
Concept explainers
A muscle enzyme called ME
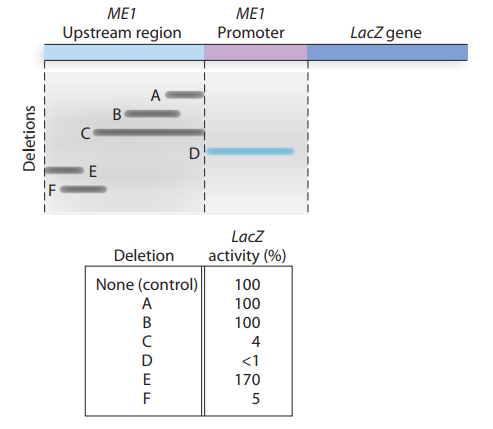
a. Does this information indicate the presence of enhancer and
b. Why does deletion D effectively eliminate transcription oflacZ?
c. Given the information available from deletion analysis, can you give a molecular explanation for the observation that ME
Want to see the full answer?
Check out a sample textbook solution
Chapter 13 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
- You are teaching a class on the regulation of eukaryotic gene expression. In order to demonstrate this complex process, you decide to draw for the class a typical eukaryotic gene/transcription unit with its major regions, such as the promoter regions, where the RNA polymerase II and transcription factors would bind From the list given - choose all components that you think are part of a typical eukaryotic gene From the list given - choose all the regulatory sequences that you think would control the expression of this eukaryotic gene From the list given - choose all of the regulatory proteins that would bind the eukaryotic gene to control its expressionarrow_forwardAn enhancer, located upstream from a gene, has the following sequence: 5′–GTAG–3′ 3′–CATC–5′ This enhancer is orientation-independent. Which of the following sequences also works as an enhancer? A. 5′–CTAC–3′ 3′–GATG–5′ B. 5′–GATG–3′ 3′–CTAC–5′ C. 5′–CATC–3′ 3′–GTAG–5′ C15.arrow_forwardMutations in bacterial promoters may increase or decrease the rate of gene transcription. Promoter mutations that increase the transcription rate are termed up-promoter mutations, and those that decrease the transcription rate are termed down-promoter mutations. As shown , the sequence of the −10 site of the promoter for the lac operon is TATGTT. Would you expect the following mutations to be up-promoter or down-promoter mutations? A. TATGTT to TATATT B. TATGTT to TTTGTT C. TATGTT to TATGATarrow_forward
- Mutations in bacterial promoters may increase or decrease the rate of gene transcription. Promoter mutations that increase the transcription rate are termed up-promoter mutations, and those that decrease the transcription rate are termed down-promoter mutations. If the promoter for the lac operon is TATGTT, would the following be uppromoters or downpromoters?A. TATGTT to TATATTB. TATGTT to TTTGTTC. TATGTT to TATGATarrow_forwardAn enhancer is surrounded by four genes (A, B, C, and D), as shown in the accompanying diagram. An insulator lies between gene C and gene D. On the basis of the positions of the genes, the enhancer, and the insulator, the transcription of which genes is most likely to be stimulated by the enhancer? Explain your reasoning.arrow_forwardWhich of the following mutations could be appropriately describedas a position effect?A. A point mutation at the –10 position in the promoter regionprevents transcription. B. A translocation places the coding sequence for a muscle-specificgene next to an enhancer that is turned on in nerve cells.C. An inversion flips a gene from the long arm of chromosome 17(which is euchromatic) to the short arm (which isheterochromatic).arrow_forward
- Name the lambda promoters whose expression is regulated by the cro protein. For each promoter you named, is cro an activator or a repressor of transcription from that promoter?arrow_forwardRegarding the process of gene transcription in eukaryotes, it is correct to state that A)The transcription process is terminated when the RNA polymerase complex reaches the final region of the gene with the poly-adenylation signal. B)The RNA polymerase II transcription elongation complex contains transcription factors such as the TATA box binding protein C)The opening of the region of DNA that will be transcribed is done by the DNA helicase, present in the transcription complex. D)The main function of the Mediator coactivator is to promote the transition between elongation and completion of the transcription process. E)Different activators and repressors can influence the transcriptional elongation complex by binding to the promoter regions of genesarrow_forwardMost eukaryotic promoters have binding sites for several different transcription factors, and the frequency with which transcription is initiated at a promoter depends on the specific combination of transcriptional regulators bound to their binding sites in that promoter. Transcription of the slither gene in garter snakes is regulated by the transcriptional activators Python and Boa and the transcriptional repressor Sidewinder. Each of these proteins has one binding site in the slither promoter; the affinity of Boa for its binding site is 30 times higher than the affinity of Python for its binding site and 6 times higher than the affinity of Sidewinder for its binding site. Under conditions where Sidewinder is 10 times more abundant than Python, and Python is 3 times as abundant as Boa, would you expect transcription of the slither gene to be activated or repressed? Show or briefly explain how/why you predicted the outcome you chose.arrow_forward
- A strain of Arabidopsis thaliana possesses a mutation in the APETALA2 gene. As a result of this mutation, much of the 3′ UTR of the mRNA transcribed from the gene is deleted. What is the most likely effect of this mutation on the expression of the APETALA2 gene?arrow_forwardThe UG4 gene is expressed in the stem and leaf tissue of the plant Arabidopsis thaliana. To identify potential mechanisms regulating UG4 gene expression, six small deletion mutations are made in cloned sequence containing the upstream regulatory region. The full length mutant segments Which mutation(s) affect an enhancer? Why? Which mutations identify the promoter? Why? Speculate about the reason for different transcription rated obtained for fusion constructs E and F.arrow_forwardA crucial step in the regulation of many bacterial genes is the binding of RNA polymerase to DNA at the promoter. Why might it be advantageous for bacteria to regulate the expression of their genes at this particular step?arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
