
EBK LIFE: THE SCIENCE OF BIOLOGY
11th Edition
ISBN: 8220103935432
Author: Sadava
Publisher: MAC HIGHER
expand_more
expand_more
format_list_bulleted
Concept explainers
Question
Chapter 17, Problem 4Q
Summary Introduction
To review:
The level of activity of the enzyme coded by ‘A’ allele of the gene compared to ‘C’ allele.
Given:
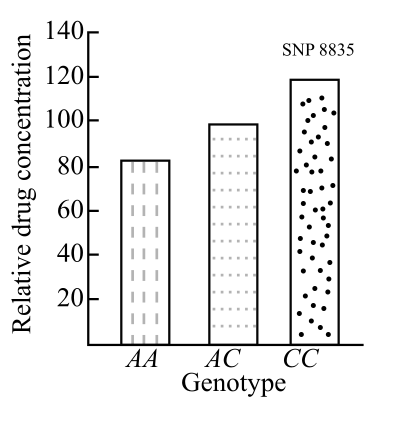
Introduction:
The rate of
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
For the genotypes and conditions (lactose present or absent) shown in the following Table 2,
predict whether the expression of beta-galactosidase is inducible (I), noninducible (N) or
constitutive (C). Explain your reason.
Table 2
Genotype
I*p*O°z*
Condition
(i)
No lactose
(ii)
I*P*O*Z+
Lactose
I*P-0*Z*
(iii)
(iv)
Lactose
ISP+O°Z+
Lactose
(v)
Ip*o°Z+
No lactose
In the case of normal RNase A, Anfisen found that oxidizing the Cys residues before slowly removing the
urea gave a very different result than oxidizing the Cys after slowly removing the urea.
When the urea was removed first, Anfisen recovered 100% of the catalytic activity. When the urea was
removed after oxidation, only about 1% of the activity (1/105) was recovered. The conclusion was that
104 of every 105 molecules was misfolded, and therefore catalytically inactive.
Let's say someone raises an objection to this interpretation, and says that perhaps, instead, something
chemically happens to the active site of RNase A when it is oxidized before refolding, and that the ~1%
activity measured represents the residual and greatly reduced enzyme activity of all of the molecules,
and the number is just coincidently similar to 1/105. (In other words, instead of 104 completely inactive
molecules for every 1 completely active molecule, all molecules are equally damaged, and only function
at…
For the genotypes and conditions (lactose present or absent) shown in the following Table 2,
predict whether the expression of beta-galactosidase is inducible (I), noninducible (N) or
constitutive (C). Explain your reason.
Table 2
Genotype
I*p*o°z+
Condition
(i)
No lactose
(ii)
(iii)
(iv)
I*P*O*Z+
I*P•O*Z*
I$P*O°Z*
Lactose
Lactose
Lactose
(v)
IP*0°Z+
No lactose
Chapter 17 Solutions
EBK LIFE: THE SCIENCE OF BIOLOGY
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- You want to study the effect of lysine methylation on gene regulation. The lysine amino acid sidechain is -CH2CH2CH2CH2NH2. To begin to study the relevant functional groups, you take free CH3CH2CH2CH2NH2 in the lab and decide to try to turn it into the product (P) CH3CH2CH2CH2NHCH3. 7. a) Suggest a simple reagent to carry out the desired transformation. (In the subsequent parts of the problem, you may call this reagent R for short). b) Draw the reaction using the arrow formalism. c) What solvent and pH do you suggest? Why? d) Is the reaction first order or second order? e) Write down a simple rate equation for dP/dt. If you include any constant(s), what are their units? f) In addition to the desired product P, name at least one other impurity (containing both carbon and nitrogen, you may call this compound U for short) that is likely to occur in substantial amounts. g) What choice of initial conditions would bias towards production of substantially more P than U? h) In biology, what…arrow_forwardWhat are histone deacetylase inhibitors (HDACi)? Give example(s)arrow_forwardIn the: Insertion of G after the 2nd T in the TATA box Explain: (a) What is the process affected? (b) What is the Effect on the process? (c) Does it affect prokaryotes, eukaryotes or both?arrow_forward
- How many ATP and GTP molecules would be hydrolyzed in the synthesis of a 125 amino acid protein? Explain why this is a good argument for careful regulation of gene expression (especially considering that most proteins are larger than 125 amino acids).arrow_forwardWhy is the position of Cys 58 important in 3GRS(GLUTATHIONE REDUCTASE)? When Cys 58 is mutated to GLY 58 how would it impact the 3D structure and function of 3GRS? explain in terms of how Cys and GLY have different properties and how it would impact the function of 3GRS (the binding sites etc.) You can see your structure(3GRS ) here or any other website: https://www.rcsb.orgarrow_forwardWhat can you determine from the r-value about the relationship between gene copy number and amylase enzyme concentration? r value = 0.935833048arrow_forward
- Consider the mechanism of the enzyme RNase: What would happen to the Km (i.e., would it increase, decrease, or stay the same) if the his12 was mutated to a lysine? Explain. What would happen to the Kcat (i.e., would it increase, decrease, or stay the same) if the his12 was mutated to a valine? Explain.arrow_forwardYou generate mutants in the metabolic pathway for starlase. You conduct some complementation tests (after testing for dominance of course) and come up with the following results: 1 2 3 4 5 6 1 2 + 3 + + a. How many complementation groups are there? [Select] 4 + 5 + + + + 6 +arrow_forwardClary Fray used the pET vector system to express her prokaryotic amylase enzyme. She added peptone into her culture broth of BL21(DE3) Escherichia coli strain to induce protein expression. At the end of the experiment, she discovered that her protein was not expressed. She repeated three more times but her protein of interest was still not produced. (i) (ii) (iii) (iv) (v) Explain the reason why Clary failed to obtain her protein of interest and suggest a solution to troubleshoot this problem. Clary plans to express her protein along with a polyhistidine-tag. Explain the importance of His-tag in protein work. Is DH5a Escherichia coli suitable to propagate the plasmid before protein expression? Besides heat shock method, elaborate another procedure Clary could utilize to transform the recombinant pET vector into the host cell. If her supervisor instructs her to express a gene from gold fish (Carassius auratus), is the expression system she is using now suitable for this experiment?…arrow_forward
- A recent genome sequencing project for the bacterium Burkholderia mallei has identified a new protein with high similarity to the lysylphosphatidylglycerol flippase enzyme. A short section of the new protein sequence is shown below. TVEVNAPGDVQKALSELQQINDGRLDIRI (a) Are any reverse turns likely to be present? Explain your answer. (b) Are any beta-strands likely to be present? Explain your answer. (c) Are any alpha helices likely to be present? Explain your answer. (d) Is any supersecondary structure likely to be present? Explain your answer. (e) Identify two residues that are likely to be buried in the core of the folded protein. Explain your answer. (f) Identify two residues that are likely to be hydrogen bonded to each other. Explain your answer.arrow_forwardThe lac genotypes are as shown below: P+OcZ-Y+A+// P¯O+Z+Y+A+ (i) The lac operon consists of three structural genes, lacZ, lacY and lacA. Which structural genes are involved in lactose metabolism? Explain. (ii) Draw and explain how lactose repress the gene expression in lac IS/I- heterozygote. (iii) What is the function of the promoter in the bacterial operon?arrow_forwardA Leu →Ala mutation at a site buried in the core of the enzyme lysozymeis found to be destabilizing. Explain the observed effect of this mutationon lysozyme stability by predicting how enthalpy (ΔH°), conformationalentropy (ΔS°peptide), and the hydrophobic effect (ΔS°solvent) are expected to change for the mutant compared to wild-type lysozyme. Explain how ΔG°for unfolding is affected by your predicted changes in enthalpy or entropy.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education

Human Anatomy & Physiology (11th Edition)
Biology
ISBN:9780134580999
Author:Elaine N. Marieb, Katja N. Hoehn
Publisher:PEARSON

Biology 2e
Biology
ISBN:9781947172517
Author:Matthew Douglas, Jung Choi, Mary Ann Clark
Publisher:OpenStax

Anatomy & Physiology
Biology
ISBN:9781259398629
Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa Stouter
Publisher:Mcgraw Hill Education,

Molecular Biology of the Cell (Sixth Edition)
Biology
ISBN:9780815344322
Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter Walter
Publisher:W. W. Norton & Company

Laboratory Manual For Human Anatomy & Physiology
Biology
ISBN:9781260159363
Author:Martin, Terry R., Prentice-craver, Cynthia
Publisher:McGraw-Hill Publishing Co.

Inquiry Into Life (16th Edition)
Biology
ISBN:9781260231700
Author:Sylvia S. Mader, Michael Windelspecht
Publisher:McGraw Hill Education
Mitochondrial mutations; Author: Useful Genetics;https://www.youtube.com/watch?v=GvgXe-3RJeU;License: CC-BY