
Concept explainers
Core Skill: Modeling The goal of this modeling challenge is to make a model that shows how a homeotic protein can interact with mediator.
Modeling Challenge: A gene, which we will call gene x, has an enhancer that is activated by a homeotic protein. The homedomain binds to the enhancer, as shown in Figure 20.15b. The enhancer is a long distance from the core promoter of gene X. After the homeotic protein binds to the enhancer, it interacts with mediator (refer back to Figure 14.16) to stimulate the transcription of gene X. Draw a model that shows how this homeotic protein interacts with mediator.
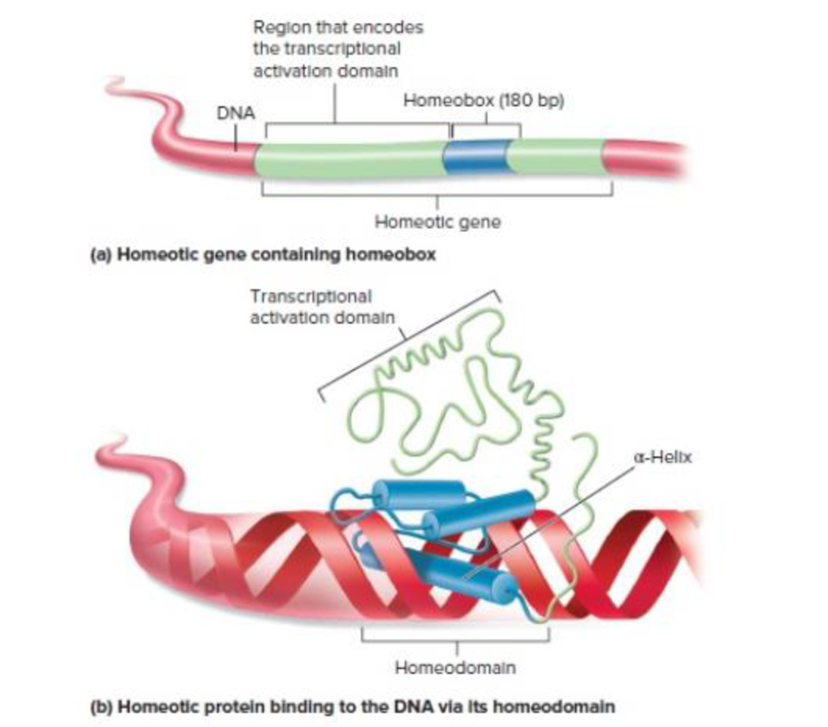
Figure 20.15 Molecular features of homeotic genes and proteins. (a) A homeotic gene (shown mostly in green) contains a 180-bp sequence called the homeobox (shown in blue). (b) Homeotic genes encode homeotic proteins that function as transcription factors. The homeotic protein contains two key domains called the homeodomain and the transcriptional activation domain. The homeodomain binds to the DNA at a regulatory site such as an enhancer. After this binding occurs, the transcriptional activation domain activates RNA polymerase to begin transcription.
Want to see the full answer?
Check out a sample textbook solution
Chapter 20 Solutions
Biology
Additional Science Textbook Solutions
Biology: Life on Earth with Physiology (11th Edition)
Marine Biology (Botany, Zoology, Ecology and Evolution)
Biology Illinois Edition (Glencoe Science)
Biology: Concepts and Investigations
Biological Science (6th Edition)
- Explain how targeted gene silencing and knockout mutations can give insights into the functions of a gene.arrow_forwardWhy do gene targeting and mutagenesis screening in mice have potential benefits for humans?arrow_forwardDiscuss how qPCR, DNA microarrays (DNA chips), and RNA-seq analysis are used to compare levels of gene expression among normal versus diseased tissues.arrow_forward
- Q.Consider this problem: You are working in the lab to study the pattern of paralysis and candidate genes involved in this process. You revealed that homozygous mutation in the SLB gene is the main cause of paralysis. The following sequence of SLB gene at the beginning of the translated region is found in individuals without paralysis. 5‟-GTA GCA TTT AAG CTT CAG TCC AAG - 3‟ (Met Thr Phe Glu Ile Gln Ser Arg). This sequence is however changed to the following sequence in affected individuals 5‟- GTA GCA TTT AAG CTT TAG TCC AAG - 3‟ (Met Thr Phe Glu Ile STOP). What mutation you will identify from this observation? Explain with reason. After identifying mutation if there is any, enlist all the possible repair mechanisms that can be helpful to repair the current situation. Elaborate your answer with that why you think that your suggested repair pathway should be used and how it can be effective for repairing in the current scenario?arrow_forwardEssay: Relate the structure of the DNA to its role as a molecule for inheritance. Elaborate on the significance of DNA Describe the molecular mechanisms involved in P53’srole as a tumor repressor protein packaging in the inheritance and transmission of traits.arrow_forwardCompare the auto induction media (AIM) with regular induced protein expression. Which is better when conducting experiments?arrow_forward
- Explain in detail the role of super-enhancers. Give some examples. How the super-enhancers can be identified?arrow_forwardDescribe in detail the role of super-enhancers. Give some examples. How the super-enhancers can be identified?arrow_forwardQ1. Bioluminescence is emitted by the marine bacterium Vibrio fischeri in response to the concentration of a chemical signal. The luxCDABEG genes form part of an operon that encodes all of the structural components necessary for light production. At high cell density, the concentration of the inducer increases and can bind to LuxR protein which then activates transcription of the operon. What kind of operon is the Lux operon? [Positive Inducible, Positive Repressible, Negative Inducible, Negative Repressible?] Q2. A population of bacteria has a Lux operon that produces bioluminesence via an autoinducer that activates a transcription factor. You grow a flask of cells that have a loss-of-function mutation in the gene that encodes the enzyme that produces an autoinducer. What is the predicted bioluminesence in this population at high and low density?arrow_forward
- genetics course 2. Describe why PARP inhibitors are considered targeted therapies (affecting only cancer cells and not healthy cells)? Describe their mechanism of action.arrow_forwardActivity 1.4: Essay.Direction: Explain your answer.Today, it is easy to make transgenic plants and animals. What are some important safety and ethical issues raised by this use of recombinant DNA technology? Explain your answer.arrow_forwardTask #1 Mc1r alleles: In Florida Gulf coast mice, there are two different alleles of the Melanocortin Receptor (Mcr1) gene. The sequence for both alleles is shown below. This is an internal segment of the coding (non-template) sequence of the Mcr1 gene. The reading frame is set and you do not need to find an AUG. M allele 5’...ATC ACC AAA AAC CGC AAC CTG CAC TCG... m allele 5’...ATC ACC AAA AAC TGC AAC CTG CAC TCG... A. Compare these two allele sequences and circle the nucleotides that are different. B. Do you think this change will cause a shift in the codon reading frame? Why or why not? C. Do you think that this change will cause an early stop in translation? Why or why not? Task #2 Flow of information: A codon table is provided above. The 5’ codon nucleotide is in the left column, and second codon nucleotide is on top. The Mcr1 M and m allele sequences are shown again in the central dogma grids below with the reading frame designated. Fill in the grids. In the second column…arrow_forward
 Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning
Biology (MindTap Course List)BiologyISBN:9781337392938Author:Eldra Solomon, Charles Martin, Diana W. Martin, Linda R. BergPublisher:Cengage Learning


