
Genetics: From Genes to Genomes
6th Edition
ISBN: 9781259700903
Author: Leland Hartwell Dr., Michael L. Goldberg Professor Dr., Janice Fischer, Leroy Hood Dr.
Publisher: McGraw-Hill Education
expand_more
expand_more
format_list_bulleted
Concept explainers
Textbook Question
Chapter 10, Problem 21P
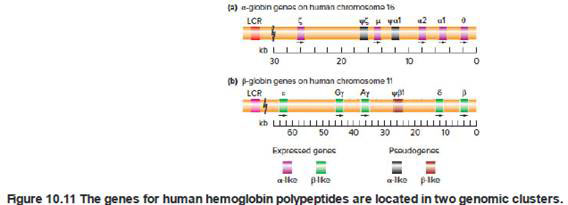
Chimpanzees have a set of hemoglobin genes very similar to the set in humans that was shown in Fig. 10.11. For example, the genomes of both species have α1, α2, β, Gγ, Aγ, δ, ε, and ζ genes.
| a. | Of the human and chimpanzee hemoglobin genes, which would be considered homologous? Which paralogous? Which orthologous? |
| b. | When comparing genomes, geneticists would usually like to know which genes are the most likely to perform similar if not identical functions in different species. This determination can be somewhat complicated in the case of gene families. Would paralogous genes or orthologous genes be more likely to be functionally equivalent? Explain. |
| c. | Which gene would have the greatest degree of |
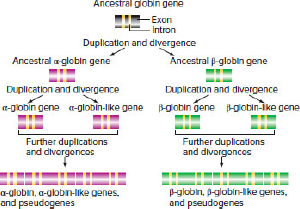
| d. | Rationalize the pattern of hemoglobin genes in the two species with the existence of duplication and divergence events among the hemoglobin genes depicted in Fig. 10.12. |
Expert Solution & Answer
Trending nowThis is a popular solution!

Students have asked these similar questions
In a study of a muscle disorder, several affected families exhibited vision problems, muscle weakness, and deafness (M. Zeviani et al. 1990. American Journal of Human Genetics 47:904–914). Analysis of the mtDNA from affected members of these families revealed that large numbers of their mtDNA molecules possessed deletions of varying lengths. Different members of the same family and even different mitochondria from the same person possessed deletions of different sizes, so the underlying defect appeared to be a tendency for the mtDNA of affected persons to have deletions. A pedigree of one of the families studied is shown below. The researchers concluded that this disorder is inherited as an autosomal dominant trait, and they mapped the diseasecausing gene to a position on chromosome 10 in the nucleus.
Q. What characteristics of the pedigree rule out inheritance of a trait encoded by a gene in the mtDNA as the cause of this disorder?
In a study of a muscle disorder, several affected families exhibited vision problems, muscle weakness, and deafness (M. Zeviani et al. 1990. American Journal of Human Genetics 47:904–914). Analysis of the mtDNA from affected members of these families revealed that large numbers of their mtDNA molecules possessed deletions of varying lengths. Different members of the same family and even different mitochondria from the same person possessed deletions of different sizes, so the underlying defect appeared to be a tendency for the mtDNA of affected persons to have deletions. A pedigree of one of the families studied is shown below. The researchers concluded that this disorder is inherited as an autosomal dominant trait, and they mapped the diseasecausing gene to a position on chromosome 10 in the nucleus.
Q. Explain how a mutation in a nuclear gene might lead to deletions in mtDNA.
Two diploid species of closely related frogs, which we will callspecies A and species B, were analyzed with regard to the genesthat encode an enzyme called hexokinase. Species A has two distinctcopies of this gene: A1 and A2. In other words, this diploidspecies is A1A1 A2A2. Species B has three copies of the hexokinasegene, which we will call B1, B2, and B3. A diploid individualof species B would be B1B1 B2B2 B3B3. These hexokinase genesfrom the two species were subjected to DNA sequencing, and thepercentage of sequence identity was compared among these genes.The results are shown here.
Percentage of DNA Sequence Identity A1 A2 B1 B2 B3A1 100 62 54 94 53A2 62 100 91 49 92B1 54 91 100 67 90B2 94 49 67 100 64B3 53 92 90 64 100…
Chapter 10 Solutions
Genetics: From Genes to Genomes
Ch. 10 - Prob. 1PCh. 10 - List three independent techniques you could use to...Ch. 10 - Figure 10.2a has numbers indicating the...Ch. 10 - Which of the enzymes from the following list would...Ch. 10 - Prob. 5PCh. 10 - a. What sequence information about a gene is...Ch. 10 - Why do geneticists studying eukaryotic organisms...Ch. 10 - Consider three different kinds of human libraries:...Ch. 10 - The human genome has been sequenced, but we still...Ch. 10 - This problem investigates issues encountered in...
Ch. 10 - For the sake of simplicity, Fig. 10.4 omitted one...Ch. 10 - Give two different reasons for the much higher...Ch. 10 - Using a cDNA library, you isolated two different...Ch. 10 - The figure that follows shows part of a modified...Ch. 10 - In Problem 14, cDNAs F and G could not be found in...Ch. 10 - Fig. 10.10 presents a model for exon shuffling in...Ch. 10 - An interesting phenomenon found in vertebrate DNA...Ch. 10 - a. If you found a zinc-finger domain which...Ch. 10 - Prob. 19PCh. 10 - In the human immune system, so-called B cells can...Ch. 10 - Chimpanzees have a set of hemoglobin genes very...Ch. 10 - Complete genome sequences indicate that the human...Ch. 10 - On your computers browser, view the page accessed...Ch. 10 - Prob. 24PCh. 10 - Prob. 25PCh. 10 - Certain individuals with mild forms of...Ch. 10 - The 1 and 2 genes in humans are identical in their...Ch. 10 - Prob. 28P
Knowledge Booster
Learn more about
Need a deep-dive on the concept behind this application? Look no further. Learn more about this topic, biology and related others by exploring similar questions and additional content below.Similar questions
- Several Drosophila species with unspotted wings are descended from a spotted ancestor. Would you predict the loss of spot formation to entail coding or noncoding changes in pigmentation genes? How would you test which is the case?arrow_forwardIf myoglobin is found in all chordates, urochordates, and cephalochordates, b-globin is found in all vertebrates, a-globin in all ostracoderm descendants, z-globin in all gnathostomes, e-globin in all viviparous mammals, and g-globin in all placental mammals, then the most ancient of these genes: must be epsilon-globin (e-globin), found only in descendants of the ancestral marsupials must be myoglobin, found only in descendants of the chordates, urochordates, and cephalochordates must be gamma-globin (g-globin), found only in descendants of the ancestral placentals must be zeta-globin (z-globin), found only in descendants of the ancestral gnathostome fishes must be beta-globin (b-globin), found only in descendants of the ancestral vertebratesarrow_forward. Figure 18-14 presents haplotype data for the G6PD genein a worldwide sample of people.a. Draw a haplotype network for these haplotypes.Label the branches on which each SNP occurs.b. Which of the haplotypes has the most connectionsto other haplotypes?c. On what continents is this haplotype found?d. Counting the number of SNPs along the branchesof your network, how many differences are therebetween haplotypes 1 and 12?arrow_forward
- In a study of a muscle disorder, several affected families exhibitedvision problems, muscle weakness, and deafness (M. Zeviani et al. 1990.American Journal of Human Genetics 47:904–914). Analysis of themtDNA from affected members of these families revealed that largenumbers of their mtDNA molecules possessed deletions of varyinglengths. Different members of the same family and even differentmitochondria from the same person possessed deletions of differentsizes, so the underlying defect appeared to be a tendency for the mtDNAof affected persons to have deletions. A pedigree of one of the familiesstudied is shown below. The researchers concluded that this disorder isinherited as an autosomal dominant trait, and they mapped the diseasecausinggene to a position on chromosome 10 in the nucleus.a. What characteristics of the pedigree rule out inheritance of a traitencoded by a gene in the mtDNA as the cause of this disorder?b. Explain how a mutation in a nuclear gene might lead to deletions…arrow_forwardFor each of the following examples, discuss whether the observed result is due to neutral mutations or mutations that have been acted on by natural selection, or both: A. When comparing sequences of homologous genes, differences in the coding sequence are most common at the wobble base (i.e., the third base in each codon). B. For a protein-encoding gene, the regions that encode portions of the polypeptide that are vital for structure and function are less likely to display mutations than other regions of the gene. C. When comparing the sequences of homologous genes, introns usually have more sequence differences than exons.arrow_forwardWhat would you predict to be the phenotype of a Drosophila larva whose mother was homozygous for a loss-of-function allele in the nanos gene?arrow_forward
- You are a researcher studying birds in an Indonesian rainforest. You have just discovered two new species whose beaks are markedly different, which you have named Laetiphonia orthorhynchus and Laetiphonia rhamphis. In particular, the beaks of L. orthorhynchus are very long, straight and pointed, whereas L. rhamphis have beaks that are quite short, wide and curved downwards. In further studies, you find that the same gene codes for beak shape in both species. In your own words, explain at least two ways that changes in gene expression could result in the differences you observe between these two species. Make sure to be specific in how your explanation applies to the bird species in this example.arrow_forwardThe most common allele of the β chain of hemoglobin in humans, ???,HbA, has glutamic acid at the sixth position. Two different amino acid substitutions at the sixth position produce two variants, ???HbS and ???,HbC, of the sickle‑cell allele. The ???HbS allele occurs when valine is substituted for glutamic acid, and the ???HbC allele occurs when lysine is substituted for glutamic acid. To answer the question, refer to the amino acid table. Match the class of amino acid and the chemical property to each possible amino acid. Not all answers will be placed.arrow_forwardA. Why are mammals hard to clone? B. What were the names of the first two cloned cows?arrow_forward
- Match the numbered concepts to the lettered facts. Explain why each pair is the best match. (1) Gene Duplication (2) Molecular Clock (3) Pseudogene (4) SINES and LINES a. Different kinds of globin chains arose over time including myoglobin, alpha globin, beta globin 1, delta globin, and beta globin 2.b. The two human beta globin genes, 1 and 2, differ at 20 base pair sites; only beta gene 1 produces a functional protein.c. In human hemoglobin, the delta polypeptide chain differs by 39 amino acid sites vs. the beta chain.d. The human alpha globin gene and the mouse alpha globin gene are orthologous to each other.arrow_forwardhow can genomes with a relatively small number of genes produce the vast complexity of phenotypes that results in living organisms, including humans?arrow_forwardExplain the likely evolutionary origin of mitochondrial and chloroplast genomes. How have the sizes of the mitochondrial and chloroplast genomes changed since their origin? How has thisoccurred?arrow_forward
arrow_back_ios
SEE MORE QUESTIONS
arrow_forward_ios
Recommended textbooks for you
 Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning
Human Heredity: Principles and Issues (MindTap Co...BiologyISBN:9781305251052Author:Michael CummingsPublisher:Cengage Learning

Human Heredity: Principles and Issues (MindTap Co...
Biology
ISBN:9781305251052
Author:Michael Cummings
Publisher:Cengage Learning
Mechanisms of Genetic Change or Evolution; Author: Scientist Cindy;https://www.youtube.com/watch?v=5FE8WvGzS4Q;License: Standard Youtube License