
Concept explainers
An electrophoretic mobility shift assay can be used to study the binding of proteins to a segment of DNA. In the experiment shown here, an EMSA was used to examine the requirements for the binding of RNA polymerase II (from eukaryotic cells) to the promoter of a protein-encoding gene. The assembly of general transcription factors and RNA polymerase II at the core promoter is described in Chapter 12 (Figure 12.14). In this experiment, the segment of DNA containing a promoter sequence was 1100 bp in length. The fragment was mixed with various combinations of proteins and then subjected to an EMSA.
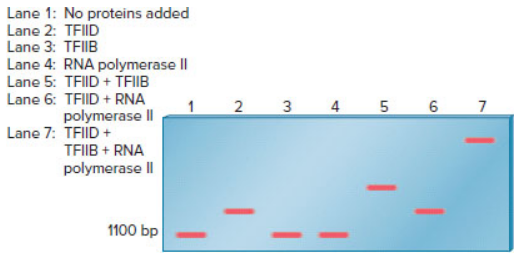
Explain which proteins (TFIID, TFIIB, or RNA polymerase II) are able to bind to this DNA fragment by themselves. Which transcription factors (i.e., TFIID or TFIIB) are needed for the binding of RNA polymerase II?
Want to see the full answer?
Check out a sample textbook solution
Chapter 21 Solutions
GENETICS:ANALYSIS+PRIN.(LL)-W/ACCESS
- SR proteins are essential to proper spliceosome assembly and are known to take part in the regulation of alternative splicing. Surprisingly, the role of SR proteins in splice-site selection and alternative splicing is affected by the promoter used for the transcription of the pre-mRNA. For example, through genetic engineering, RNA polymerase II promoters that have somewhat different sequences can be created. When pre-mRNAs with exactly the same sequences are transcribed from two different RNA polymerase II promoters that differ slightly in sequence, which promoter is used can affect how the pre-mRNA is spliced. Propose a mechanism by which the DNA sequence of an RNA polymerase II promoter could affect alternative splicing of pre-mRNA.arrow_forwardGal4 is a transcription factor that activates transcription of galactose metabolism genes in yeast. These genes are ‘turned on’ when yeast cells need to metabolize galactose. To identify promoter sequences necessary for regulation of transcription of GAL1, reporter gene fusions were made and introduced into yeast cells. Deletions of GAL1 promoter were cloned upstream of LacZ gene. β-Galactosidase activity was measured in presence of galactose. Shown below is a representation of the results obtained. In the diagrams below (not to scale!): • Construct 1 contains ~ 130bp of the promoter, which is predicted to have all the predicted/putative proximal promoter elements (indicated by the solid boxes) needed to regulate transcription of GAL1.• The stippled box is the core promoter.• The arrow represents the transcriptional start site for the reporter gene Lac Z• Number of + signs represents level of transcription• Star represents a mutation in DNA sequence at that location (few nucleotides…arrow_forwardRNA knockdown has become a powerful tool in the arsenal of methods used to repress gene expression. Briefly describe how gene expression can be knocked down. What effect would introducing siRNAs to TSC1 have on human cells?arrow_forward
- Here is the model for TfR mRNA regulation (see Fig.), for your convenience. Orange protein (IRP or IREBP), when bound to RNA, protects mRNA from degradation. When Fe is bound to the protein, it cannot bind the mRNA. If you made a radioactive DNA probe specific to TfR mRNA and performed a Northern blot analysis (to detect mRNA) from cells treated with high iron (Fe) concentration what would you expect to see? (a) Low iron RNase A (b) High iron O High levels of RNase O Low levels of TfR mRNA Fe RNase O The result would be the same as untreated cells Degradation O High levels of IRP (or IREBP - iron responsive element binding protein) O High levels of TfR mRNAarrow_forwardWhich serine in the CTD of RNA polymerase II is phosphorylated to recruit enzymes to add the 5’ guanosine cap? How is this phosphorylation different than the CTD phosphorylation in promoter clearance?arrow_forwardExplain how a transcriptional fusion to a reporter gene, combined with site directed mutagenesis, can indicate key nucleotides in a regulatory region. Name one in vitro technique that can be used to further confirm the significance of these regions.arrow_forward
- Suppose you had isolated a new transcription factor and wanted to know which genes this protein might regulate. Is there any way that you could use a cDNA microarray of the type shown in the picture to approach this question?arrow_forwardThe following diagram show what is required for an active promoter of a gene of interest, where:A1 = Activator 1A2 = Activator 2Med = MediatorRep = Repressor Based on the following data, predict: Chromatin conformation Methylation state of the proximal promoter If protein A1 is present or absent If protein A2 is present or absent If the mediator is present or absent If the repressor is absence or present If this gene is likely to be transcribed or notPlease note that this is an "all or nothing" bonus question, and no partial credit will be awarded. Selecting all answers will result in zero points awarded. Selecting at least one incorrect answer will result in zero points being awarded. Question 28 options: Euchromatin Heterochromatin Methylated promoter Unmethylated promoter Activator A1 present Activator A1 absent Activator A2 present…arrow_forwardWhich of the following statements is true regarding the lys2-128d reporter? Select all that apply. a.) Mutants that activate this reporter are likely to have mutations in PIC components. b.) This reporter is sensitive to changes in chromatin maintenance because the start site is located within the ORF. c.) Mutants that activate this reporter cause downstream shifts in transcription start site selection. d.) This reporter is being used to help us rule out mutations that are likely impacting chromatin structure.arrow_forward
- The IMD2 promoter contains three upstream transcription start sites (TSS) that are utilized under high GTP conditions and a single downstream TSS (-106) that is normally only utilized under low GTP conditions. In a wild type cell, expression of IMD2 mRNA only occurs if transcription initiates from the -106 TSS. In 300 words or less, describe: 1.) The normal function of Ssl2, and 2.) why a mutation in Ssl2, that increases its catalytic rate, would allow expression of the IMD2 ORF under high GTP conditions. (Conditions under which the IMD2 ORF is NOT expressed in the wild type.)arrow_forwardBacterial DNA containing an operon encoding three enzymes is introduced into chromosomal DNA in yeast (a eukaryote) in such a way that it is properly flanked by a promoter and a transcriptional terminator. The bacterial DNA is transcribed and the RNA correctly processed, but only the protein nearest the promoter is produced. Can you suggest why?arrow_forwardIf the binding activity was purified, suggest two experimental approaches that could be taken to verify that this factor is in fact a transcription factor? Explain.arrow_forward
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
