
Concept explainers
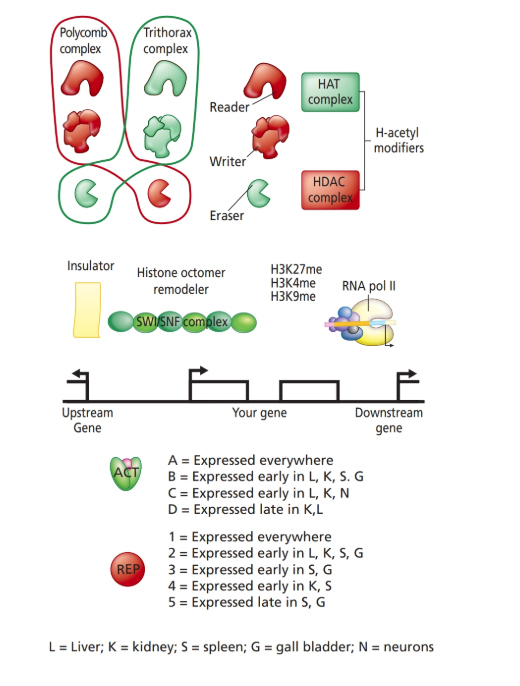
Using the components in the accompanying diagram, design regulatory modules (i.e., enhancer
a. How will the gene be activated in the proper cell type?
b. How will its expression be maintained?
c. How will expression be prevented in other cell types?
Want to see the full answer?
Check out a sample textbook solution
Chapter 13 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- Which of the following statements is true regarding the lys2-128d reporter? Select all that apply. a.) Mutants that activate this reporter are likely to have mutations in PIC components. b.) This reporter is sensitive to changes in chromatin maintenance because the start site is located within the ORF. c.) Mutants that activate this reporter cause downstream shifts in transcription start site selection. d.) This reporter is being used to help us rule out mutations that are likely impacting chromatin structure.arrow_forwardA gene, which we will call gene C, can be epigenetically modified in such a way that its expression in some cells is permanently silenced. Describe how you could conduct cell-fusion experiments to determine if a cis- or a trans-epigenetic mechanism is responsible for maintaining the silencing of gene C.arrow_forwardHow do I draw the sequence and explain the process for each step? Draw the sequence of molecular events that occurs to induce STAT transcription factor localization to the nucleus. To complete this, you will draw the first step that occurs, then draw a new figure with the second step that occurs, then draw a new figure with the third step that occurs, and so on until you have completed all of the steps. On the drawing, briefly label each molecular event (each drawing). For this brief label, explain what is happening during each step.arrow_forward
- A procedure called chromatin immunoprecipitation allows scientists to determine where aparticular protein is located in the genome. You conduct this procedure for a human regulatorytranscription factor and a histone acetyltransferase, and you find that the two proteins arepresent together at the promoters of many genes. Is the transcription factor more likely to bean activator or a repressor? Why?arrow_forwardYou are interested in studying a novel gene that appears to be involved in cancer. There is no information about the function of this gene. What would you do to obtain the cDNA for this gene? How would you express this gene and what expression systems might you utilize to study its function and why? How would determine the subcellular localization of this gene in eukaryotic cells? What are alternative methods in case one doesn't work? How would you purify and determine the 3-dimensional structure of this protein?arrow_forwardWhich of the following statements is most consistent with the pattern of gene expression shown in the given graph? (A) Repressors that bind to a regulatory sequence of Gene X are present in brain tissue but not in heart tissue. (B) Gene X is located within heterochromatin in brain tissue and within euchromatin in heart tissue. (C) Small RNAs that help degrade Gene X mRNA are present in brain tissue but not in heart tissue (D)Activators that bind to an enhancer of Gene X are present in brain tissue but not in heart tissue.arrow_forward
- Describe how chromatin remodeling complexes allowgene expression to occur.arrow_forwardGive typing answer with explanation and conclusion What characteristics of the DSX protein enable the female- and male-specific isoforms of DSX to regulate the same genes but with different outcomes in female and male development? [multiple answers possible] A.The two isoforms are different alleles of the same gene B.The two isoforms have different DNA-binding domains C.The two isoforms are lncRNAs involved in dosage compensation D.The two isoforms differ in their activation domain E.The two isoforms share the same activation domain F. The two isoforms share the same DNA-binding domainarrow_forwardDiscuss the following argument: “if the expression of every gene depends on a set of transcription regulators, then the expression of these regulators must also depend on the expression of other regulators, and their expression must depend on the expression of still other regulators, and so on. cells would therefore need an infinite number of genes, most of which would code for transcription regulators.” how does the cell get by without having to achieve the impossible?arrow_forward
- Describe the 4 types of suppressor high copy suppression, bypass suppression, nonsense suppression, protein interaction . in your Own words. Often geneticists look for suppressors to find interactive proteins. Which of the type(s) of suppressors you put for part a will help to identify interacting proteins, and which type(s) will not? What are two (or one, if we don’t get a chance to talk about two of them in class) other techniques (not necessarily “genetic” techniques, but at least, lab techniques) that help to identify identifying proteins?arrow_forwardMutations in the CFTR gene result in cystic fibrosis in humans, a conditions in which abnormal secretions are present in the lungs, pancreas, and sweat glands. The gene was mapped to a 500-kb region on chromosome 7 containing 3 candidate genes. a)Using your knowledge of the disease symptoms, how would you distinguish between the candidate genes to decide which is most likely to encode the CFTR gene? b)How would you prove that your chosen candidate is the CFTR gene?arrow_forwardA translocation mutation results in a silencer being inserted just upstream of the promoter of a gene (Gene X). How would you predict the expression of gene X be affected by this mutation? Gene X will not be expressed in any cell that has this mutation Gene X will not be expressed in any cell that has this mutation and the appropriate repressor protein Gene X will be expressed in any cell that has this mutation Gene X will be expressed in any cell that has this mutation and the appropriate repressor proteinarrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





