
Biochemistry: Concepts and Connections (2nd Edition)
2nd Edition
ISBN: 9780134641621
Author: Dean R. Appling, Spencer J. Anthony-Cahill, Christopher K. Mathews
Publisher: PEARSON
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 8, Problem 11P
The following data describe the catalysis of cleavage of peptide bonds small peptides by the enzyme elastase.
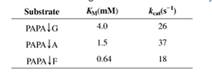
The arrow indicates the peptide bond cleaved each case.
a. If a mixture of these three substrates was presented to elastase with the concentration of each peptide equal to 0.5 mM, which would be digested most rapidly? Which most slowly? (Assume enzyme present in excess.)
b. On basis of these data, suggest what features of amino acid sequence dictate the specificity of proteolytic cleavage by elastase.
c. Elastase is closely related to chymotrypsin. Suggest two kinds of amino acid residues you might expect to find in or near the active site.
Expert Solution & Answer
Want to see the full answer?
Check out a sample textbook solution
Students have asked these similar questions
Considering the chemical characteristics of the amino acids valine and glutamic acid (see Figure 5.14), propose a possible explanation for the dramatic effect on protein function that occurs when valine is substituted for glutamic acid.
Remembering that the amino acid side chains projecting from each polypeptide backbone in a β sheet point alternately above and below the plane of the sheet, consider the following protein sequence: Leu-Lys-Val-Asp-Ile-Ser-Leu-Arg- Leu-Lys-Ile-Arg-Phe-Glu. Do you find anything remarkable about the arrangement of the amino acids in this sequence when incorporated into a β sheet? Can you make any predictions as to how the β sheet might be arranged in a protein?
you were given an unknown peptide. Explain in detail how you would determine the amino acid sequence of this peptide by means of Edman degradation. (You can do this by showing how you would determine the N-terminal amino acid of the peptide shown below in figure 2 with Edman degradation.
Chapter 8 Solutions
Biochemistry: Concepts and Connections (2nd Edition)
Ch. 8 - Prob. 1PCh. 8 - The enzyme urease catalyzes the hydrolysis of urea...Ch. 8 - An enzyme contains an active site aspartic acid...Ch. 8 - The folding and unfolding rate constants for a...Ch. 8 - In some reactions, in which a protein molecule is...Ch. 8 - Would you expect an “enzyme” designed to bind to...Ch. 8 - The initial rate for an enzyme-catalyzed reaction...Ch. 8 - a. If the total enzyme concentration in Problem 7...Ch. 8 - Prob. 9PCh. 8 - Prob. 10P
Ch. 8 - The following data describe the catalysis of...Ch. 8 - At 37 oC, the serine protease subtilisin has kcat...Ch. 8 - The accompanying figure shows three...Ch. 8 - The steady-state kinetics of an enzyme are studied...Ch. 8 - The same enzyme as in Problem 14 is studied in the...Ch. 8 - Enalapril is an anti-hypertension “pro-drug"...Ch. 8 - Initial rate data for an enzyme that obeys...Ch. 8 - Prob. 18PCh. 8 - Suggest the effects of each of the following...Ch. 8 - The inhibitory effect of an uncompetitive...Ch. 8 - Prob. 21PCh. 8 - Prob. 22PCh. 8 - Prob. 23PCh. 8 - In kinetics experiments, the hydrolysis of the...
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biochemistry and related others by exploring similar questions and additional content below.Similar questions
- b. Compounds A, B, C, and D are known to be intermediates in the pathway for production of protein E. To determine where the block in protein-E production occurred in each individual, the various intermediates were given to each individuals cel Is in culture. After a few weeks of growth with the intermediate, the cells were assayed for the production of protein E. The results for each individuals cells are given in the following table. A plus sign means that protein E was produced after the cells were given the intermediate listed at the top of the column. A minus sign means that the cells still could not produce protein E even after being exposed to the intermediate at the top of the column. Denote the point in the pathway in which each individual is blocked.arrow_forwarda. Compounds A, B, C, and D are known to be intermediates in the pathway for production of protein E. To determine where the block in protein-E production occurred in each individual, the various intermediates were given to each individuals cel Is in culture. After a few weeks of growth with the intermediate, the cells were assayed for the production of protein E. The results for each individuals cells are given in the following table. A plus sign means that protein E was produced after the cells were given the intermediate listed at the top of the column. A minus sign means that the cells still could not produce protein E even after being exposed to the intermediate at the top of the column. Draw the pathway leading to the production of protein E.arrow_forwardThe following data describe the catalysis of cleavage of peptide bonds in small peptides by the enzyme UTSAse (the arrow indicates the peptide bond cleaved in each case). Substrate Km(mM) kcat(s-1) PAPA↓G 4.0 26 PAPA↓A 1.5 37 PAPA↓F 0.64 18 what features of amino acid sequence dictate the specificity of the proteolytic cleavage? Large hydrophilic R-groups Large hydrophobic R-groups Neutral R-groups Small hydrophilic R-groups Large hydrophobic R-groups Negatively charged R-groups Positively charged R-groupsarrow_forward
- An enzyme that catalyzes disulfide– sulfhydryl exchange reactions, called protein disulfide isomerase (PDI), has been isolated. PDI rapidly converts inactive scrambled ribonuclease into enzymatically active ribonuclease. In contrast, insulin is rapidly inactivated by PDI. What does this important observation imply about the relation between the amino acid sequence of insulin and its threedimensional structure?arrow_forwardA heptapeptide when treated with trypsin produced two peptides. T1 (D, G, Y) and T2 (K, F, V, A). When the heptapeptide was treated with chymotrypsin, three peptides were produced: CT1 (K,,Y, G), CT2 (F,A, V), and CT3 (D). The sequences of these peptides is not known, however. When the peptide was treated with Sanger’s Reagent and hydrolyzed, DNP-K and DNP-A were recovered. What is the amino acid sequence of the heptapeptide?arrow_forwardA peptide composed of 40 amino acids (aa) was isolated from the venom of the Gila Monster(Heloderma suspectum, then submitted to various biochemical and chemical treatments.1) After a treatment with a highly specific chymotrypsin, four peptides were generated:- peptide A (6 aa)- peptide B (16 aa)- peptide C (3 aa)- , and peptide D (15 aa)2) Treatment with carboxypeptidases A or B resulted in the release of:- phenylalanine for peptides A and B- tryptophan for peptide C- glycine for peptide D and the 40 amino acids venom peptide3) One cycle of Edman degradation either for peptide A or for the 40 amino acidspeptide gave the exact PTH derivative indicated below: 4) The 40 amino acids peptide treated with Asp-N endopeptidase followed by massspectrometry MS/MS revealed the following composition:- two molecules of glutamic acid,- various peptides indicated below sorted by masses:HG,DLSKQM,EGTFTS,EAVRLFI,EWLKNGGPSSGAPPPSG Information- Chymotrypsin (endopeptidase) hydrolyzes peptide bonds on…arrow_forward
- When performing his experiments on protein refolding, Christian Anfinsen obtained a quite different result when reduced ribonuclease was reoxidized while it was still in 8 M urea and the preparation was then dialyzed to remove the urea. Ribonuclease reoxidized in this way had only 1% of the enzymatic activity of the native protein. Why were the outcomes so different when reduced ribonuclease was reoxidized in the presence and absence of urea?arrow_forwardCarboxypeptidase, which sequentially removes carboxyl-terminal amino acid residues from its peptide substrates, is a single polypeptide of 307 amino acids. The two essential catalytic groups in the active site arefurnished by Arg 145 and Glu 270 .(a) If the carboxypeptidase chain were a perfect α helix, how far apart (in Å) would Arg 145 and Glu 270 be? (b) Explain how the two amino acid residues can catalyze a reaction occurring in the space of a few angstromsarrow_forwardusing, 3’ TGAGGCGCTAGGCCAAGCGGTAAGGATGCATGGTCGTGGTAG , What would be the resultant type of error on the amino acid chain?arrow_forward
- PLP is a cofactor for a number of enzymes involved in amino acid metabolism. Give an example of reaction in which PLP participates in cleavage of the a, b, and c bonds of an amino acid, as diagrammed in shown Fig.arrow_forwardWhat is the expected molecular weight of the protein encoded by the sequence provided here: ATGGCGCACGCTGGGAGAACAGGGTACGATAACCGGGAGATAGTGATGAAGTACATCCATTATAAGCTGTCGCAGAGGGGCTACGAGTGGGATGCGGGAGATGTGGGCGCCGCGCCCCCGGGGGCCGCCCCCGCACCGGGCATCTTCTCCTCCCAGCCCGGGCACACGCCCCATCCAGCCGCATCCCGGGACCCGGTCGCCAGGACCTCGCCGCTGCAGACCCCGGCTGCCCCCGGCGCCGCCGCGGGGCCTGCGCTCAGCCCGGTGCCACCTGTGGTCCACCTGACCCTCCGCCAGGCCGGCGACGACTTCTCCCGCCGCTACCGCCGCGACTTCGCCGAGATGTCCAGCCAGCTGCACCTGACGCCCTTCACCGCGCGGGGACGCTTTGCCACGGTGGTGGAGGAGCTCTTCAGGGACGGGGTGAACTGGGGGAGGATTGTGGCCTTCTTTGAGTTCGGTGGGGTCATGTGTGTGGAGAGCGTCAACCGGGAGATGTCGCCCCTGGTGGACAACATCGCCCTGTGGATGACTGAGTACCTGAACCGGCACCTGCACACCTGGATCCAGGATAACGGAGGCTGGGTAGGTGCACTTGGTGATGTGAGTCTGGGCTGAarrow_forwardIn some proteins, the side chain of serine appears to undergo ionization. Explain why ionization would be facilitated by the presence of an aspartate residue nearby.arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning
Biomolecules - Protein - Amino acids; Author: Tutorials Point (India) Ltd.;https://www.youtube.com/watch?v=ySNVPDHJ0ek;License: Standard YouTube License, CC-BY