
Hepatocellular carcinoma is the most frequent form of liver cancer. In a patient with heritable hepatocellular carcinoma, formation of the tumor was associated with eight genetic alterations affecting two different oncogenes and three different tumor-suppressor genes. These alterations are:
| i. | Mitotic recombination |
| ii. | A deletion of a chromosomal region |
| iii. | Trisomy |
| iv. | A duplication of a chromosomal region |
| v. | Uniparental disomy (see Fig. 20.24) |
| vi. | A point mutation |
| vii. | Another point mutation |
| viii. | Yet another point mutation |
For parts a–c below, supply all possible correct answers from the preceding list. Remember that the majority of point mutations are loss-of-function mutations.
| a. | Which of the mutations from the preceding list is likely to affect a proto-oncogene? |
| b. | Which of the mutations from the preceding list is likely to involve a tumor-suppressor gene? |
| c. | Which of the mutations from the preceding list involves copy-neutral loss-of-heterozygosity (that is, a loss-of-heterozygosity in which the genomes of the cancerous cells still have two copies of the gene in question, whether or not those copies are functional)? |
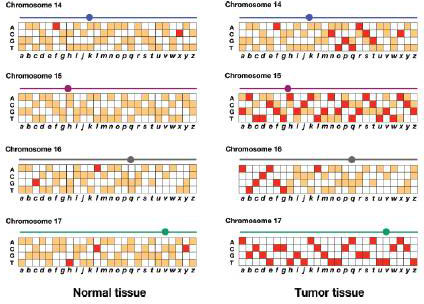
Genomic DNA is prepared from normal white blood cells and from a biopsy of the tumor in this patient. These genomic DNAs are prepared as fluorescent probes that are each hybridized to an ASO microarray of polymorphisms in the human genome (review Figs. 11.16 and 11.17). The results for SNPs a–z on chromosomes 14, 15, 16, and 17 are shown in the accompanying figure. Red and orange represent different levels of fluorescence.
| d. | Based on the microarray data, provide the most accurate localization of the first five types of genetic alterations in the list (i–v). For example, if an alteration involves markers a–e of chromosome 15, write 15a–e. |
| e. | As precisely as possible, indicate the location of the mitotic recombination event involved in the genesis of this cancer |
| f. | If these data allow you to map any of the three cancer-promoting point mutations, provide the most accurate mutation location(s) possible. |
| g. | Of all the genetic alterations i–viii, for which one do you see clear-cut evidence that the mutation or other event was inherited from a parent of the patient? |
| h. | For a tumor-suppressor gene to play a role in cancer, normally both of the copies in the tumor cells must be nonfunctional. For each of the three tumor-suppressor genes contributing to the cancer in this patient, provide a scenario explaining which two hits (i–viii in the list, with vi–viii equivalent) could be responsible, the order in which the hits must have occurred, and whether the hits in question could be inherited or could have occurred somatically. |
Want to see the full answer?
Check out a sample textbook solution
Chapter 20 Solutions
EBK GENETICS: FROM GENES TO GENOMES
- Tumor suppressor genes and oncogenes are implicated in carcinogenesis. However, one can predict whether a gene potentially encodes for a protein that influences carcinogenesis by examining their mutational profile. You sequence the genome of 4 cancers and identify 3 genes of interest. Which of the following genes has the best potential to an oncogene? Tumor 1 Tumor 2 Tumor 3 Tumor 4 Gene A S24F, N465T R33T T345S, G366R P367E, P368Y Gene B S34R, F360I S34R V254I S34E, T67Y Gene C S24F, I322E C255I, E344D S34E, P367Earrow_forwardDescribe the steps by which the TP53 gene responds to DNA damage and/or cellular stress to promote cell-cycle arrest and apoptosis. Given that TP53 is a recessive gene and is not located on the X chromosome, why would people who inherit just one mutant copy of a recessive tumor-suppressor gene be at higher risk of developing cancer than those without the recessive gene?arrow_forwardThe following chromosomal aberration is found in nearly 90-95% of all patients who have chronic myelogenous leukemia. This is because the change brings the BCR and ABL genes in close proximity. BCR is responsible for cell growth, and ABL is a proto-oncogene... this favors uncontrolled growth. Which is the most accurate description of the aberration... chio moso me 9 Philad elphia chromosome chromosome 22 BCR ABL 22q11.2 (BCR) 9934 1 (ABL) Deletion Translocation Inversion Duplication DELL O O O Carrow_forward
- Explain why mutations in tumor suppressor genes are recessive (both copies of the gene must be defective for the regulation of cell division to be defective), whereas mutations in oncogenes are dominant.arrow_forwardPart 3C: Below is data from a pedigree trace of patients affected with mouse model of pancreatic cancer, and below are the observations regarding a new gene called 318 that is found on chromosome 5. Patient II-4 has two copies of loss of function variant alleles (or is homozygous for loss-of-function variant 318 allele). Patient I-2 has one copy of loss of function variant and one copy of unaffected allele (is heterozygous for loss-of-function variant 318 allele). Patient II-1 has two copies of unaffected allele (or is homozygous for unaffected 318 allele). |-1 1-2 Il-1 Il-2 Il-3 |l-4 II-5 II-1 III-2 III-3 Based upon data above, what is the mode of inheritance for mouse model for pancreatic cancer? Why?arrow_forwardThe propensity to develop retinal cancer can run in pedigrees of certain families. You have learned about the role of the retinoblastoma (Rb) protein in the cell cycle. A heterozygous individual (genotype = Rbm / Rb+) that inherits a mutant allele of the Rb gene (Rbm) as well as a normal allele of the Rb gene (Rb+) is at significant risk for developing cancer of the retina. The best explanation for this is that… A. the Rb(m) allele is dominant to the Rb(+) allele. B. a mutation in the Rb(+) allele can result in a Rb(m) / Rb(m) genotype in certain cells. C. the retina is frequently exposed to UV rays D. the Rb(m) allele directly converts the Rb(+) allele to another Rb(m) allele. E. the cells of the retina carry more than two copies of the Rb genearrow_forward
- Match the following statements to the type of gene they describe. ✓Encode proteins that promote cell cycle progression ✓Encode proteins that regulate cell cycle checkpoints ✓Mutations are loss-of-function and recessive ✓Mutations are gain-of-function and dominant ✓RB1 is an example of this type of gene A.proto-oncogene B.tumor suppressor genearrow_forwardWhich of the following mutations will result in cancer? a. homozygous recessive mutation in a tumor-suppressor gene coding for a nonfunctional protein b. dominant mutation in a tumor-suppressor gene in which the normal protein product is overexpressed c. homozygous recessive mutation in which there is a deletion in the coding region of a proto-oncogene, leaving it nonfunctional d. dominant mutation in a proto-oncogene in which the normal protein product is overexpressedarrow_forwardSome cancers are consistently associated with the deletion of a particularpart of a chromosome. Does the deleted region contain an oncogene or atumor-suppressor gene? Explain.arrow_forward
- Please explain the relationship between proto-oncogenes and the cell cycle. Then describe three different types of alterations in the genome in relation to proto-oncogenes that alter the cell cycle and cause cancer.arrow_forwardMutations in proto-oncogenes that turn them into oncogenes tend to be dominant, while cancer-causing mutations in tumor suppressor genes tend to be recessive. Please explain why.arrow_forwardThere are three broad categories of cancer-related genes: proto-oncogenes, tumor suppressor genes, and DNA stability/repair genes. Define each of these categories and indicate which one you think the RB1 gene belongs to and why.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
