
GENETIC ANALYSIS: INTEGRATED - ACCESS
3rd Edition
ISBN: 9780135349298
Author: Sanders
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 15, Problem 15P
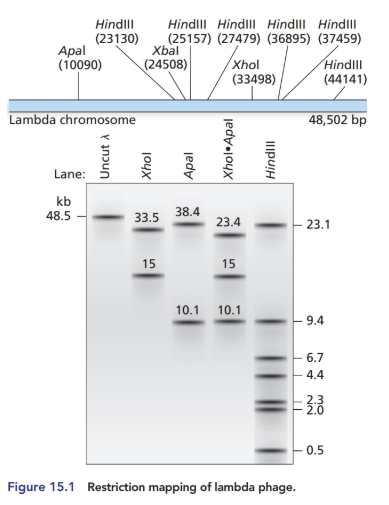
The bacteriophage lambda genome can exist in either a linear form (see Figures
How many fragments will be formed by restriction enzyme digestion with XhoI alone, with XbaI alone and with both XhoI and XbaIin the linear and circular forms of the lambda genome?
Diagram the resulting fragments as they would appear on an agarose gel after electrophoresis.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
A 2.0kb bacterial plasmid ‘BS1030’ is digested with the restriction endonuclease Sau3A; the plasmid map is depicted in the diagram below and the Sau3A (S) restriction sites are indicated.
Which of the following DNA fragments do you expect to see on an agarose gel when you run Sau3A-digested plasmid ‘BS1030’ DNA?
a.
250 bp, 450 bp, 550 bp, 1.1 kb, 1.5 kb and 2.0 kb
b.
2.0kb
c.
250 bp, 400 bp, 450 bp, 500 bp and 550 bp
d.
100 bp, 200 bp, 250 bp, 400 bp, 500 bp and 550 bp
A 12 kb linear DNA fragment is subject to single or double RE digest and agarose gelelectrophoresis, to yield the gel profile shown below. The first lane contains the size marker(M).a) Explain how the name of the enzyme EcoRI is derived.b) How many sites are there for EcoRI and PvuII respectively on this DNA fragment?c) Use the sizes of the DNA bands on the gel to compile a restriction enzyme map of the DNAfragment. Indicate the positions of the restriction enzymes sites for EcoRI and PvuII on themap.
Consider the following plasmid (size 8000 bp), with restriction sites at the positions indicated: (see image)
a) This plasmid is digested with the enzymes listed below. Indicate how many fragments will begenerated in each case, and give the sizes of the fragments.PstIXhoICombination of PstI + XhoI + EcoRI (triple digest)
b) Draw the banding pattern you would expect to observe if each of these digestions is loaded into a separate well of an agarose gel, and the fragments separated by electrophoresis. In the first well you load a DNA marker (M) containing fragments with sizes of 1000 bp, 2000 bp, 4000 bp and 8000 bp.
c) This gel is transferred to a membrane in a Southern blot experiment, and hybridised to a radioactively labelled 200 bp probe, which anneals to the plasmid DNA at the position indicated on the diagram above. Draw the autoradiographic profile you would expect to observe for the membrane.
Chapter 15 Solutions
GENETIC ANALYSIS: INTEGRATED - ACCESS
Ch. 15 - 15.1 What purpose do the bla and lacZ genes serve...Ch. 15 - The human genome is 3109 bp in length. How many...Ch. 15 - 15.3 Ligase catalyzes a reaction between the...Ch. 15 - You have constructed four different libraries: a...Ch. 15 - Using the genomic libraries in Problem 4, you wish...Ch. 15 - The human genome is 3109bp. You wish to design a...Ch. 15 - 15.7 Using animal models of human diseases can...Ch. 15 - 15.8 Compare methods for constructing homologous...Ch. 15 - 15.9 Chimeric genefusion products can be used for...Ch. 15 - 15.10 Why are diseases of the blood simpler...
Ch. 15 - Injection of double-stranded RNA can lead to gene...Ch. 15 - Compare and contrast methods for making transgenic...Ch. 15 - 15.13 It is often desirable to insert cDNAs into a...Ch. 15 - 15.14 A major advance in the s was the development...Ch. 15 - 15.15 The bacteriophage lambda genome can exist in...Ch. 15 - 15.16 The restriction enzymes Xho and Sal cut...Ch. 15 - 15.17 The bacteriophage has a single-stranded DNA...Ch. 15 - 15.18 To further analyze the CRABS CLAW gene (see...Ch. 15 - You have isolated a genomic clone with an EcoR I...Ch. 15 - 15.20 You have identified a cDNA clone that...Ch. 15 - 15.21 You have isolated another cDNA clone of the...Ch. 15 - 15.22 You have identified five genes in S....Ch. 15 - You have generated three transgenic lines of maize...Ch. 15 - 15.24 Bacterial Pseudomonas species often possess...Ch. 15 - 15.25 Two complaints about some transgenic plants...Ch. 15 - 15.26 In Drosophila, lossoffunction Ultrabithorax...Ch. 15 - Prob. 27PCh. 15 - The highlighted sequence shown below is the one...Ch. 15 - Vitamin E is the name for a set of chemically...Ch. 15 - The RAS gene encodes a signaling protein that...Ch. 15 - 15.31 You have cloned a gene for an enzyme that...Ch. 15 - 15.32 About of occurrences of nonautoimmune type...Ch. 15 - Describe how having the Cas 9 gene at a genomic...Ch. 15 - 15.34 Would a gene drive system spread rapidly...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Bacterial systems serve as an excellent model to express proteins but has a disadvantage – what is that disadvantage? List the points to differentiate three classes of restriction enzymes? Give an example of restriction enzyme that has ability to generate blunt and cohesive ends after digestion of DNA?arrow_forwardIt is desired to isolate genomic DNA from liquid culture of S. cerevisiae yeast. A commercial kit will be used to isolate genomic DNA from this liquid culture. Answer the following questions to understand the strategy used by commercial kits for genomic DNA isolation. a) List all the steps from cell pellet preparation to DNA elution. b) With which feature can the membrane in the column that comes with the commercial kit bind DNA? c) Which component in the kit would you use to recover the DNA from the membrane of the column to which the DNA was attached?arrow_forwardExamine the structure of the pBR322 plasmid depicted below. Assume total size of the plasmid is 4,361 bp and the blue numbers indicate locations of restriction sites relative to the O point at the top of the plasmid. What size fragments would be generated by the following restriction digestion reactions? 1. Sall 2. Sal 1 + BamH1 3. Sal 1 + EcoR1 4. Sal I + BamH1 + EcoR1 PstI 3607 3000 4000 amp ori HindIII Edit View Insert Format Tools Table EcoRI EcoRV 4359 0 29 185 pBR322 4361 bp 2295 NdeI tet 2000 BamHI 375 651 SalI 1000 Type a short answer in the space provided below.arrow_forward
- The map of plasmid pUC19 is shown below. The restriction site coordinate is the position of the 5’base on the top strand of each site sequence. The restriction enzyme sites are in bold type if there is only one site in pUC19. Please list the fragments in order of size, largest to smallest, which will result from a complete digestion by the restriction enzyme PvuII. Please list the fragments in order of size, largest to smallest, which will result from a complete digestion by the restriction enzyme DrdI.arrow_forwardBacteriophage lambda (λ) consists primarily of a head, which contains the genomic DNA, and a tail that is involved in phage attachment to bacterial cells. The diagram to the right represents the bacteriophage lambda genomic DNA, showing the locations of important gene clusters (Ausubel et al. 1998). Arrows below mark the sites where the restriction enzyme HindIII cuts the DNA, and the numbers indicate the number of base pairs in each fragment. The restriction enzyme HindIII cuts the λ genome 7 times, how many fragments are produced following this reaction?arrow_forwardA small DNA molecule was cleaved with several different restriction nucleases, and the size of each fragment was determined by gel electrophoresis.The following data were obtained. (a) Is the original molecule linear or circular?(b) Draw a map of restriction sites (showing distances between sites) that isconsistent with the data given.(c) How many additional maps are compatible with the data?(d) What would have to be done to locate the cleavage sites unambiguouslywith respect to each other?arrow_forward
- In the formation of recombinant DNA, a restriction endonuclease cuts a bacterial plasmid to give sticky ends. The DNA segments that are to be added to the plasmid are cleaved with the same restriction endonuclease. What aresticky ends and why is it important that the target DNA and the plasmid it will be incorporated into have complementary sticky ends?arrow_forwardConstruct a detail linear restriction map according to the given information in Table 1 below. Table 1 Restriction enzyme EcoRI No, of fragments 2 Size of fragment (kb) 2,9 ВатHI 3 6, 1,4 Xhol 8, 3 EcoRI + BamHI 4 2, 4, 1,4 EcoRI + Xhol 3 2, 6, 3arrow_forwardRestriction endonucleases are bacterial enzymes that cleave duplex (double-stranded) DNA at specific nucleotide sequences. The mode of replication of the animal virus SV40 has been investigated by using restriction endonucleases that cleave SV40 DNA into a number of unique segments. Like most viruses, SV40 DNA is circular. The map positions of the 11 fragments produced by a pair of restriction endonucleases are shown on the next page. Immediately following a 5 or 10 minute pulse of radioactively labeled thymidine, labeled SV40 molecules that have completed replication during the pulse are isolated. These newly replicated DNA molecules are digested by the restriction endonucleases and the resulting fragments are analyzed for the relative amounts of pulse label they contain. The results are in the table below. Assume that at the time the label was added there was a random population of replicating SV40 DNA molecules in all possible stages of synthesis. From the information given below,…arrow_forward
- All are correct about DNA gyrase in E. coli EXCEPT: It works to remove positive supercoiling introduced by the DnaB protein (helicase). It is a topoisomerase that hydrolyzes ATP during its reaction mechanism. Its mechanism involves the breaking of a single phosphoester bond in one strand of dsDNA. It works to relieve supercoiling in DNA to overcome the torsion stress imposed upon unwinding.arrow_forwardHow would you expect your results to change if the enzyme did not have time to cut all of the sites? (Hint, remember that there are many pieces of the lambda DNA in the tube – draw yourself a simplified model where there are four restriction sites, and imagine what would happen if the enzyme only cut at one, two, three or all four, randomly, each time). How would you expect your results to change if you treated them with both EcoRI and HindIII in the same tube? Why might it be safer for the virus to have its DNA become circular when it enters the host, rather than remain linear?arrow_forwardA plasmid DNA and a linear DNA (both of the same size) have one site for a restriction endonuclease. When cut and separated on agarose gel electrophoresis, plasmid shows one DNA band while linear DNA shows two fragments. Explain.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license