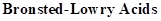 : If a species loses a proton then it is considered as
: If a species loses a proton then it is considered as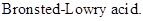
Strong Acids: Acids that dissociates into ions completely which results in easy donation of protons are considered as strong acids. Strong acid forms weaker conjugated base.
Weak Acids: Acids that do not easily dissociate into ions completely which has difficulty in proton donation are considered as weak acids. Weak acid forms stronger conjugated base
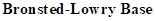 : If a species receives one proton, then it is considered as
: If a species receives one proton, then it is considered as 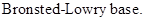
Strong Base: Bases that has strong attraction towards protons and accepts readily. Strong base forms weaker conjugated acid.
Weak Base: Bases that has little affinity towards protons. Weak base forms stronger conjugated acid.
If a base receives one proton, then the formed species is a conjugate acid whereas an acid lose one proton, then the formed species is a conjugated base.
Equilibrium: A state of a
The 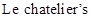 principle says that when a disturbance is given to equilibrium of a chemical reaction, the equilibrium will respond by shifting to the direction at which the effect of disturbance is nullified
principle says that when a disturbance is given to equilibrium of a chemical reaction, the equilibrium will respond by shifting to the direction at which the effect of disturbance is nullified
Balanced Chemical Reaction: On accordance with law of conservation of mass, for any chemical reaction, total masses of reactants and products must be equal.
Want to see the full answer?
Check out a sample textbook solution
Chapter 10 Solutions
Fundamentals of General, Organic, and Biological Chemistry, Books a la Carte Edition; Modified Mastering Chemistry with Pearson eText -- ValuePack ... and Biological Chemistry (4th Edition)
- Consider the following pH titration curve of a diprotic acid. What is the approximate values for pka 1 and pka 2? the curve is attached below.arrow_forwardGiven the titration curve of the hypothetical polyprotic acid X at 0.100 M concentration (pKa1=4.0, pKa2=8.0, pKa3=12.0) titrated with 0.600 M NaOH, identify the pH at point C, H, E, and M.arrow_forwardGlycine hydrochloride (Cl− H3N+CH2COOH) is a diprotic acid that contains a carboxylic acid group and an ammonium group and is therefore called an amino acid. It is often used in biochemical buffers. Solve, (b) Write the chemical equations describing the dissociation of the first and second protons of Cl−H3N+CH2COOH.arrow_forward
- Identify the acid and conjugate base in each reaction. Calculate the pKa for each acid. List them in order from the strongest to weakest acid. The acid-ionization constants, Ka, at 25°C are listed for each. a. HC2H3O2 + H2O ↔ H3O+ + C2H3O2-acetic acid, KA = 1.7 x 10-5 b. HC7H5O2 + H2O H₂O+ + C7H5O2-benzoic acid, KA= 6.3 x 10-5 c. HC6H4NO2 + H2O ↔ H3O++ C6H4NO2-nicotinic acid, KA = 1.4 x 10-5arrow_forwardRefer to the following titration curve below: 13 12 11 10 9 7 6 5 4 3 4 6 8 10 12 14 16 18 20 22 24 26 28 30 Volume of Titrant / mL - Unknown Acid 0.10 mol/L - titrant = NaOH 0.1 mol/L At ph 10.0, which form of histidine is most abundant? His2+ His- His° His+arrow_forwardSketch out the titration curve below and the structure(s) of the major ionic form(s) of glutamic acid that exist in solution at the noted parts of the titration curve. Refer to the fully protonated glutamic acid and the pKaarrow_forward
- Glycine hydrochloride (Cl− H3N+CH2COOH) is a diprotic acid that contains a carboxylic acid group and an ammonium group and is therefore called an amino acid. It is often used in biochemical buffers. Solve, In analogy with Figure , sketch the titration curve of this diprotic acid.arrow_forwardAn enzyme uses a lysine and a histidine as general acid-base catalytic residues. In one direction of the reaction, the lysine and histidine residues are typically protonated (~NH3*, ~Im), but in the other direction of the reaction, the two residues are "reverse protonated" (~NH2, ~Hlm*) (see below scheme). A pH profile of the reaction at various pH values yielded estimates of 7.57 and 9.30 for the macroscopic ionization constants pk,' and pK2, respectively. A separate set of experiments with site-speciffic variants of the enzyme yielded an estimate of 8.78 for the microscopic ionization constant pKp. Given this information, what is the value of the microscopic ionization constant pKA? Report your answer to the nearest hundredth. Lys-NH3* His-HIm" KB H KA H* Lys-NH2 Lys-NH3* His-HIm" His-Im Kc H* Kp Lys-NH2 His-Imarrow_forwardThe ionization of p-nitrophenol is shown below (pKa = 7.0): a. Identify the weak acid and conjugate base. b. At pH 7, what are the relative concentrations of ionized and un-ionized p-nitrophenol? c. If enough concentrated hydrochloric acid is added to a solution of p-nitrophenol to lower the pH from 7 to 5, what will happen to the relative concentrations of the ionized and un-ionized forms? d. Ionized p-nitrophenol has a yellow color, while the un-ionized form is colorless. The yellow color can be measured using a spectrophotometer at 400nm. In order to determine the total amount of p-nitrophenol in a solution, would you perform the spectrophotometer reading at an acidic or basic pH? Clearly explain why? e. A solution of p-nitrophenol at pH 7.95 was found to have an A400 of 0.255 . What is the total concentration (in µM) of p-nitrophenol (ionized plus un-ionized) in the solution? The molar extinction coefficient of p-nitrophenol is 18,500 M-1cm-1 and the pKa is 7.arrow_forward
- Mixtures of amino acids can be analyzed by first separating the mixture into its components through ion‑exchange chromatography. Amino acids placed on a cation‑exchange resin containing sulfonate (−SO−3)(−SO3−) groups flow down the column at different rates because of two factors that influence their movement: (1) ionic attraction between the sulfonate residues on the column and positively charged functional groups on the amino acids, and (2) aggregation of nonpolar amino acid side chains with the hydrophobic backbone of the polystyrene resin. Note that the ionic attraction is more important than hydrophobicity for this column media. For each pair of amino acids, identify which will be eluted first from a cation‑exchange column using a pH 7.0pH 7.0 buffer.arrow_forwardThe enzyme urease is widely used for determining urea concentration in blood. The Michaelis constant for urease at room temperature is 2.0 mM and k2 = 2.8 × 104 s-1 at pH = 7.5. Based on the kinetic experiment, when 0.34 mM urease was used, the initial rate of the reaction was determined to be 3.49 M/sec. What is the concentration of urea in blood in unit of mM? Please keep your answer to two decimal place. Please convert all concentration unit to mM in calculation.arrow_forwardWhat is the ratio of concentrations of acetate ion and undissociated acetic acid in a solution that has a pH of 5.12?arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





