
Concept explainers
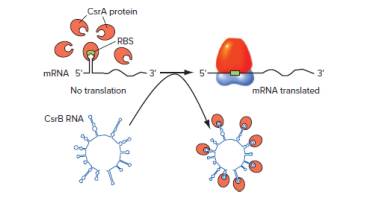
Great variation exists in the mechanisms by which RNAs can mediate gene regulation. In one recently discovered example shown in the following diagram, the genes CsrA and CsrB are global regulators of suites of target genes that are involved in the use of carbon atoms. The product of CsrA is the CsrA protein, which binds to the ribosome binding site (RBS) of target gene mRNA, preventing target gene expression. The product of the CsrB
gene is the CsrB RNA, which contains 22 binding sites for CsrA protein. CsrB RNA can thus compete with target mRNAs for CsrA protein binding. In the presence of high CsrB RNA concentrations, CsrA protein cannot bind to mRNA binding sites, so expression of the target genes is turned on.
| a. For the CsrA and CsrB genes, indicate what kind of molecule the gene product is (DNA, RNA, protein, small molecule), whether it acts as a positive or negative regulator, what stage of gene expression it affects, and whether it acts in cis or in trans. (It will be interesting to compare your answers to those for Problem 30.) |
| b. CsrA/CsrB regulate glycogen biosynthesis and breakdown; glycogen is a To what environmental factor do you think that the CsrA/CsrB system is most likely to respond? Suggest a possible way that this system might be repressible, and then suggest a different hypothesis for how this system might be inducible. (Assume in both cases that CsrB expression is modulated.) Which of your hypotheses would be most consistent with the target genes being involved in glycogen biosynthesis (an anabolic pathway), and which is most consistent with glycogen breakdown (a catabolic pathway)? Explain. |
Want to see the full answer?
Check out a sample textbook solution
Chapter 16 Solutions
Genetics: From Genes to Genomes
- The yeast gene SER3, whose product has a role in serine biosynthesis, is repressed during growth in nutrient-rich medium, so little transcription takes place, and little SER3 enzyme is produced, under these conditions. In an investigation of the repression of the SER3 gene, a region of DNA upstream of SER3 was found to be heavily transcribed when SER3 is repressed ). Within this upstream region is a promoter that stimulates the transcription of an RNA molecule called SRG1 RNA (for SER3 regulatory gene 1). This RNA molecule has none of the sequences necessary for translation. Mutations in the promoter for SRG1 result in the disappearance of SRG1 RNA, and these mutations remove the repression of SER3. When RNA polymerase binds to the SRG1 promoter, the polymerase travels downstream, transcribing the SGR1 RNA, and passes through and transcribes the promoter for SER3. This activity leads to the repression of SER3. Propose a possible explanation for how the transcription of SGR1 might…arrow_forwardThe insertion of transposable elements into genes can alter the normal pattern of expression. In the following situations, describe the possible consequences on gene expression.a. A LINE inserts into an enhancer of a human gene. b. A transposable element contains a binding site for a transcriptional repressor and inserts adjacent to a promoter. c. An Alu element inserts into the 3′ splice (AG) site of an intron in a human gene. d. A Ds element that was inserted into the exon of a maize gene excises imperfectly and leaves three base pairs behind in the exon. e. Another excision by that same Ds element leaves two base pairs behind in the exon. f. A Ds element that was inserted into the middle of an intron excises imperfectly and leaves five base pairs behind in the intron.arrow_forwardConsider Figure 3, which shows some features of a eukaryotic gene. A, B, C are exons while 1, 2 are introns. E marks an enhancer. The 5’ UTR and 3’ UTR are also marked. Which one of the following features would you expect to NOT be included in the primary RNA (before any processing has occurred) that results from transcription of this gene? Region marked E Region marked 5’ UTR Regions marked A, B and C Regions marked 1 and 2arrow_forward
- For each statement about gene expression mechanisms, choose the correct end to the sentence. For each gene, the template strand for transcription is determined by…. The direction of translation is determined by…… The tissue-specificity of protein production is determined by…. choices: a. location of the start codon b. location of the promoter c. direction of polymerization by RNA polymerase d. none of these e. direction of movement of ribosomes f. overall orientation of the chromosomearrow_forwardYou have isolated a protein that binds to DNA in the region upstream of the promoter sequence of a gene sys. If this protein is a positive regulator, which of the following would be true (multiple selection possible)? Loss-of-function mutations in the gene encoding the DNA-binding protein would cause constitutive expression of sys. Loss-of-function mutations in the gene encoding the DNA-binding protein would result in little or no expression of sys.arrow_forwardCompare the control of gene regulation in eukaryotes and prokaryotes at the level of initiation of transcription. How do the regulatory mechanisms work? What are the similarities and differences in these two types of organisms in terms of the specific components of the regulatory mechanisms? Address how the differences or similarities relate to the biological context of the control of gene expression.arrow_forward
- A. A mutation is recovered in the gene that encodes the lactose operon repressor protein (LacI). Which of the following lac phenotypes would lead to the conclusion that the mutation causes an inability of the repressor to bind lactose? The lac structural genes would never be expressed. The lac structural genes would be expressed continuously. The lac structural genes would be expressed efficiently only in the absence of lactose. The lac structural genes would be expressed efficiently only in the presence of lactose. B. A mutation is recovered in the gene that encodes the lactose operon repressor protein (LacI). Which of the following lac phenotypes would lead to the conclusion that the mutation causes an inability of the repressor to bind Operator DNA? The lac genes would be expressed efficiently only in the presence of lactose. The lac genes would be expressed efficiently only in the absence of lactose. The lac genes would never be expressed. The lac genes would be…arrow_forwardA mutation in the trp repressor gene that prevents the apo-repressor from binding the co-repressor What is the correct option from the choices below? causes the Trp repressor to permanently bind the trp operator. leads to a high level of transcription initiation from the trp promoter, even when tryptophan is present in the medium. does not lead to a high concentration of mRNA molecules that code for the structural genes trp E, D, C, B, A, when tryptophan is present in the medium, because attenuation of the operon will still work. a and b, but not c b and c, but not aarrow_forwardTranscriptional repressor proteins (e.g., lac repressor), antisense RNA, and feedback inhibition are three different mechanisms that turn off the expression of genes and gene products. Which of these three mechanisms will be most effective in each of the following situations? A. Shutting down the synthesis of a polypeptide B. Shutting down the synthesis of mRNA C. Shutting off the function of a protein For your answers to parts A–C that list more than one mechanism, which mechanism will be the fastest or the most efficient?arrow_forward
- An enhancer, located upstream from a gene, has the following sequence: 5′–GTAG–3′ 3′–CATC–5′ This enhancer is orientation-independent. Which of the following sequences also works as an enhancer? A. 5′–CTAC–3′ 3′–GATG–5′ B. 5′–GATG–3′ 3′–CTAC–5′ C. 5′–CATC–3′ 3′–GTAG–5′ C15.arrow_forwardRegarding transcriptional promoter sites, which of the following statements are true? Select one or more than one: a)They are located in the gene (DNA) whose information will be transcribed b)They are found at the 3 'end of the gene that will be transcribed c)Some of them are called 'TATA box' d)They are found in the DNA, 'upstream' of the gene to be transcribed. e)They are proteins of the cytoplasmarrow_forwardExplain how the following mutations would affect transcription of the yeast GAL1 gene in the presence of galactose. (a) A deletion within the GAL4 gene that removes the region encoding amino acids 1 to 100. (b) A deletion of the entire GAL3 gene. (c) A mutation within the GAL80 gene that blocks the ability of Gal80 protein to interact with Gal3p. (d) A deletion of one of the four UASG elements upstream from the GAL1 gene. (e) A point mutation in the GAL1 core promoter that alters the sequence of the TATA box.arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





