
Concept explainers
The gel electrophoresis pattern in Figure 4.23 was determined by soaking the gel in a solution of ethidium bromide (EtBr). This is a fluorescent molecule with a planar structure:
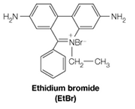
The flat molecule intercalates, or fits directly between two adjacent base pairs in a double helix. In doing so, it unwinds the double helix by 26o for each ethidium molecule bound.
a. If EtBr was added to relaxed, closed circular DNA, would you expect positive or negative supercoiling to occur? Explain.
b. If the circular DNA were nicked (had a single-stranded break) on one strand, what would be the effect on supercoiling?
c. If negatively supercoiled DNA is titrated with EtBr, the electrophoretic mobility decreases at first but then increases at higher EtBr concentrations. Explain.

Want to see the full answer?
Check out a sample textbook solution
Chapter 4 Solutions
Biochemistry: Concepts and Connections
- Explain how DNA-binding proteins can make sequence-specific contacts to a double-stranded DNA molecule without breaking the hydrogen bonds that hold the bases together. indicate how, through such contacts, a protein can distinguish a T-A from a C-G pair. indicate the parts of the nucleotide base pairs that could form noncovalent interactions— hydrogen bonds, electrostatic attractions, or hydrophobic interactions -with a DNA-binding protein.arrow_forwardThe two sides of the DNA double helix are connected by pairs of bases (adenine, thymine, cytosine, and guanine). Because of the geometric shape of these molecules, adenine bonds with thymine and cytosine bonds with guanine. The figure (Figure 1) shows the thymine-adenine bond. Each charge shown is ±e, and the H−N distance is 0.110 nm . Calculate the net force that thymine exerts on adenine. To keep the calculations fairly simple, yet reasonable, consider only the forces due to the O−H−N and the N−H−N combinations, assuming that these two combinations are parallel to each other. Remember, however, that in the O−H−N set, the O− exerts a force on both the H+ and the N−, and likewise along the N−H−N set. Express your answer in newtons. Is the net force attractive or repulsive?arrow_forwardIn the following sequence, a cytosine was deaminated and is now a uracil (underlined). 5’-GGTAUTAAGC-3’ a. Which repair pathway(s) could restore this uracil to cytosine? b. If the uracil is not removed before a DNA replication fork passes through, what will be the sequences of the two resulting double helices? Provide the sequences of both strands of both helices. Label the old and new strands and underline the mutation(s). c. Could the mismatch repair pathway fix the mutations you’ve indicated in part b? d. If the cell undergoes mitosis, and the replicated DNAs are distributed into the two daughter cells. Will 0, 1, or 2 daughter cells have a mutation in this sequence?arrow_forward
- To answer the prompts below, you will need to draw the chemical structure of the trinucleotide 5' - ACG - 3', labeling the 5' and 3' ends. Opposite this structure, draw the complementary trinucleotide to make a double-stranded DNA molecule. A. What is the complementary trinucleotide sequence from 5' to 3' (enter answer as e.g. CGA)? TCG B. How many non-covalent hydrogen bonds stabilize this structure? 8 C. How many covalent phosphate linkages stabilize this structure? 10 D. Which type of bond takes less energy to break? Phosphate linkage O Non-covalent hydrogen bondsarrow_forwardA closed circular duplex DNA has a 100-bp segment of alternating C and G residues. On transfer to a high salt solution, the segment undergoes a transition from the B conformation to the Z conformation. What is the change in its linking number, writhing number, and twist?arrow_forwardFor a linear B-DNA molecule of 50,000 kb, calculate (a) the contour length and (b) the length of the DNA as packaged in nucleosomes with linker histones present.arrow_forward
- Given the sequence shown below, write the complementary DNA sequence, using the base-pairing rules, as well as the directionality of the strands: 5'- CGAGGCTAGGTTAACCTG-3'arrow_forwardA circular, double-stranded DNA contains 2100 base pairs. The solution conditions are such that DNA has 10.5 bp/turn. (a) What is Lo for this DNA? (b) The DNA is found to have 12 left-handed superhelical turns. What is the superhelix density o?arrow_forwardA solution contains DNA polymerase and the Mg ²+ salts of dATP, dGTP, dCTP, and TTP. The following DNA molecules are added to aliquots of this solution. Which of them would lead to DNA synthesis? (a) A single-stranded closed circle containing 1000 nucleotide units. (b) A double-stranded closed circle containing 1000 nucleotide pairs. (c) A single-stranded closed circle of 1000 nucleotides base-paired to a linear strand of 500 nucleotides with a free 3' -OH terminus. (d) A double-stranded linear molecule of 1000 nucleotide pairs with a free 3’-OH group at each end.arrow_forward
- If a given double stranded DNA undergoes enzymatic hydrolysis targeting only the "b" side in the phosphodiester bond, what are the consequent nucleotide products of the hydrolysis of the 2 strands? (Please write the answers using only the letters corresponding to the bases with either the p or OH beside it to indicate the phosphate and OH groups respectively.) Sequence in one strand: TCGATCAGarrow_forwarda) It is known that double stranded DNA is denatured at low pH. pKa values should allow the determination of whether this is due to perturbation of the hydrogen bonding in A-T and/or G-C base pairs. The table gives values for the pKas of different protonated groups in the nucleobases.Nucleobase Position & pKa A N1, 3.5 G N7, 1.6; N1, 9.2 C N3, 4.2 T N3, 9.7a) Draw the A-T and G-C base pairs. - Label the bases with the one-letter code. – - Number the atoms in the rings and label the atom that attaches to the sugar. - Mark the groups that interact in normal…arrow_forwardThe base analog 5-bromouracil (5BU), which sterically resembles thymine, more readily undergoes tautomerization from its keto form to its enol form than does thymine. 5BU can be incorporated into newly synthesized DNA when it pairs with adenine on the template strand. However, the enol form of 5BU pairs with guanine rather than adenine. (a) Draw the 5BU · G base pair. (b) What type of mutation results?arrow_forward
 BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman
BiochemistryBiochemistryISBN:9781319114671Author:Lubert Stryer, Jeremy M. Berg, John L. Tymoczko, Gregory J. Gatto Jr.Publisher:W. H. Freeman Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman
Lehninger Principles of BiochemistryBiochemistryISBN:9781464126116Author:David L. Nelson, Michael M. CoxPublisher:W. H. Freeman Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY
Fundamentals of Biochemistry: Life at the Molecul...BiochemistryISBN:9781118918401Author:Donald Voet, Judith G. Voet, Charlotte W. PrattPublisher:WILEY BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305961135Author:Mary K. Campbell, Shawn O. Farrell, Owen M. McDougalPublisher:Cengage Learning BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON
Fundamentals of General, Organic, and Biological ...BiochemistryISBN:9780134015187Author:John E. McMurry, David S. Ballantine, Carl A. Hoeger, Virginia E. PetersonPublisher:PEARSON





