
Concept explainers
Interpretation: An explanation regarding the cleavage of the given amide substrate A by chymotrypsin reveals no burst by using stopped-flow kinetic methods is to be stated.
Concept introduction: The organic compounds that contain
Proteolytic enzymes commonly known as proteases are used for the cleavage of protein. Chymotrypsin is an enzyme that is used for the cleavage of protein through hydrolysis reaction.
Answer to Problem 1P
The cleavage of the given amide substrate, A, by chymotrypsin reveals no burst because the acyl-enzyme intermediate formation occurs at a slower rate as compared to the hydrolysis for the given amide substrate.
Explanation of Solution
Given:
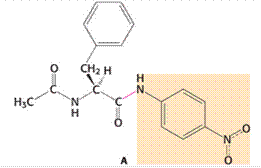
Chymotrypsin is an enzyme that is used for the cleavage of protein through hydrolysis reaction. In case of large hydrophobic amino acids, the cleavage of the peptide bond occurs particularly at the carboxyl-terminal side. In the given amide substrate A, the cleavage by chymotrypsin shows no burst because the hydrolysis for this substrate of amide is faster than the generation of the acyl-enzyme intermediate. In other words, the acyl-enzyme intermediate formation occurs at a slower rate as compared to the hydrolysis for the given amide substrate. Hence, no burst is found. In case of ester substrates, the acyl-enzyme intermediate formation occurs at a faster rate.
Want to see more full solutions like this?
- The nitrogenase complex can reduce molecules other than N2. Provide the structures for the products for each of the following substrates (real and hypothetical) that contain triple bonds: hydrogen cyanide, dinitrogen, and acetylene.arrow_forwardSubstrate KM (M) N-Acetylvaline ethyl ether 8.8 X 10 -2 N-Acetyltyrosine ethyl ether 6.6 X 10-4 Which substrate has the higher apparent affinity for the enzyme? Explain. Which substrate is likely to give a higher value for Vmax?arrow_forward(b) The values of kinetic parameters for a variety of synthetic ester and peptide substrates ofa- chymotrypsin are compared for the reaction scheme below where ES is the Michaelis complex, ES' E+S Substrate K₁ N-Ac-Trp–OC₂H5 N-Ac-Phe-OC2H5 N-Ac-Leu-OC2H5 N-Ac-Phe-CONH2 N-Ac-Tyr-p-nitro-anilide ES K₂ 3.5 13.0 3.2 1.7 K-₁ is the acylenzyme, P₁ is the first product to be released, P2 is the second product, k2 is the acylation rate constant, k3 is the deacylation rate constant, and kcat = k2 k3/( k2 + k³). In the table below, N- Ac = N-acetyl; -CONH₂ = carboxamide; -OC₂H5 = ethyl ester; p-nitroanilide = −NH-C6H4-NO2 simulating a peptide group. ES' K2 (S-¹) K3 (S-1) 0.84 2.2 0.19 P₁ K3 0.073 H → E+ P2 Kcat (S-1) 0.82 1.9 0.18 0.070 0.038 KM (MM) 0.08 1.3 4.2 24.0 0.35 1 Kcat/ KM (mM-¹ s¯¹) 10.3 1.5 0.04 0.003 0.11 The value of kcat for N-Ac-Phe-OC2H5 is two-fold greater than that for the L-tryptophanyl analog and more than 10-fold greater than the value of kcat for the ester substrate…arrow_forward
- Based on your understanding of the structure and function of the enzyme chymotrypsin, explain why proteolytic activation by Trypsin in the intestine is required for chymotrypsin to bind its substrates with high affinity. Your answer should include a description of the amino acid residue or residues involved AND a kinetic experiment that suggested this.arrow_forward(i) Describe the mechanism of chymotrypsin in cleaving a peptide bond, highlighting the roles of the catalytice triad for the two phases of the catalytic reactions. Explain the significance of the oxyanion hole for the catalysis. (ii) All serine proteases contain the catalytic triad and these amino acids are positioned in the exact same conformation. Since this is true, why do trypsin and chymotrypsin have such different substrate specificity? What features of the enzyme allow for this situation?arrow_forwardUsing the ActiveModel for aldose reductase, describe the structure of the TIM barrel motif and the structure and location of the active site.arrow_forward
- Extending the Mechanism of Methylmalonyl-CoA Mutase to Similar Reactions Based on the mechanism for the methylmalonyl-CoA mutase (see problem 14), write reasonable mechanisms for the following reactions shown.arrow_forwardThe substitution of His 64 of carbonic anhydrase II with Ala results in a sharp decrease in the activity of the enzyme in HEPES buffer (molecular weight of HEPES = 238.3 g/mol). However, increasing concentrations of imidazole (molecular weight = 68.1 g/mol) restores the reaction rate close to that of the wild-type enzyme. Propose an explanation for these results.arrow_forwardLysine degradation requires removal of two amino groups. Removal of itsε-amino group gives a-aminoadipic semialdehyde. This product is thendegraded to acetoacetate by the same chemical strategy used to degrade thebranched-chain amino acids. Draw the proposed intermediates in this pathway (Hint: see Figure 18.14).arrow_forward
- Shown below is a substrate for a Trypsin. Draw the mechanism for this serine protease using the artificial substrate. Be sure to draw the catalytic triad, and show the role of the oxyanion hole. Draw the complete structure of every intermediate and product and PUSH ARROWS!!!!! Do not abbreviate structures using R and R' H₂N _N_CH. сно CH₂ CH₂ CH₂ NH d=19H₂ NH₂ O CH- H₂C HN O CHarrow_forwardWhat will be the products? Which enzyme will catalyzes this reaction?arrow_forwardLysozyme is an antibacterial enzyme found in animals. Residues Asp 101 and Arg 114 are required for efficient catalysis, although they are located at some distance from the active site Glu 35 and Asp 52. Substituting Ala for either Asp 101 or Arg 114 does not significantly alter the enzyme's tertiary structure, but significantly reduces its catalytic activity. Explain.arrow_forward
 BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
BiochemistryBiochemistryISBN:9781305577206Author:Reginald H. Garrett, Charles M. GrishamPublisher:Cengage Learning
