
Concept explainers
Interpretation:
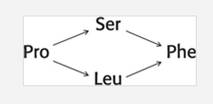
A single-stranded RNA from tobacco mosaic virus was treated with a chemical mutagen and mutants were obtained having serine or leucine instead of proline at specific position.Further treatment of these mutants with same mutagen yielded phenylanaline at this position. Hence, the plausible codonassignments for the given four amino acids are to be depicted.
Concept introduction:
A codonis defined as the sequence ofthree nucleotides which encode for anamino acid duringprotein synthesis or translation.For example,CCT, and CTT are the codons which codes for proline and leucine respectively.
A chemical mutagen is a substance that can cause alteration of the nitrogenous base incorporated in the DNA.
Want to see the full answer?
Check out a sample textbook solution
Chapter 29 Solutions
SAPLINGPLUS F/BIOCHEM+ICLICKER REEF-CODE
- Polymerase inhibition. Cordycepin inhibits poly(A) synthesis at low concentrations and RNA synthesis at higher concentrations. NH2 H. он Cordycepin (3'-deoxyadenosine) a. What is the basis of inhibition by cordycepin? b. Why is poly(A) synthesis more sensitive than the synthesis of other RNAS to the presence of cordycepin? c. Does cordycepin need to be modified to exert its effect?arrow_forwardBackward? Bacteriophage T7 helicase moves along DNA in the 5'-to- 3'5'-to-3' direction. Other helicases have been reported to move in the 3'-to-5'3'-to-5' direction. Is there any fundamental reason why you would expect helicases to move in one direction or the other?arrow_forward. The following synthetic polynucleotide is synthesized and used as a template for peptide synthesis in a cell-free system from E. coli. .AUAUAUAUAUAUAU-. What polypeptide would you expect to be produced? Precisely what information would this give you about the code?arrow_forward
- please help me with this question. As this is a non-directional cloning, recombinant plasmids can contain an insert ligated into the vector in two different orientations. Provide two diagrams to illustrate the two potential recombinant plasmids, with the inserts ligated in opposite orientations. Include all RE sites and distances between sites on the diagram.arrow_forwardRNA is transcribed. Label the 5′ and 3′ ends of each strand. 17. The following sequence of nucleotides is found in a single-stranded DNA template: ATTGCCAGATCATCCCAATAGAT Assume that RNA polymerase proceeds along this template from left to right. a. Which end of the DNA template is 5′ and which end is 3′? b. Give the sequence and identify the 5′ and 3′ ends of the RNA transcribed from this template.arrow_forwardI am more confused. how about we start from begining, you post answers on here, and then we go from there? 1. Identify the open reading frame in the following DNA sequence, the protein that this gene encodes for, its function, and the source. 2. "Look carefully at the DNA sequence and identify the start site for transcription" 3. Click on the DNA sequence from the start site of transcription, select all of the sequence, and copy the sequence. Go to the National Center for Biotechnology Information (NCBI) website http://www.ncbi.nlm.nih.gov/. Click on BLAST on the right-hand side under “Popular Resources.” BLAST is a program that will allow you to find the protein sequence for the DNA sequence (gene) you submit. Next click on blastx (translated nucleotide protein). Paste the DNA sequence into the box under “Entry Query Sequence.” Scroll down and click BLAST. The search may take a few seconds; the page will keep updating until the search is completed. You do not need to enter any…arrow_forward
- please help me with thi question. What advantages do CRISPR‑Cas systems have over restriction enzymes and engineered nucleases for editing DNA? The options are attached. Multiple answers can be chosenarrow_forwardOriginal sequence: Consider the following coding 71 nucleotide DNA template sequence (It does not contain a translational start): 5’-GTTTCCCCTATGCTTCATCACGAGGGCACTGACATGTGTAAACGAAATTCCAACCTGAGCGGCGT GTTGAG-3’ Question: 4) In a mutant you discovered that the underlined nucleotide has been deleted. What would the resulting peptide sequence be? What type of mutation is this? 5’-GTTTCCCCTATGCTTCATCACGAGGGCACTGACATGTGTAAACGAAATTCCAACCTGAGCGGCGT GTTGAG-3arrow_forwardprotein. You create a mouse line with Cas9 under control of a brain-specific enhancer, while the short guide RNA complementary to the first exon of Gene Y is expressed in all tissues. You subsequently sequence Gene Y in both brain and liver tissue. What would expect in each tissue? You can assume that the CRISPRICas9 system will impact both copies of Gene Y in cells, and that the first exon of Gene Y is necessary for Gene Ys function. a. Liver: Functional Gene Y; Brain: Functional Gene Y b. Liver: Nonfunctional Gene Y; Brain: Funtional Gene Y c. Liver: Functional Gene Y; Brain: Nonfunctional Gene Y d. Liver: Nonfunctional Gene Y; Brain: Nonfunctional Gene Yarrow_forward
- Remember remove the introns! All introns start with GT and end with AG.arrow_forwardPart I. Structure-Function Relationships in Genes 1. Consider the "two-line model" of a gene shown below - each line represents one strand of a DNA double helix, and the transcription start site is indicated as +1. Use the two-line models provided when answering the following questions. 3' 5' +1 Assume that you know RNA polymerase will move to the right during transcription. On the diagram above, do the following: • Label "upstream" and "downstream" on this gene • Label where you would find the promoter min I • Draw a box where you would expect to find the TATA box • Draw a third line below the model representing the RNA transcript (label the ends!) • Label one of the DNA strands as the template strand 3' 2. Now, let's try that again! This time assume that you know RNA polymerase will move to the left during transcription. Repeat the same tasks as before on the diagram below: 5' 5' 3' +1 I I 5' 3'arrow_forwardgene. If the JM109 strain is transformed by the PBKSK plasmid, the strain will produce the B-galactosidase (from the lac gene) and will hydrolyze X-gal to produce the blue compound. Therefore, colonies that were transformed and contain the pBSKS wil you appear blue. IPTG & X-Gal & NO colonies Amp E. coli JM109 E. coli JM109 50 mM calcium chloride-15% glycerol lac lac lac IPTG & I Recovery X-Gal solution at -702C PBSKS White colonies E. coli JM109 E. coli JM109 ampR amp I amp lac lac Heat Shock Non-transformed 42°C E. coli JM109 E. coli JM109 amps amps lac lac IPTG & X-Gal lac I Recovery lac PBSKS BLUE colonies PBSKS ampRI (amp Transformed IPTG & X-Gal & BLUE colonies Amp Hypotheses: Circle the correct answer 1. If PBSKS is transformed into JM109 cells, colonies will be (able/not able) to grow in the presence of ampicillin. a. Why? _ 2. If PBSKS is transformed into JM109 cells, colonies in media with IPTG (will/will not) induce the production the B- galactosidase enzyme. a. Why?_ 3. If…arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
