
Microbiology: An Introduction
12th Edition
ISBN: 9780321929150
Author: Gerard J. Tortora, Berdell R. Funke, Christine L. Case
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 9, Problem 2CAE
Using the restriction enzyme ECORI, the following gel electrophoresis patterns were obtained from digests of various DNA molecules from a transformation experiment. Can you conclude from these data that transformation occurred? Explain why or why not.
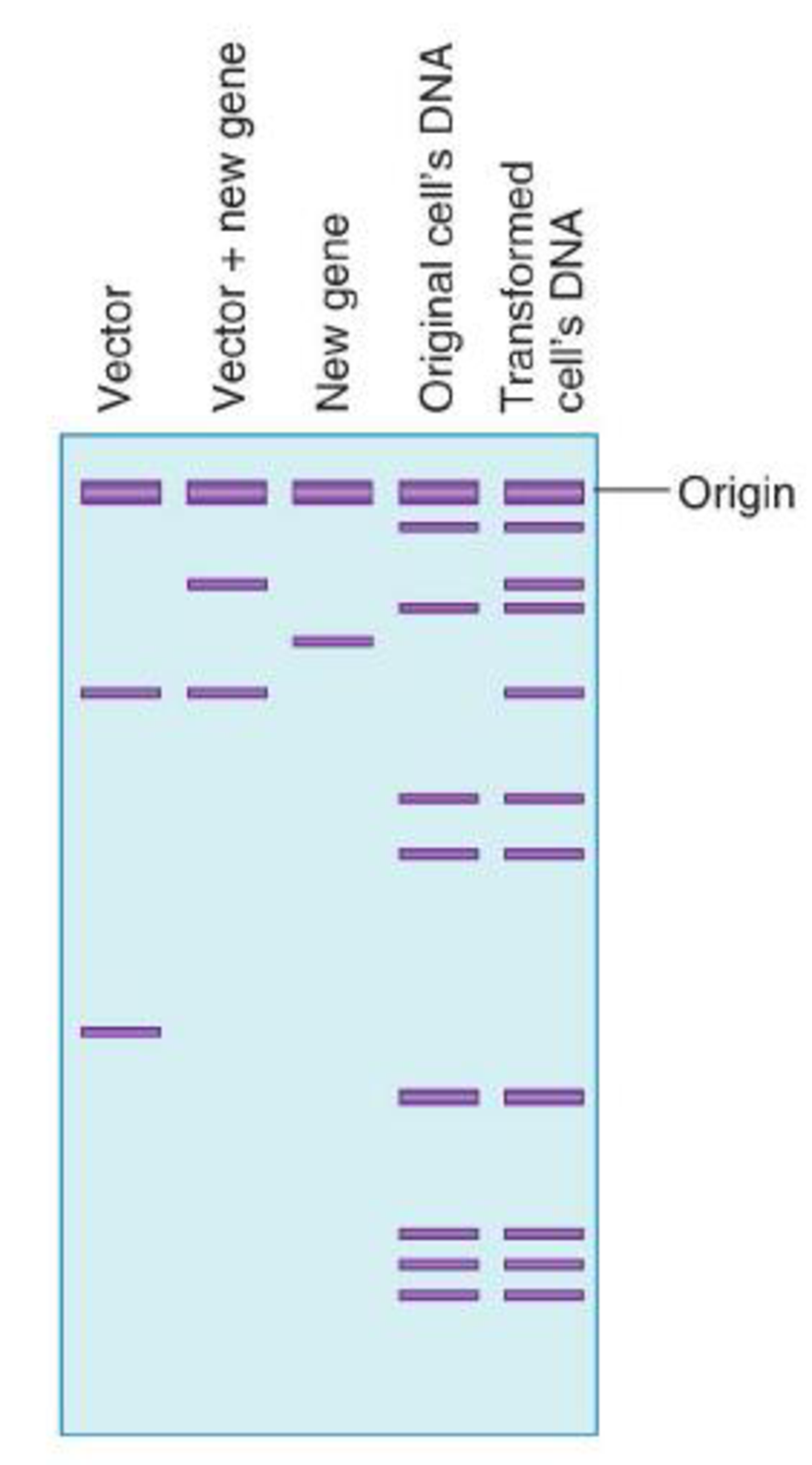
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
How would the results of this activity have been different if the DNA sequence you digested were circular instead of linear?
When restriction digests are performed and analyzed in the laboratory, one lane of the agarose gel is loaded with DNA ladders, which are DNA fragments of known sizes. What is the point of including these size standards?
Restriction sites are palindromic; that is, they read the same in the5' to 3' direction on each strand of DNA. What is the advantage ofhaving restriction sites organized this way?
A molecule of double-stranded DNA that is 5 million base pairs long has a base composition that is 62% G + C. How many times, on average, are restriction sites for the following restriction enzymes likely to be present in this DNA molecule?
a. HindIII (recognition sequence is AAGCTT)
Chapter 9 Solutions
Microbiology: An Introduction
Ch. 9 - Compare and contrast the following terms: a. cDNA...Ch. 9 - Differentiate the following terms. Which one is...Ch. 9 - Some commonly used restriction enzymes are listed...Ch. 9 - Suppose you want multiple copies of a gene you...Ch. 9 - Which enzyme makes the smallest fragment...Ch. 9 - Describe a recombinant DNA experiment in two or...Ch. 9 - List at least two examples of the use of rDNA in...Ch. 9 - You are attempting to insert a gene for saltwater...Ch. 9 - How does RNAi silence a gene?Ch. 9 - Prob. 10R
Ch. 9 - Restriction enzymes were first discovered with the...Ch. 9 - The DNA probe, 3-GGCTTA, will hybridize with which...Ch. 9 - Which of the following is the fourth basic step to...Ch. 9 - The following enzymes are used to make cDNA. What...Ch. 9 - If you put a gene in a virus, the next step in...Ch. 9 - You have a small gene that you want replicated by...Ch. 9 - Pieces of human DNA stored in yeast cells. a....Ch. 9 - A population of cells carrying a desired plasmid....Ch. 9 - Self-replicating DNA for transmitting a gene from...Ch. 9 - A gene that hybridizes with mRNA. a. antisense b....Ch. 9 - Design an experiment using vaccinia virus to make...Ch. 9 - Why did the use of DNA polymerase from the...Ch. 9 - The following picture shows bacterial colonies...Ch. 9 - Prob. 1CAECh. 9 - Using the restriction enzyme ECORI, the following...
Additional Science Textbook Solutions
Find more solutions based on key concepts
Identify each of the following reproductive barriers as prezygotic or postzygotic. a. One lilac species lives o...
Campbell Essential Biology with Physiology (5th Edition)
Physiology a. deals with the processes or functions of living things. b. is the scientific discipline that inve...
SEELEY'S ANATOMY+PHYSIOLOGY
Review the Chapter Concepts list on page 422. These all center on quantitative inheritance and the study and an...
Essentials of Genetics (9th Edition) - Standalone book
The correct term for production of offspring. Introduction: Reproduction is an important life process for most ...
Biology Illinois Edition (Glencoe Science)
Describe Mendels conclusions about how traits are passed from generation to generation.
Concepts of Genetics (11th Edition)
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Restriction endonuclease digestion of a DNA sequence yielded fragments of the following sizes: 1. 5.2 kb 2. 0.8 kb 3. 1.2 kb 4. 3.8 kb 5. 3.1 kb After gel electrophoresis, what would be the order in which these fragments would be found—the last fragment listed being furthest from the negative pole.arrow_forwardWhen circular DNA is sequenced, the nucleotide base pairs are numbered starting from a fixed position on the DNA, all the way around, usually in a clockwise manner. a DNA molecule that is 3133 base pairs long is digested with RsaI restriction enzyme recognition sites at base numbers 366, 1534, and 2207. What are the sizes of the DNA fragments that will be produced after the DNA is digested with RsaI?arrow_forwardIf restriction endonucleases are produced by bacteria within a host, why don’t these enzymes chew up the genomic DNA of their host? What is the role of DNA methyltransferase in this? Indicate the answerarrow_forward
- This plasmid was digested using different restriction enzymes whose sites have been mapped. The plasmid is 7896 base pairs long. This is a long question so u can count this as two or even three but please answer the question? Determine the size (base pairs) and number of fragments that would be produced if the plasmid was digested with the following enzymes: a) EcoRI b) BamHI c) HindIII d) EcoRI and HindIII e)EcoRI, HindIII, and BamHI *Hint- this is actually an EASY question, since the restriction map is already drawn for you!arrow_forwardHow would you expect your results to change if the enzyme did not have time to cut all of the sites? (Hint, remember that there are many pieces of the lambda DNA in the tube – draw yourself a simplified model where there are four restriction sites, and imagine what would happen if the enzyme only cut at one, two, three or all four, randomly, each time). How would you expect your results to change if you treated them with both EcoRI and HindIII in the same tube? Why might it be safer for the virus to have its DNA become circular when it enters the host, rather than remain linear?arrow_forwardIn making recombinant DNA molecules that combine restriction fragments from different organisms, researchers usually prefer restriction enzymes like BamHI or HindIII that generate fragments with “sticky ends” (ends with overhangs) rather than enzymes like HpaI or SmaI (Table 12.1) that generate fragments with “blunt ends” (ends without overhangs). Can you think of a reason for this preference?arrow_forward
- When performing cloning experiments, it is not always necessary to treat sources of DNA with the same restriction enzyme. For example, DNA treated with EcoRI can be combined with DNA from a treatment using FunII. Explain why this is possible.arrow_forwardIf a restriction enzyme cuts at this DNA sequence: TACGGAT, in general what is this sequence called?arrow_forwardA molecule of double-stranded DNA that is 5 million base pairs long has a base composition that is 62% G + C. How many times, on average, are restriction sites for the following restriction enzymes likely to be present in this DNA molecule? a. HpaII (recognition sequence is CCGG)arrow_forward
- If restriction endonucleases are produced by bacteria within a host, why don’t these enzymes chew up the genomic DNA of their host? What is the role of DNA methyltransferase in this?arrow_forwardA small DNA molecule was cleaved with several different restriction nucleases, and the size of each fragment was determined by gel electrophoresis. The following data were obtained.arrow_forwardRestriction enzymes and DNA ligase play essential roles in DNA cloning. How is it that a bacterium that produces a restriction enzyme does not cut its own DNA? Describe some general features of restriction sites.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning
Biology Today and Tomorrow without Physiology (Mi...BiologyISBN:9781305117396Author:Cecie Starr, Christine Evers, Lisa StarrPublisher:Cengage Learning Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

Biology Today and Tomorrow without Physiology (Mi...
Biology
ISBN:9781305117396
Author:Cecie Starr, Christine Evers, Lisa Starr
Publisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning
Bacterial Genomics and Metagenomics; Author: Quadram Institute;https://www.youtube.com/watch?v=_6IdVTAFXoU;License: Standard youtube license